Abstract
With the rapid development of DNA microarray technology, large amount of genomic data has been generated. Classification of these microarray data is a challenge task since gene expression data are often with thousands of genes but a small number of samples. In this paper, an effective gene selection method is proposed to select the best subset of genes for microarray data with the irrelevant and redundant genes removed. Compared with original data, the selected gene subset can benefit the classification task. We formulate the gene selection task as a manifold regularized subspace learning problem. In detail, a projection matrix is used to project the original high dimensional microarray data into a lower dimensional subspace, with the constraint that the original genes can be well represented by the selected genes. Meanwhile, the local manifold structure of original data is preserved by a Laplacian graph regularization term on the low-dimensional data space. The projection matrix can serve as an importance indicator of different genes. An iterative update algorithm is developed for solving the problem. Experimental results on six publicly available microarray datasets and one clinical dataset demonstrate that the proposed method performs better when compared with other state-of-the-art methods in terms of microarray data classification.

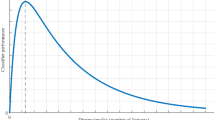
The graphical abstract of this work



Similar content being viewed by others
Notes
CLL_SUB_111 and Lung can be downloaded from: http://featureselection.asu.edu/datasets.php; Breast nd GCM can be downloaded from: http://portals.broadinstitute.org/cgi-bin/cancer/datasets.cgi; Tumors-11 and SRBCT can be downloaded from:http://datam.i2r.a-star.edu.sg/datasets/krbd/index.html.
References
Lj VTV, Dai H, Mj VDV, He YD, Hart AA, Mao M, Peterse HL, Van DKK, Marton MJ, Witteveen AT (2002) Gene expression profiling predicts clinical outcome of breast cancer. Nature 415(6871):530–536
Kolali KM, Bazrafkan M (2016) A novel sparse coding algorithm for classification of tumors based on gene expression data. Med Biol Eng Comput 54(6):869
Kurgan LA, Cios KJ, Tadeusiewicz R, Ogiela M, Goodenday LS (2001) Knowledge discovery approach to automated cardiac spect diagnosis. Artif Intell Med 23(2):149–169
Liao JC, Boscolo R, Yang YL, Tran LM, Sabatti C, Roychowdhury VP (2003) Network component analysis: reconstruction of regulatory signals in biological systems. Proc Natl Acad Sci USA 100(26):15522–15527
Guo S, Guo D, Chen L, Jiang Q (2017) A l1-regularized feature selection method for local dimension reduction on microarray data. Comput Biol Chem 67:92–101
Jiang X, Gao J, Hong X, Cai Z (2014) Gaussian processes autoencoder for dimensionality reduction. In: Pacific-asia conference on knowledge discovery and data mining, pp 62–73
Jiang X, Song X, Gao J, Cai Z, Zhang D (2016) Nonparametrically guided autoencoder with laplace approximation for dimensionality reduction. In: International joint conference on neural networks, pp 3378–3384
Ramos J, Castellanos-Garzón JA, González-Briones A, Paz JFD, Corchado JM (2017) An agent-based clustering approach for gene selection in gene expression microarray. Interdisciplinary Sci Comput Life Sci 9(1):1–13
Wang WZ, Yang BP, Feng CL, Wang JG, Xiong GR, Zhao TT, Zhang SZ (2017) Efficient sugarcane transformation via bar gene selection. Trop Plant Biol 10:1–9
Sharbaf FV, Mosafer S, Moattar MH (2016) A hybrid gene selection approach for microarray data classification using cellular learning automata and ant colony optimization. Genomics 107(6):231
Lv J, Peng Q, Chen X, Sun Z (2016) A multi-objective heuristic algorithm for gene expression microarray data classification. Expert Syst Appl Int J 59:13–19
Wang H, Jing X, Niu B (2017) A discrete bacterial algorithm for feature selection in classification of microarray gene expression cancer data. Know-Based Syst 126:8–19
Zhou LT, Cao YH, Lv LL, Ma KL, Chen PS, Ni HF, Lei XD, Liu BC Feature selection and classification of urinary mrna microarray data by iterative random forest to diagnose renal fibrosis: a two-stage study, Scientific Reports 7
Duda RO, Hart PE, Stork DG (2001) Pattern Classification (2nd Edition). Wiley, New York
Dy JG, Brodley CE (2004) Feature selection for unsupervised learning. J Mach Learn Res 5:845–889
He X, Cai D, Niyogi P (2005) Laplacian score for feature selection. NIPS 18:507–514
Mitra P, Murthy C, Pal SK (2002) Unsupervised feature selection using feature similarity. IEEE Trans Pattern Anal Mach Intell 24(3):301–312
Nie F, Xiang S, Jia Y, Zhang C, Yan S (2008) Trace ratio criterion for feature selection. In: NCAI, pp 671–676
Oh IS, Lee JS, Moon BR (2004) Hybrid genetic algorithms for feature selection. IEEE Trans Pattern Anal Mach Intell 26(11):1424–37
Bolón-Canedo V, Sánchez-Maroño N, Alonso-Betanzos A, Benítez JM, Herrera F (2014) A review of microarray datasets and applied feature selection methods. Inf Sci 282(5):111–135
Cai D, Zhang C, He X (2010) Unsupervised feature selection for multi-cluster data. In: SIGKDD, pp 333–342
Zhao Z, Wang L, Liu H et al (2010) Efficient spectral feature selection with minimum redundancy. In: AAAI, pp 673–678
Zhao Z, Liu H (2007) Spectral feature selection for supervised and unsupervised learning. In: ICML, pp 1151–1157
Li Z, Yang Y, Liu J, Zhou X, Lu H (2012) Unsupervised feature selection using nonnegative spectral analysis. In: NCAI, pp 1026–1032
Golub TR, Slonim DK, Tamayo P, Huard C, Gaasenbeek M, Mesirov JP, Coller H, Loh ML, Downing JR, Caligiuri MA (1999) Molecular classification of cancer: Class discovery and class prediction by gene expression monitoring. Brain Res 501(2):205–14
Thomas JG, Olson JM, Tapscott SJ, Zhao LP (2001) An efficient and robust statistical modeling approach to discover differentially expressed genes using genomic expression profiles. Genome Res 11(7):1227
Dudoit S, Yang YH, Callow MJ, Speed TP (2000) Statistical methods for identifying differentially expressed genes in replicated cdna microarray experiments. Stat sinica 12(1):111–139
Long AD, Mangalam HJ, Chan BY, Tolleri L, Hatfield GW, Baldi P (2001) Improved statistical inference from dna microarray data using analysis of variance and a bayesian statistical framework. analysis of global gene expression in escherichia coli k12. J Biol Chem 276(23):19937–44
Cai R, Hao Z, Yang X, Wen W (2009) An efficient gene selection algorithm based on mutual information. Neurocomputing 72(4-6):991–999
Chuang LY, Yang CH, Li JC, Yang CH (2012) A hybrid bpso-cga approach for gene selection and classification of microarray data. J Comput Biol A J Comput Mol Cell Biol 19(1):68
Wang Y, Tetko IV, Hall MA, Frank E, Facius A, Mayer KFX, Mewes HW (2005) Gene selection from microarray data for cancer classification-a machine learning approach. Comput Biol Chem 29(1):37–46
Gevaert O, Smet FD, Timmerman D, Moreau Y, Moor BD (2006) Predicting the prognosis of breast cancer by integrating clinical and microarray data with bayesian networks. Bioinformatics 22(14):e184—90
Peng H, Long F, Ding C (2005) Feature selection based on mutual information criteria of max-dependency, max-relevance, and min-redundancy. IEEE Trans Pattern Anal Mach Intell 27(8):1226–1238
Battiti R (1994) Using mutual information for selecting features in supervised neural net learning. IEEE Trans Neural Netw 5(4):537–550
Guyon I, Weston J, Barnhill S, Vapnik V (2002) Gene selection for cancer classification using support vector machines. Mach Learn 46(1):389–422
Ghosh D, Chinnaiyan AM (2005) Classification and selection of biomarkers in genomic data using lasso. J Biomed Biotechnol 2005(2):147
Wang YX, Liu JX, Gao YL, Zheng CH, Shang JL (2016) Differentially expressed genes selection via laplacian regularized low-rank representation method. Comput Biol Chem 65(1):185–192
Wang D, Liu JX, Gao YL, Yu J, Zheng CH, Xu Y (2016) An nmf-l2,1-norm constraint method for characteristic gene selection. Plos One 11(7):e0158494
Zheng CH, Ng TY, Zhang D, Shiu CK (2011) Tumor classification based on non-negative matrix factorization using gene expression data. IEEE Trans Nanobioscience 10(2):86–93
Du S, Ma Y, Li S, Ma Y (2017) Robust unsupervised feature selection via matrix factorization. Neurocomputing 241:115–127
Zhu P, Zuo W, Zhang L, Hu Q, Shiu SCK (2015) Unsupervised feature selection by regularized self-representation. Pattern Recogn 48(2):438–446
Shang R, Zhang Z, Jiao L, Liu C, Li Y (2016) Self-representation based dual-graph regularized feature selection clustering. Neurocomputing 171(1):1242–1253
Zhu P, Zhu W, Wang W, Zuo W, Hu Q (2017) Non-convex regularized self-representation for unsupervised feature selection. Image Vis Comput 60(1):22–29
Liu Y, Liu K, Zhang C, Wang J, Wang X (2017) Unsupervised feature selection via diversity-induced self-representation. Neurocomputing 219:350–363
Zhu X, Li X, Zhang S, Ju C, Wu X (2017) Robust joint graph sparse coding for unsupervised spectral feature selection. IEEE Trans Neural Netw Learn Syst 28(6):1263–1275
Lee DD, Seung HS (1999) Learning the parts of objects by non-negativ matrix factorization. Nature 401 (6755):788
Cai D, He X, Han J, Huang TS (2011) Graph regularized nonnegative matrix factorization for data representation. IEEE Trans Pattern Anal Mach Intell 33(8):1548–1560
Belkin M, Niyogi P (2002) Laplacian eigenmaps and spectral techniques for embedding and clustering. Adv Neural Inf Proces Syst 14(6):585–591
He X, Niyogi P (2003) Locality preserving projections. In: Advances in Neural Information Processing Systems, pp 186–197
Hestenes MR (1969) Multiplier and gradient methods. J Optim Theory Appl 4(5):303–320
Ito K, Kunisch K (2010) Lagrange multiplier approach to variational problems and applications. Society for Industrial and Applied Mathematics
Tang C, Wang P, Zhang C, Li W (2017) Salient object detection via weighted low rank matrix recovery. IEEE Signal Process Lett 24(4):490–494
Tang C, Cao L, Chen J, Zheng X (2017) Speckle noise reduction for optical coherence tomography images via non-local weighted group low-rank representation. Laser Phys Lett 14(5):056002
Boyd S, Vandenberghe L (2004) Convex Optimization. Cambridge University Press, Cambridge
Cortes C, Vapnik V (1995) Support-vector networks. Mach Learn 20(3):273–297
Platt JC (1999) Probabilistic outputs for support vector machines and comparisons to regularized likelihood methods. Advances in Large Margin Classifiers 10(4):61–74
Ho TK (2002) Random decision forests. In: International Conference on Document Analysis and Recognition, p 278
Ho TK (1998) The random subspace method for constructing decision forests. IEEE Trans Pattern Anal Mach Intell 20(8):832–844
Altman NS (1992) An introduction to kernel and nearest-neighbor nonparametric regression. Am Stat 46 (3):175–185
Geisser S (1993) Predictive inference : an introduction. Chapman and Hall, London
Kohavi R (1995) A study of cross-validation and bootstrap for accuracy estimation and model selection. In: International Joint Conference on Artificial Intelligence, pp 1137–1143
Devijver PA, Kittler J (1982) Pattern recognition: a statistical approach. Prentice/hall International, New Jersey
Cheng WC, Tsai ML, Chang CW, Huang CL, Chen CR, Shu WY, Lee YS, Wang TH, Hong JH, Li CY (2010) Microarray meta-analysis database (m(2)db): a uniformly pre-processed, quality controlled, and manually curated human clinical microarray database. Bmc Bioinformatics 11(1):421
Guo S, Guo D, Chen L, Jiang Q (2016) A centroid-based gene selection method for microarray data classification. J Theor Biol 400:32–41
Chang CC, Lin CJ (2011) Libsvm: A library for support vector machines. ACM Trans Intell Syst Technol 2(27):1–27
Zhou X, Tuck DP (2007) Msvm-rfe: extensions of svm-rfe for multiclass gene selection on dna microarray data. Bioinformatics 23(9):1106–1114
Cao KAL, Bonnet A, Gadat S (2009) Multiclass classification and gene selection with a stochastic algorithm. Comput Stat Data Anal 53(10):3601–3615
Sun S, Peng Q, Shakoor A (2014) A kernel-based multivariate feature selection method for microarray data classification. Plos One 9(9):e102541
Zhao G, Wu Y Feature subset selection for cancer classification using weight local modularity, Scientific Reports 6
An S, Wang J, Wei J (2017) Local-nearest-neighbors-based feature weighting for gene selection. IEEE/ACM Trans Comput Biol Bioinform PP(99):1–1
Chen KH, Wang KJ, Tsai ML, Wang KM, Adrian AM, Cheng WC, Yang TS, Teng NC, Tan KP, Chang KS (2014) Gene selection for cancer identification: a decision tree model empowered by particle swarm optimization algorithm. Bmc Bioinform 15(1):49
Li X, Li M, Yin M (2016) Multiobjective ranking binary artificial bee colony for gene selection problems using microarray datasets. IEEE/CAA J Automatica Sinica PP(99):1–16
Golub GH, Van Loan CF (1996) Matrix computations (3rd ed.) Johns Hopkins University Press, Baltimore
Acknowledgments
This research was supported by the Fundamental Research Funds for the Central Universities, China University of Geosciences (Wuhan) (No. CUG170654) and the National Natural Science Foundation of China (No. 61701451 and No. 61601261).
Author information
Authors and Affiliations
Corresponding author
Additional information
Chang Tang and Lijuan Cao contributed equally to this work.
Rights and permissions
About this article
Cite this article
Tang, C., Cao, L., Zheng, X. et al. Gene selection for microarray data classification via subspace learning and manifold regularization. Med Biol Eng Comput 56, 1271–1284 (2018). https://doi.org/10.1007/s11517-017-1751-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11517-017-1751-6