Abstract
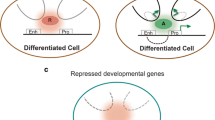
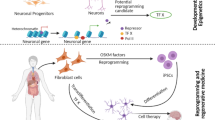
Over 50 years of efforts, cellular reprogramming opens a new door for disease modeling and regenerative medicine. Although induction of pluripotency by transcription factors has become common, only a small portion of basic mechanisms of epigenetic modifications during this process have been revealed. To clearly understand reprogramming and devise ways to promote full transition towards pluripotency, we must gain insight from comprehensive characterizations of cells at distinct reprogramming stages, which involves gene expression profiling, chromatin state maps of key activating and repressive marks, and DNA modifications. Here, we review recent advances in epigenetic reprogramming to pluripotency with a focus on the principal molecular regulators and attach importance to the combination of high-throughput sequencing and systematic biology approaches in uncovering underlying molecular mechanisms of this unique platform in future researches.
Similar content being viewed by others
References
Takahashi K, Yamanaka S (2006) Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126:663–676
Zhao XY, Li W, Lv Z et al (2009) iPS cells produce viable mice through tetraploid complementation. Nature 461:86–90
Boland MJ, Hazen JL, Nazor KL et al (2009) Adult mice generated from induced pluripotent stem cells. Nature 461:U91–U94
Kang L, Wang J, Zhang Y et al (2009) iPS cells can support full-term development of tetraploid blastocyst-complemented embryos. Cell Stem Cell 5:135–138
Soldner F, Hockemeyer D, Beard C et al (2009) Parkinson's disease patient-derived induced pluripotent stem cells free of viral reprogramming factors. Cell 136:964–977
Carvajal-Vergara X, Sevilla A, D'Souza SL et al (2010) Patient-specific induced pluripotent stem-cell-derived models of LEOPARD syndrome. Nature 465:808–812
Raya A, Rodriguez-Piza I, Guenechea G et al (2009) Disease-corrected haematopoietic progenitors from Fanconi anaemia induced pluripotent stem cells. Nature 460:53–59
Lee G, Papapetrou EP, Kim H et al (2009) Modelling pathogenesis and treatment of familial dysautonomia using patient-specific iPSCs. Nature 461:402–406
Ebert AD, Yu J, Rose FF et al (2009) Induced pluripotent stem cells from a spinal muscular atrophy patient. Nature 457:277–280
Smith ZD, Nachman I, Regev A et al (2010) Dynamic single-cell imaging of direct reprogramming reveals an early specifying event. Nat Biotechnol 28:521–526
Samavarchi-Tehrani P, Golipour A, David L et al (2010) Functional genomics reveals a BMP-driven mesenchymal-to-epithelial transition in the initiation of somatic cell reprogramming. Cell Stem Cell 7:64–77
Stadtfeld M, Maherali N, Breault DT et al (2008) Defining molecular cornerstones during fibroblast to iPS cell reprogramming in mouse. Cell Stem Cell 2:230–240
Li R, Liang J, Ni S et al (2010) A mesenchymal-to-epithelial transition initiates and is required for the nuclear reprogramming of mouse fibroblasts. Cell Stem Cell 7:51–63
Brambrink T, Foreman R, Welstead GG et al (2008) Sequential expression of pluripotency markers during direct reprogramming of mouse somatic cells. Cell Stem Cell 2:151–159
Buganim Y, Faddah DA, Cheng AW et al (2012) Single-cell expression analyses during cellular reprogramming reveal an early stochastic and a late hierarchic phase. Cell 150:1209–1222
Hanna J, Saha K, Pando B et al (2009) Direct cell reprogramming is a stochastic process amenable to acceleration. Nature 462:595–601
Yamanaka S (2009) Elite and stochastic models for induced pluripotent stem cell generation. Nature 460:49–52
Rais Y, Zviran A, Geula S et al (2013) Deterministic direct reprogramming of somatic cells to pluripotency. Nature 502:65–70
Le Guezennec X, Vermeulen M, Brinkman AB et al (2006) MBD2/NuRD and MBD3/NuRD, two distinct complexes with different biochemical and functional properties. Mol Cell Biol 26:843–851
Kaji K, Nichols J, Hendrich B (2007) Mbd3, a component of the NuRD co-repressor complex, is required for development of pluripotent cells. Development 134:1123–1132
Guo S, Zi X, Schulz VP et al (2014) Nonstochastic reprogramming from a privileged somatic cell state. Cell 156:649–662
Goldberg AD, Allis CD, Bernstein E (2007) Epigenetics: a landscape takes shape. Cell 128:635–638
Kouzarides T (2007) Chromatin modifications and their function. Cell 128:693–705
Strahl BD, Allis CD (2000) The language of covalent histone modifications. Nature 403:41–45
Koche RP, Smith ZD, Adli M et al (2011) Reprogramming factor expression initiates widespread targeted chromatin remodeling. Cell Stem Cell 8:96–105
Polo JM, Anderssen E, Walsh RM et al (2012) A molecular roadmap of reprogramming somatic cells into iPS cells. Cell 151:1617–1632
Onder TT, Kara N, Cherry A et al (2012) Chromatin-modifying enzymes as modulators of reprogramming. Nature 483:598–602
Mansour AA, Gafni O, Weinberger L et al (2012) The H3K27 demethylase Utx regulates somatic and germ cell epigenetic reprogramming. Nature 488:409–413
Bulger M, Groudine M (2011) Functional and mechanistic diversity of distal transcription enhancers. Cell 144:327–339
Roeder RG (2005) Transcriptional regulation and the role of diverse coactivators in animal cells. FEBS Lett 579:909–915
Rada-Iglesias A, Bajpai R, Swigut T et al (2011) A unique chromatin signature uncovers early developmental enhancers in humans. Nature 470:279–283
Creyghton MP, Cheng AW, Welstead GG et al (2010) Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc Natl Acad Sci USA 107:21931–21936
Heintzman ND, Hon GC, Hawkins RD et al (2009) Histone modifications at human enhancers reflect global cell-type-specific gene expression. Nature 459:108–112
Whyte WA, Bilodeau S, Orlando DA et al (2012) Enhancer decommissioning by LSD1 during embryonic stem cell differentiation. Nature 482:221–225
Lee JE, Wang CC, Xu SLY et al (2013) H3K4 mono- and di-methyltransferase MLL4 is required for enhancer activation during cell differentiation. Elife 2:e01503
Chen J, Liu H, Liu J et al (2013) H3K9 methylation is a barrier during somatic cell reprogramming into iPSCs. Nat Genet 45:34–42
Wang T, Chen K, Zeng X et al (2011) The histone demethylases Jhdm1a/1b enhance somatic cell reprogramming in a vitamin-C-dependent manner. Cell Stem Cell 9:575–587
Esteban MA, Wang T, Qin B et al (2010) Vitamin C enhances the generation of mouse and human induced pluripotent stem cells. Cell Stem Cell 6:71–79
Huangfu D, Maehr R, Guo W et al (2008) Induction of pluripotent stem cells by defined factors is greatly improved by small-molecule compounds. Nat Biotechnol 26:795–797
Huangfu D, Osafune K, Maehr R et al (2008) Induction of pluripotent stem cells from primary human fibroblasts with only Oct4 and Sox2. Nat Biotechnol 26:1269–1275
Mali P, Chou BK, Yen J et al (2010) Butyrate greatly enhances derivation of human induced pluripotent stem cells by promoting epigenetic remodeling and the expression of pluripotency-associated genes. Stem Cells 28:713–720
Liang G, Taranova O, Xia K et al (2010) Butyrate promotes induced pluripotent stem cell generation. J Biol Chem 285:25516–25521
Zhang Z, Wu WS (2013) Sodium butyrate promotes generation of human induced pluripotent stem cells through induction of the miR302/367 cluster. Stem Cells Dev 22:2268–2277
Tonge PD, Corso AJ, Monetti C et al (2014) Divergent reprogramming routes lead to alternative stem-cell states. Nature 516:192–197
Hussein SM, Puri MC, Tonge PD et al (2014) Genome-wide characterization of the routes to pluripotency. Nature 516:198–206
Yang PK, Kuroda MI (2007) Noncoding RNAs and intranuclear positioning in monoallelic gene expression. Cell 128:777–786
Wu H, Zhang Y (2011) Mechanisms and functions of Tet protein-mediated 5-methylcytosine oxidation. Genes Dev 25:2436–2452
Pastor WA, Aravind L, Rao A (2013) TETonic shift: biological roles of TET proteins in DNA demethylation and transcription. Nat Rev Mol Cell Biol 14:341–356
Kim J, Woo AJ, Chu J et al (2010) A Myc network accounts for similarities between embryonic stem and cancer cell transcription programs. Cell 143:313–324
Sridharan R, Tchieu J, Mason MJ et al (2009) Role of the murine reprogramming factors in the induction of pluripotency. Cell 136:364–377
Mikkelsen TS, Hanna J, Zhang X et al (2008) Dissecting direct reprogramming through integrative genomic analysis. Nature 454:49–55
Piccolo FM, Bagci H, Brown KE et al (2013) Different roles for Tet1 and Tet2 proteins in reprogramming-mediated erasure of imprints induced by EGC fusion. Mol Cell 49:1023–1033
Hu X, Zhang L, Mao SQ et al (2014) Tet and TDG mediate DNA demethylation essential for mesenchymal-to-epithelial transition in somatic cell reprogramming. Cell Stem Cell 14:512–522
Doege CA, Inoue K, Yamashita T et al (2012) Early-stage epigenetic modification during somatic cell reprogramming by Parp1 and Tet2. Nature 488:652–655
Costa Y, Ding J, Theunissen TW et al (2013) NANOG-dependent function of TET1 and TET2 in establishment of pluripotency. Nature 495:370–374
Gao Y, Chen J, Li K et al (2013) Replacement of Oct4 by Tet1 during iPSC induction reveals an important role of DNA methylation and hydroxymethylation in reprogramming. Cell Stem Cell 12:453–469
Chen J, Gao Y, Huang H et al (2014) The combination of Tet1 with Oct4 generates high-quality mouse induced pluripotent stem cells (iPSCs). Stem Cells 33:686–698
Jackson SA, Sridharan R (2013) The nexus of Tet1 and the pluripotency network. Cell Stem Cell 12:387–388
Papp B, Plath K (2013) Epigenetics of reprogramming to induced pluripotency. Cell 152:1324–1343
Maherali N, Sridharan R, Xie W et al (2007) Directly reprogrammed fibroblasts show global epigenetic remodeling and widespread tissue contribution. Cell Stem Cell 1:55–70
Han DW, Greber B, Wu G et al (2011) Direct reprogramming of fibroblasts into epiblast stem cells. Nat Cell Biol 13:66–71
Nichols J, Smith A (2009) Naive and primed pluripotent states. Cell Stem Cell 4:487–492
Chen Q, Gao S, He W et al (2014) Xist repression shows time-dependent effects on the reprogramming of female somatic cells to induced pluripotent stem cells. Stem Cells 32:2642–2656
Hoffman LM, Hall L, Batten JL et al (2005) X-inactivation status varies in human embryonic stem cell lines. Stem Cells 23:1468–1478
Shen Y, Matsuno Y, Fouse SD et al (2008) X-inactivation in female human embryonic stem cells is in a nonrandom pattern and prone to epigenetic alterations. Proc Natl Acad Sci USA 105:4709–4714
Silva SS, Rowntree RK, Mekhoubad S et al (2008) X-chromosome inactivation and epigenetic fluidity in human embryonic stem cells. Proc Natl Acad Sci USA 105:4820–4825
Anguera MC, Sadreyev R, Zhang Z et al (2012) Molecular signatures of human induced pluripotent stem cells highlight sex differences and cancer genes. Cell Stem Cell 11:75–90
Mekhoubad S, Bock C, de Boer AS et al (2012) Erosion of dosage compensation impacts human iPSC disease modeling. Cell Stem Cell 10:595–609
Tomoda K, Takahashi K, Leung K et al (2012) Derivation conditions impact X-inactivation status in female human induced pluripotent stem cells. Cell Stem Cell 11:91–99
Orkin SH, Hochedlinger K (2011) Chromatin connections to pluripotency and cellular reprogramming. Cell 145:835–850
Skene PJ, Henikoff S (2013) Histone variants in pluripotency and disease. Development 140:2513–2524
Shinagawa T, Takagi T, Tsukamoto D et al (2014) Histone variants enriched in oocytes enhance reprogramming to induced pluripotent stem cells. Cell Stem Cell 14:217–227
Gaspar-Maia A, Qadeer ZA, Hasson D et al (2013) MacroH2A histone variants act as a barrier upon reprogramming towards pluripotency. Nat Commun 4:1565
Buganim Y, Markoulaki S, van Wietmarschen N et al (2014) The developmental potential of iPSCs is greatly influenced by reprogramming factor selection. Cell Stem Cell 15:295–309
Wu T, Liu Y, Wen D et al (2014) Histone variant H2A.X deposition pattern serves as a functional epigenetic mark for distinguishing the developmental potentials of iPSCs. Cell Stem Cell 15:281–294
Gao Y, Gao S (2014) Quality control: H2A.X links to better iPSCs. Cell Stem Cell 15:259–260
Kagey MH, Newman JJ, Bilodeau S et al (2010) Mediator and cohesin connect gene expression and chromatin architecture. Nature 467:430–435
Schoenfelder S, Sexton T, Chakalova L et al (2010) Preferential associations between co-regulated genes reveal a transcriptional interactome in erythroid cells. Nat Genet 42:53–61
Gaspar-Maia A, Alajem A, Meshorer E et al (2011) Open chromatin in pluripotency and reprogramming. Nat Rev Mol Cell Biol 12:36–47
Meshorer E, Yellajoshula D, George E et al (2006) Hyperdynamic plasticity of chromatin proteins in pluripotent embryonic stem cells. Dev Cell 10:105–116
Lieberman-Aiden E, van Berkum NL, Williams L et al (2009) Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326:289–293
Dixon JR, Selvaraj S, Yue F et al (2012) Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485:376–380
de Wit E, Bouwman BA, Zhu Y et al (2013) The pluripotent genome in three dimensions is shaped around pluripotency factors. Nature 501:227–231
Apostolou E, Ferrari F, Walsh RM et al (2013) Genome-wide chromatin interactions of the Nanog locus in pluripotency, differentiation, and reprogramming. Cell Stem Cell 12:699–712
Wei Z, Gao F, Kim S et al (2013) Klf4 organizes long-range chromosomal interactions with the oct4 locus in reprogramming and pluripotency. Cell Stem Cell 13:36–47
Sexton T, Cavalli G (2013) The 3D genome shapes up for pluripotency. Cell Stem Cell 13:3–4
Acknowledgments
This work was supported by the National Natural Science Foundation of China (31325019, 91319306 and 31401247) and Ministry of Science and Technology of China (2015CB964800 and 2014CB964601). We are grateful to our colleagues in the laboratory for their assistance with the preparation of this manuscript.
Conflict of interest
The authors declare that they have no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
SPECIAL TOPIC: Stem cell, Basis and Application
About this article
Cite this article
Gao, R., Liu, X. & Gao, S. Progress in understanding epigenetic remodeling during induced pluripotency. Sci. Bull. 60, 1713–1721 (2015). https://doi.org/10.1007/s11434-015-0919-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11434-015-0919-4