Abstract
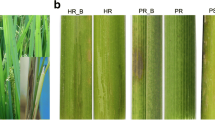
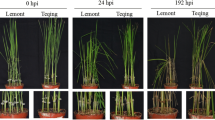
Rice blast is one of the most devastating diseases in the world and outbreaks occur frequently. The differences in protein expression between blast resistant and susceptible near-isogenic lines (NILs) of japonica rice var. Yunyin infected with Magnaporthe oryzae were analyzed using proteomics, indicated that 67 different proteins were identified from 75 obtained proteins using the matrix-assisted laser desorption/ionization time-of-flight mass spectrometry technique. Seven specific expression proteins, 43 up-regulated proteins and 17 down-regulated proteins were identified among the 67 proteins. The bioinformatical analysis demonstrated that these 67 different proteins were involved in many biological physiological processes including five proteins related to photosynthesis, 25 proteins related to metabolism, six proteins related to anti-oxidants, 10 proteins related to protein synthesis and modification, five proteins related to signal transduction, four proteins related to adversity stress and 12 non-functional proteins. These identified proteins were directly or indirectly related to stress. Five proteins related to different physiological processes were selected. Their cDNA sequences were predicted and their expression patterns were analyzed using real-time PCR, demonstrated that the genes would response to M. oryzae and the response blindingly different between blast resistant and blast susceptible NILs.





Similar content being viewed by others
References
Ventelon-Debout M, Delalande F, Brizard JP et al (2004) Proteome analysis of cultivar-specific degradation of Oryza sativa indica and O. sativa japonica cellular suspensions undergoing rice yellow mottle virus infection. Proteomics 4:216–225
Tanaka N, Konishi H, Khan MM et al (2004) Proteome analysis of rice tissues by two-dimensional electrophoresis: an approach to the investigation of gibberellin regulated proteins. Mol Gen Genomics 270:485–496
Rakwal R, Komatsu S (2000) Role of jasmonate in the rice (Oryza sativa L.) self-defense mechanism using proteome analysis. Electrophoresis 21:2492–2500
Konishi H, Komatsu S (2003) A proteomics approach to investigating promotive effects of brassinolide on lamina inclination and root growth in rice seedlings. Biol Pharm Bull 26:401–408
Yang G, Inoue A, Takasaki H et al (2005) A proteomic approach to analyze auxin-and zinc-responsive protein in rice. J Proteome Res 4:456–463
Leone A, Costa A, Tucci M et al (1994) Comparative analysis of short-and long-term changes in gene expression caused by low water potential in potato (Solanum tuberosum) cell-suspension cultures. Plant Physiol 106:703–712
Ramagopal S (1987) Salinity stress induced tissue-specific proteins in barley seedlings. Plant Physiol 84:324–331
Zhu YS, Cai QH, Xue WM et al (2012) Preliminary genotype analysis of broad-spectrum rice blast resistance variety yunyin and its near-isogenic lines construction. Fujian J Agric Sci 27:1–6
Zhang W, Wang Z, Cai Y et al (2004) Life and reproductivity tables of Magnaporthe grisea in CO39 near-isogenic rice lines. Acta Phytopathol Sin 34:61–68
Li YF, Zhang ZH, Nie YF et al (2012) Proteomic analysis of salicylic acid-induced resistance to Magnaporthe oryzae in susceptible and resistant rice. Proteomics 12:2340–2354
Damerval C, de Vienne D, Zivy M et al (1986) Paper symposium: two dimensional electrophoresis of plant proteins. Electrophoresis 7:52–54
Giavalisco P, Nordhoff E, Lehrach H et al (2003) Extraction of proteins from plant tissues for two-dimensional electrophoresis analysis. Electrophoresis 24:207–216
Lowry OH, Rosebrough NJ, Farr AL et al (1951) Protein measurement with the Folin reagent. J Biol Chem 193:265–275
Bevan M, Bancroft I, Bent E et al (1998) Analysis of 1.9 Mb of contiguous sequence from chromosome 4 of Arabidopsis thaliana. Nature 391:485–488
Sasaki T, Burr B (2005) International rice genome sequencing project: the effort to completely sequence the rice genome. Curr Opin Plant Biol 3:138–141
Chen DX, Chen XW, Ma BT et al (2010) Genetic transformation of rice with Pi-d2 gene enhances resistance to rice blast fungus Magnaporthe oryzae. Rice Sci 17:19–27
Kim S, Ahn IP, Lee YH (2001) Analysis of genes expressed during rice–Magnaporthe oryzae interactions. Mol Plant Microbe Interact 14:1340–1346
Nishizawa Y, Hirai A (1989) The nucleotide sequences and expression of genes for the beta and epsilon subunits of ATP synthase from rice (Oryza sativa L.). Jpn J Genet 64:223–229
Forsthoefel NR, Cushman MA, Cushman JC (1995) Posttranscriptional and posttranslational control of enolase expression in the facultative Crassulacean acid metabolism plant Mesembryanthemum crystallinum L. Plant Physiol 108:1185–1195
Norbeck J, Blomberg A (1997) Metabolic and regulatory changes associated with growth of Saccharomyces cerevisiae in 1.4 M NaCl. J Biol Chem 272:5544–5554
Lal SK, Lee C, Sachs MM (1998) Differential regulation of enolase during anaerobiosis in maize. Plant Physiol 118:1285–1293
Lowry OH, Rosebrough NJ, Farr AL (1951) Protein measurement with the Folin reagent. J Biol Chem 193:265–275
Neuhoff V, Arold N, Taube D et al (1988) Improved staining of proteins in polyacrylamide gels including isoelectric focusing gels with clear background at nanogram sensitivity using Coomassie Brilliant Blue G-250 and R-250. Electrophoresis 9:255–262
Asada K, Takehashi M (1987) Production and scavenging of active oxygen in photosynthesis. Photoinhibition 9:227–288
Asada K (1992) Ascorbate peroxidase-a hydrogen peroxide-scavenging enzyme in plants. Physiol Plantarum 85:235–241
Acknowledgements
This work was supported by the National Basic Research Program of China (2012CB723003), the National Natural Sciences Foundation of China (30871509) and the Fujian Provincial Sciences Foundation (2007J0005, 2009J06011).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Weimin Xue and Xiaohui Mao have contributed equally to this work
About this article
Cite this article
Xue, W., Mao, X., Wei, Y. et al. Proteomic analysis of blast-resistant near-isogenic lines derived from japonica rice, var. Yunyin, infected with Magnaporthe oryzae . Chin. Sci. Bull. 59, 4312–4322 (2014). https://doi.org/10.1007/s11434-014-0447-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11434-014-0447-7