Abstract
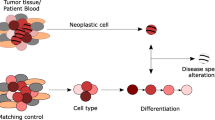
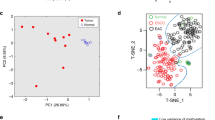
There are several kinds of epigenetic networks in the human body including the cell differentiation epigenetic network (DiEN) and the host adaptation epigenetic network (AdEN). DiEN networks are static and cell/tissue-specific. AdEN networks are variable and dependent upon environmental factors. DiEN and AdEN alterations can respectively serve as biomarkers for different kinds of diseases. Cancer is a consequence of accumulated pathophysiological adaptations of tissue stem cells to exposure of environmental carcinogens. Cancer cells are de-differentiated cells that obtain the capacity of unrestricted proliferation, local invasion, and distant migration/metastasis. Both DiEN and AdEN changes can be observed in cancer tissues. Alterations of DNA methylation are the most stable epigenetic modifications and can be sensitively detected in a small cell population. These advantages make DNA methylation the optimal biomarkers for detection of initiated cells in precancerous lesions and metastasis stem cells in cancer tissues. It has been proven that p16 methylation can be used as a diagnostic biomarker to determine malignant potential of epithelium dysplasia in many organs including the stomach. In a large-scale validation study on the DNA methylome of gastric carcinomas (GC), the methylation status of more than 90 CpG islands has been analyzed by DHPLC. Furthermore, GFRA1 demethylation and methylation of SRF and ZNF382 are frequent events during gastric carcinogenesis and consistently correlate to GC metastasis and overall survival of GC patients from China, Japan, and Korea, respectively. In a population study, it has been demonstrated that gradual increasing of plasma miR-211 and other miRNA levels may be an early risk predictor for GC development.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Hemberger M, Dean W, Reik W. Epigenetic dynamics of stem cells and cell lineage commitment: Digging Waddington’s canal. Nat Rev Mol Cell Biol, 2009, 10: 526–537
Faul K C, Dolinoy D C. Timing is everything: The when and how of environmentally induced changes in the epigenome of animals. Epigenetics, 2011, 6: 791–797
The ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature, 2012, 489: 57–74
Deng D J, Liu Z J, Du Y T. Epigenetic alterations as cancer diagnostic, prognostic, and predictive biomarkers. Adv Genet, 2010, 71: 125–176
Sun Y, Deng D, You W C, et al. Methylation of p16 CpG islands associated with malignant transformation of gastric dysplasia in a population-based study. Clin Cancer Res, 2004, 10: 5087–5093
Cao J, Zhou J, Gao Y, et al. Methylation of p16 CpG island associated with malignant progression of oral epithelial dysplasia: A prospective cohort study. Clin Cancer Res, 2009, 15: 5178–5183
Hall G L, Shaw R J, Field E A, et al. p16 Promoter methylation is a potential predictor of malignant transformation in oral epithelial dysplasia. Cancer Epidemiol Biomarkers Prev, 2008, 17: 2174–2179
Belinsky S A, Liechty K C, Gentry F D, et al. Promoter hypermethylation of multiple genes in sputum precedes lung cancer incidence in a high-risk cohort. Cancer Res, 2006, 66: 3338–3344
Jin Z, Cheng Y, Gu W, et al. A multicenter, double-blinded validation study of methylation biomarkers for progression prediction in Barrett’s esophagus. Cancer Res, 2009, 69: 4112–4115
Zhou J, Cao J, Lu Z M, et al. A 115-bp MethyLight assay for detection of p16 (CDKN2A) methylation as a diagnostic biomarker in human tissues. BMC Med Genet, 2011, 12: 67
Lister R, Pelizzola M, Dowen R H, et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature, 2009, 462: 315–322
Laurent L, Wong E, Li G, et al. Dynamic changes in the human methylome during differentiation. Genome Res, 2010, 20: 320–331
Chodavarapu R K, Feng S, Bernatavichute Y V, et al. Relationship between nucleosome positioning and DNA methylation. Nature, 2010, 466: 388–392
Barski A, Cuddapah S, Cui K, et al. High-resolution profiling of histone methylations in the human genome. Cell, 2007, 129: 823–837
Statham A L, Robinson M D, Song J Z, et al. Bisulphite-sequencing of chromatin immunoprecipitated DNA (BisChIP-seq) directly informs methylation status of histone-modified DNA. Genome Res, 2012, 22: 1120–1127
Brinkman A B, Gu H, Bartels S J J, et al. Sequential ChIP-bisulfite sequencing enables direct genome-scale investigation of chromatin and DNA methylation cross-talk. Genome Res, 2012, 22: 1128–1138
Bhutani N, Burns D M, Blau H M. DNA demethylation dynamics. Cell, 2011, 146: 866–872
Xu Y, Wu F, Tan L, et al. Genome-wide regulation of 5hmC, 5mC, and gene expression by Tet1 hydroxylase in mouse embryonic stem cells. Mol Cell, 2011, 42: 451–464
Ficz G, Branco M R, Seisenberger S, et al. Dynamic regulation of 5-hydroxymethylcytosine in mouse ES cells and during differentiation. Nature, 2011, 473: 398–402
Irizarry R A, Ladd-Acosta C, Wen B, et al. The human colon cancer methylome shows similar hypo- and hypermethylation at conserved tissue-specific CpG island shores. Nat Genet, 2009, 41: 178–186
Doi A, Park I H, Wen B, et al. Differential methylation of tissue- and cancer-specific CpG island shores distinguishes human induced pluripotent stem cells, embryonic stem cells and fibroblasts. Nat Genet, 2009, 41: 1350–1353
Yang P, Ma J, Zhang B, et al. CpG Site-specific hypermethylation of p16INK4? in peripheral blood lymphocytes of PAH-exposed workers. Cancer Epidemiol Biomarkers Prev, 2012, 21: 182–190
Yan P S, Shi H, Rahmatpanah F, et al. Differential distribution of DNA methylation within the RASSF1A CpG island in breast cancer. Cancer Res, 2003, 63: 6178–6186
Lu Z M, Zhou J, Wang X, et al. Nucleosomes correlate with in vivo progression pattern of de novo methylation of p16 CpG islands in human gastric carcinogenesis. PLoS One, 2012, 7: e35928
Bai H, Zhou J, Deng D J. Nucleosome positions and differential methylation status of various regions within MLH1 CpG island. Chin J Cancer Res, 2008, 20: 237–242
Egger G, Aparicio A M, Escobar S G, et al. Inhibition of histone deacetylation does not block resilencing of p16 after 5-aza-2′-deoxycytidine treatment. Cancer Res, 2007, 67: 346–353
De Carvalho D D, Sharma S, You J S, et al. DNA methylation screening identifies driver epigenetic events of cancer cell survival. Cancer Cell, 2012, 21: 655–667
Costello J F, Frühwald M C, Smiraglia D J, et al. Aberrant CpG-island methylation has non-random and tumour-type-specific patterns. Nat Genet, 2000, 24: 132–138
Gitan R S, Shi H, Chen C M, et al. Methylation-specific oligonucleotide microarray: A new potential for high-throughput methylation analysis. Genome Res, 2002, 12: 158–164
Yu L, Liu C, Vandeusen J, et al. Global assessment of promoter methylation in a mouse model of cancer identifies ID4 as a putative tumor-suppressor gene in human leukemia. Nat Genet, 2005, 37: 265–274
Shen L, Kondo Y, Guo Y, et al. Genome-wide profiling of DNA methylation reveals a class of normally methylated CpG island promoters. PLoS Genet, 2007, 3: 2023–2036
Liu Z J, Gao Y H, Zhang J, et al. DNA methylation markers as predictors of initiation and progression of gastric carcinomas. In: Noh H S, Mok Y J, Yang H K, eds. 9th International Gastric Cancer Congress. Pianoro, Bologna, Italy: MEDIMOND s.r.l., 2011. 13–17
Deng D J, Deng G R, Smith M F, et al. Simultaneous detection of CpG methylation and single nucleotide polymorphism by denaturing high performance liquid chromatography. Nucleic Acids Res, 2002, 30: 13E
Luo D, Zhang B, Lv L, et al. Methylation of CpG islands of p16 associated with progression of primary gastric carcinomas. Lab Invest, 2006, 86: 591–598
Deng D, Liu Z. Methods and nucleotide fragments of predicting the ability of tumor invasion and metastasis in vitro (in Chinese). PCT patent, WO2011095067, 2011-03-30
Deng D, Gao Y, Zhang B, et al. Methods and nucleotides for assessing tumor metastasis or prognosis in vitro (in Chinese). PCT patent, WO2012097477, 2011-01-21
Deng D, Zhang J, Liu Z, et al. Predicting ability of tumor development, metastasis, and patient’s survival, by extracting DNA from biological samples, detecting demethylation degree of cytosine in CpG island of DNA sequence of GFRA1 and determining tested samples (in Chinese). PCT patent, PCT/CN2012/000169, 2011-03-08
Du Y, Liu Z, Gu L, et al. Characterization of human gastric carcinoma-related methylation of 9 miR CpG islands and repression of their expressions in vitro and in vivo. BMC Cancer, 2012, 12: 249
Dong C X, Deng D J, Pan K F, et al. Promoter methylation of p16 associated with Helicobacter pylori infection in precancerous gastric lesions: A population-based study. Int J Cancer, 2009, 124: 434–439
Niwa T, Tsukamoto T, Toyoda T, et al. Inflammatory processes triggered by Helicobacter pylori infection cause aberrant DNA methylation in gastric epithelial cells. Cancer Res, 2010, 70: 1430–1440
Song M Y, Pan K F, Su H J, et al. Identification of serum microRNAs as novel non-invasive biomarkers for early detection of gastric cancer. PLoS One, 2012, 7: e33608
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at Springerlink.com
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Deng, D., Lu, Z. Differentiation and adaptation epigenetic networks: Translational research in gastric carcinogenesis. Chin. Sci. Bull. 58, 1–6 (2013). https://doi.org/10.1007/s11434-012-5578-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11434-012-5578-0