Abstract
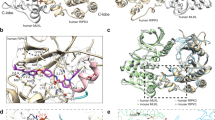
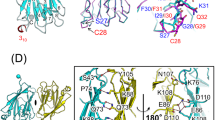
RNA-dependent protein kinase (PKR) is crucial for the innate immune response, cell growth, proliferation, signal transduction and apoptosis. The activation process of PKR has been studied for many years and is still under debate. To obtain new insight into the mechanism of PKR activation, we solved the crystal structure of a latent mutant of the PKR kinase domain (PKR-KD) in the apo form at a resolution of 2.9 Å. The overall structure of PKR-KD is similar to previously reported structures. Structural analysis revealed a classical back-to-back dimer and a newly defined face-to-face dimer. Analytical ultracentrifugation experiments, electrostatic surface maps and the model of PKR-KD in complex with the eIF2α substrate all support that the face-to-face dimer is more reflective of PKR in solution. Our results provide new information on PKR dimerization and its activation mechanism.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Shi Y. Serine/threonine phosphatases: Mechanism through structure. Cell, 2009, 139: 468–484
Nanduri S, Rahman F, Williams B R, et al. A dynamically tuned double-stranded RNA binding mechanism for the activation of antiviral kinase PKR. EMBO J, 2000, 19: 5567–5574
Taghavi N, Samuel C E. Protein kinase PKR catalytic activity is required for the PKR-dependent activation of mitogen-activated protein kinases and amplification of interferon beta induction following virus infection. Virology, 2012, 427: 208–216
Pfaller C K, Li Z, George C X, et al. Protein kinase PKR and RNA adenosine deaminase ADAR1: New roles for old players as modulators of the interferon response. Curr Opin Immunol, 2011, 23: 573–582
Dever T E. Gene-specific regulation by general translation factors. Cell, 2002, 108: 545–556
Dey M, Trieselmann B, Locke E G, et al. PKR and GCN2 kinases and guanine nucleotide exchange factor eukaryotic translation initiation factor 2B (eIF2B) recognize overlapping surfaces on eIF2alpha. Mol Cell Biol, 2005, 25: 3063–3075
Nallagatla S R, Toroney R, Bevilacqua P C. Regulation of innate immunity through RNA structure and the protein kinase PKR. Curr Opin Struct Biol, 2011, 21: 119–127
Nakamura T, Furuhashi M, Li P, et al. Double-stranded RNA-dependent protein kinase links pathogen sensing with stress and metabolic homeostasis. Cell, 2010, 140: 338–348
Dar A C, Dever T E, Sicheri F. Higher-order substrate recognition of eIF2alpha by the RNA-dependent protein kinase PKR. Cell, 2005, 122: 887–900
Robertson H D, Mathews M B. The regulation of the protein kinase PKR by RNA. Biochimie, 1996, 78: 909–914
Cole J L. Activation of PKR: An open and shut case? Trends Biochem Sci, 2007, 32: 57–62
Dey M, Cao C, Dar A C, et al. Mechanistic link between PKR dimerization, autophosphorylation, and eIF2alpha substrate recognition. Cell, 2005, 122: 901–913
Romano P R, Garcia-Barrio M T, Zhang X, et al. Autophosphorylation in the activation loop is required for full kinase activity in vivo of human and yeast eukaryotic initiation factor 2alpha kinases PKR and GCN2. Mol Cell Biol, 1998, 18: 2282–2297
McKenna S A, Lindhout D A, Shimoike T, et al. Viral dsRNA inhibitors prevent self-association and autophosphorylation of PKR. J Mol Biol, 2007, 372: 103–113
VanOudenhove J, Anderson E, Krueger S, et al. Analysis of PKR structure by small-angle scattering. J Mol Biol, 2009, 387: 910–920
Otwinowski Z, Minor W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol, 1997, 276: 307–326
McCoy A J, Grosse-Kunstleve R W, Adams P D, et al. Phaser crystallographic software. J Appl Crystallogr, 2007, 40: 658–674
Emsley P, Lohkamp B, Scott W G, et al. Features and development of coot. Acta Crystallogr D Biol Crystallogr, 2010, 66: 486–501
Adams P D, Afonine P V, Bunkoczi G, et al. PHENIX: A comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr D Biol Crystallogr, 2010, 66: 213–221
Laskowski R A, MacArthur M W, Moss D S, et al. PROCHECK: A program to check the stereochemical quality of protein structures. J Appl Crystallogr, 1993, 26: 283–291
Schuck P. Size-distribution analysis of macromolecules by sedimentation velocity ultracentrifugation and Lamm equation modeling. Biophys J, 2000, 78: 1606–1619
Winn M D, Ballard C C, Cowtan K D, et al. Overview of the CCP4 suite and current developments. Acta Crystallogr D Biol Crystallogr, 2011, 67: 235–242
Author information
Authors and Affiliations
Corresponding authors
Additional information
These authors contributed equally to this work.
This article is published with open access at Springerlink.com
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Li, F., Li, S., Wang, Z. et al. Structure of the kinase domain of human RNA-dependent protein kinase with K296R mutation reveals a face-to-face dimer. Chin. Sci. Bull. 58, 998–1002 (2013). https://doi.org/10.1007/s11434-012-5461-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11434-012-5461-z