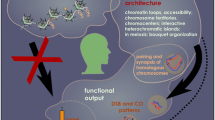
Abstract
Unlike animals, plants do not set aside germ cells early in development. In angiosperm species, reproduction occurs in the adult plant upon flowering. The multicellular male and female gametophytes differentiate from meiotic products within reproductive floral organs. Double fertilization is another remarkable feature of most angiosperm species. The zygote derived from fertilization of the egg cell by one of the sperm cells and the endosperm from fertilization of the central cell by the second sperm cell develop in a coordinated manner together and enclosed in the sporophytic maternal integuments, forming the seed. Understanding plant reproduction is biologically pertinent and agronomically and ecologically important. Here, we describe the known functions of histone lysine methylations in various steps of reproduction in the reference plant Arabidopsis thaliana. It is emerging that histone lysine methylation is key for understanding epigenetic regulation networks of genome function.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Johnson L, Mollah S, Garcia B A, et al. Mass spectrometry analysis of Arabidopsis histone H3 reveals distinct combinations of posttranslational modifications. Nucleic Acids Res, 2004, 32: 6511–6518
Martin C, Zhang Y. The diverse functions of histone lysine methylation. Nat Rev Mol Cell Biol, 2005, 6: 838–849
Baumbusch L O, Thorstensen T, Krauss V, et al. The Arabidopsis thaliana genome contains at least 29 active genes encoding SET domain proteins that can be assigned to four evolutionarily conserved classes. Nucleic Acids Res, 2001, 29: 4319–4333
Liu C, Lu F, Cui X, et al. Histone methylation in higher plants. Annu Rev Plant Biol, 2010, 61: 395–420
Yu Y, Pu Z, Shen W H, et al. An update on histone lysine methylation in plants. Prog Nat Sci, 2009, 19: 407–413
Baurle I, Dean C. The timing of developmental transitions in plants. Cell, 2006, 125: 655–664
Michaels S D, Amasino R M. FLOWERING LOCUS C encodes a novel MADS domain protein that acts as a repressor of flowering. Plant Cell, 1999, 11: 949–956
Sheldon C C, Burn J E, Perez P P, et al. The FLF MADS box gene: A repressor of flowering in Arabidopsis regulated by vernalization and methylation. Plant Cell, 1999, 11: 445–458
Kim S Y, He Y, Jacob Y, et al. Establishment of the vernalization-responsive, winter-annual habit in Arabidopsis requires a putative histone H3 methyl transferase. Plant Cell, 2005, 17: 3301–3310
Zhao Z, Yu Y, Meyer D, et al. Prevention of early flowering by expression of FLOWERING LOCUS C requires methylation of histone H3 K36. Nat Cell Biol, 2005, 7: 1256–1260
Xu L, Zhao Z, Dong A, et al. Di- and tri-but not monomethylation on histone H3 lysine 36 marks active transcription of genes involved in flowering time regulation and other processes in Arabidopsis thaliana. Mol Cell Biol, 2008, 28: 1348–1360
Pien S, Fleury D, Mylne J S, et al. ARABIDOPSIS TRITHORAX1 dynamically regulates FLOWERING LOCUS C activation via histone 3 lysine 4 trimethylation. Plant Cell, 2008, 20: 580–588
Johanson U, West J, Lister C, et al. Molecular analysis of FRIGIDA, a major determinant of natural variation in Arabidopsis flowering time. Science, 2000, 290: 344–347
Jiang D, Gu X, He Y. Establishment of the winter-annual growth habit via FRIGIDA-mediated histone methylation at FLOWERING LOCUS C in Arabidopsis. Plant Cell, 2009, 21: 1733–1746
Ko J H, Mitina I, Tamada Y, et al. Growth habit determination by the balance of histone methylation activities in Arabidopsis. EMBO J, 2010, 29: 3208–3215
Choi K, Kim J, Hwang H J, et al. The FRIGIDA complex activates transcription of FLC, a strong flowering repressor in Arabidopsis, by recruiting chromatin modification factors. Plant Cell, 2011, 23: 289–303
Saleh A, Alvarez-Venegas R, Yilmaz M, et al. The highly similar Arabidopsis homologs of trithorax ATX1 and ATX2 encode proteins with divergent biochemical functions. Plant Cell, 2008, 20: 568–579
Berr A, Xu L, Gao J, et al. SET DOMAIN GROUP25 encodes a histone methyltransferase and is involved in FLOWERING LOCUS C activation and repression of flowering. Plant Physiol, 2009, 151: 1476–1485
Tamada Y, Yun J Y, Woo S C, et al. ARABIDOPSIS TRITHORAXRELATED7 is required for methylation of lysine 4 of histone H3 and for transcriptional activation of FLOWERING LOCUS C. Plant Cell, 2009, 21: 3257–3269
Schuettengruber B, Chourrout D, Vervoort M, et al. Genome regulation by polycomb and trithorax proteins. Cell, 2007, 128: 735–745
Kohler C, Villar C B. Programming of gene expression by Polycomb group proteins. Trends Cell Biol, 2008, 18: 236–243
Xu L, Shen W H. Polycomb silencing of KNOX genes confines shoot stem cell niches in Arabidopsis. Curr Biol, 2008, 18: 1966–1971
Bratzel F, Lopez-Torrejon G, Koch M, et al. Keeping cell identity in Arabidopsis requires PRC1 RING-finger homologs that catalyze H2A monoubiquitination. Curr Biol, 2010, 20: 1853–1859
Chen D, Molitor A, Liu C, et al. The Arabidopsis PRC1-like ringfinger proteins are necessary for repression of embryonic traits during vegetative growth. Cell Res, 2010, 20: 1332–1344
Sung S, Amasino R M. Remembering winter: Toward a molecular understanding of vernalization. Annu Rev Plant Biol, 2005, 56: 491–508
Bastow R, Mylne J S, Lister C, et al. Vernalization requires epigenetic silencing of FLC by histone methylation. Nature, 2004, 427: 164–167
Wood C C, Robertson M, Tanner G, et al. The Arabidopsis thaliana vernalization response requires a polycomb-like protein complex that also includes VERNALIZATION INSENSITIVE 3. Proc Natl Acad Sci USA, 2006, 103: 14631–14636
Finnegan E J, Dennis E S. Vernalization-induced trimethylation of histone H3 lysine 27 at FLC is not maintained in mitotically quiescent cells. Curr Biol, 2007, 17: 1978–1983
Mylne J S, Barrett L, Tessadori F, et al. LHP1, the Arabidopsis homologue of HETEROCHROMATIN PROTEIN1, is required for epigenetic silencing of FLC. Proc Natl Acad Sci USA, 2006, 103: 5012–5017
Jiang D, Wang Y, He Y. Repression of FLOWERING LOCUS C and FLOWERING LOCUS T by the Arabidopsis Polycomb repressive complex 2 components. PLoS ONE, 2008, 3: e3404
Alvarez-Venegas R, Avramova Z. Methylation patterns of histone H3 Lys 4, Lys 9 and Lys 27 in transcriptionally active and inactive Arabidopsis genes and in atx1 mutants. Nucleic Acids Res, 2005, 33: 5199–5207
Liu F, Quesada V, Crevillen P, et al. The Arabidopsis RNA-binding protein FCA requires a lysine-specific demethylase 1 homolog to downregulate FLC. Mol Cell, 2007, 28: 398–407
Jiang D, Yang W, He Y, et al. Arabidopsis relatives of the human lysine-specific Demethylase1 repress the expression of FWA and FLOWERING LOCUS C and thus promote the floral transition. Plant Cell, 2007, 19: 2975–2987
Noh B, Lee S H, Kim H J, et al. Divergent roles of a pair of homologous jumonji/zinc-finger-class transcription factor proteins in the regulation of Arabidopsis flowering time. Plant Cell, 2004, 16: 2601–2613
Jeong J H, Song H R, Ko J H, et al. Repression of FLOWERING LOCUS T chromatin by functionally redundant histone H3 lysine 4 demethylases in Arabidopsis. PLoS ONE, 2009, 4: e8033
Lu F, Cui X, Zhang S, et al. JMJ14 is an H3K4 demethylase regulating flowering time in Arabidopsis. Cell Res, 2010, 20: 387–390
Liu J, He Y, Amasino R, et al. siRNAs targeting an intronic transposon in the regulation of natural flowering behaviour in Arabidopsis thaliana. Genes Dev, 2004, 18: 2873–2878
Swiezewski S, Crevillen P, Liu F, et al. Small RNA-mediated chromatin silencing directed to the 3′ region of the Arabidopsis gene encoding the developmental regulator, FLC. Proc Natl Acad Sci USA, 2007, 1104: 3633–3638
Schmitz R J, Hong L, Fitzpatrick K E, et al. DICER-LIKE 1 and DICER-LIKE 3 redundantly act to promote flowering via repression of FLOWERING LOCUS C in Arabidopsis thaliana. Genetics, 2007, 176: 1359–1362
Heo J B, Sung S. Vernalization-mediated epigenetic silencing by a long intronic noncoding RNA. Science, 2011, 331: 76–79
Grini P E, Thorstensen T, Alm V, et al. The ASH1 HOMOLOG 2 (ASHH2) histone H3 methyltransferase is required for ovule and anther development in Arabidopsis. PLoS ONE, 2009, 4: e7817
Berr A, McCallum E J, Menard R, et al. Arabidopsis SET DOMAIN GROUP2 is required for H3K4 trimethylation and is crucial for both sporophyte and gametophyte development. Plant Cell, 2010, 22: 3232–3248
Guo L, Yu Y, Law J A, et al. SET DOMAIN GROUP2 is the major histone H3 lysine 4 trimethyltransferase in Arabidopsis. Proc Natl Acad Sci USA, 2010, 107: 18557–18562
Schiefthaler U, Balasubramanian S, Sieber P, et al. Molecular analysis of NOZZLE, a gene involved in pattern formation and early sporogenesis during sex organ development in Arabidopsis thaliana. Proc Natl Acad Sci USA, 1999, 96: 11664–11669
Yang W C, Ye D, Xu J, et al. The SPOROCYTELESS gene of Arabidopsis is required for initiation of sporogenesis and encodes a novel nuclear protein. Genes Dev, 1999, 13: 2108–2117
Ito T, Wellmer F, Yu H, et al. The homeotic protein AGAMOUS controls microsporogenesis by regulation of SPOROCYTELESS. Nature, 2004, 430: 356–360
Grossniklaus U, Vielle-Calzada J P, Hoeppner M A, et al. Maternal control of embryogenesis by MEDEA, a polycomb group gene in Arabidopsis. Science, 1998, 280: 446–450
Luo M, Bilodeau P, Koltunow A, et al. Genes controlling fertilization-independent seed development in Arabidopsis thaliana. Proc Natl Acad Sci USA, 1999, 96: 296–301
Kiyosue T, Ohad N, Yadegari R, et al. Control of fertilizationindependent endosperm development by the MEDEA polycomb gene in Arabidopsis. Proc Natl Acad Sci USA, 1999, 96: 4186–4191
Huh J H, Bauer M J, Hsieh T F, et al. Cellular programming of plant gene imprinting. Cell, 2008, 132: 735–744
Kohler C, Hennig L, Bouveret R, et al. Arabidopsis MSI1 is a component of the MEA/FIE Polycomb group complex and required for seed development. EMBO J, 2003, 22: 4804–4814
Wang D, Tyson M D, Jackson S S, et al. Partially redundant functions of two SET-domain polycomb-group proteins in controlling initiation of seed development in Arabidopsis. Proc Natl Acad Sci USA, 2006, 103: 13244–13249
Kohler C, Hennig L, Spillane C, et al. The Polycomb-group protein MEDEA regulates seed development by controlling expression of the MADS-box gene PHERES1. Genes Dev, 2003, 17: 1540–1553
Kohler C, Page D R, Gagliardini V, et al. The Arabidopsis thaliana MEDEA Polycomb group protein controls expression of PHERES1 by parental imprinting. Nat Genet, 2005, 37: 28–30
Pillot M, Baroux C, Vazquez M A, et al. Embryo and endosperm inherit distinct chromatin and transcriptional states from the female gametes in Arabidopsis. Plant Cell, 2010, 22: 307–322
Gehring M, Bubb K L, Henikoff S. Extensive demethylation of repetitive elements during seed development underlies gene imprinting. Science, 2009, 12: 1447–1451
Hsieh T F, Ibarra C A, Silva P, et al. Genome-wide demethylation of Arabidopsis endosperm. Science, 2009, 324: 1451–1454
Author information
Authors and Affiliations
Corresponding authors
Additional information
This article is published with open access at Springerlink.com
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Yao, X., Shen, W. Crucial function of histone lysine methylation in plant reproduction. Chin. Sci. Bull. 56, 3493–3499 (2011). https://doi.org/10.1007/s11434-011-4814-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11434-011-4814-3