Abstract
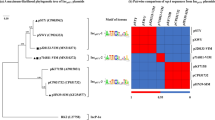
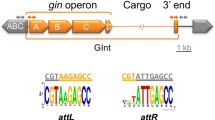
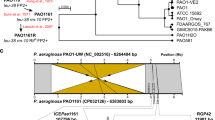
Mobile genomic islands (GIs) can be excised from the chromosome, then form a circular intermediate and be reintegrated into the chromosome by the GI internal integrase. Some mobile GIs can also be transferred into a new receptor cell by transformation, conjugation, or transduction. The action sites of the integrase are usually flanked direct repeats (DRs) of the GIs. Accurate localization of the flanking sequences is a precondition for determining the mobility of the GI. Mobile GIs are generally associated with transfer RNAs (tRNAs). Based on the correlation between flanking sequences and tRNA sequences, the flanking sequences of 11 putative mobile GIs in Pseudomonas aeruginosa PAO1, P. aeruginosa PA14, P. fluorescens Pf-5 and P. fluorescens Pf0-1 were identified. Among the 11 GIs, Pf0-1GI-1 is responsible for benzoate degradation. PAO1GI-1, Pf5GI-2, Pf5GI-3, and Pf5GI-4 were confirmed experimentally to be excised from a chromosome to form a circular intermediate. The action sites of the integrases are these GIs direct repeats. Due to distinct DRs, cutting sites for the internal integrase of PAO1GI-1, Pf5GI-2, Pf5GI-3 and Pf5GI-4 were determined outside the T-loop of the tRNAGly gene, outside the anticodon loop of the tRNASer gene and tRNALys gene, and at the asymmetric 3′-end of the tRNALeu gene, respectively. PAO1GI-1 and other mobile GIs may be transferred into many different strains that belong to different phyla because of the clear flanking sequences. This study describes basic information about the action sites of the integrases, assesses the mobility of GIs, and can help design and transfer mobile GIs to candidate strains.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
de la Cruz F, Davies J. Horizontal gene transfer and the origin of species: Lessons from bacteria. Trends Microbiol, 2000, 8: 128–133
Juhas M, van der Meer J R, Gaillard M, et al. Genomic islands: Tools of bacterial horizontal gene transfer and evolution. FEMS Microbiol Rev, 2009, 33: 376–393
Dobrindt U, Hochhut B, Hentschel U, et al. Genomic islands in pathogenic and environmental microorganisms. Nat Rev Microbiol, 2004, 2: 414–424
Glöckner G, Albert-Weissenberger C, Weinmann E, et al. Identification and characterization of a new conjugation/type IVA secretion system (trb/tra) of Legionella pneumophila Corby localized on two mobile genomic islands. Int J Med Microbiol, 2008, 298: 411–428
Hacker J, Blum-Oehler G, Mühldorfer I, et al. Pathogenicity islands of virulent bacteria: Structure, function and impact on microbial evolution. Mol Microbiol, 1997, 23: 1089–1097
Lyczak J B, Cannon C L, Pier G B. Establishment of Pseudomonas aeruginosa infection: Lessons from a versatile opportunist. Microbes Infect, 2000, 2: 1051–1060
Stover C K, Pham X Q, Erwin A L, et al. Complete genome sequence of Pseudomonas aeruginosa PAO1, an opportunistic pathogen. Nature, 2000, 406: 959–964
Lee D G, Urbach J M, Wu G, et al. Genomic analysis reveals that Pseudomonas aeruginosa virulence is combinatorial. Genome Biol, 2006, 7: R90
Paulsen I T, Press C M, Ravel J, et al. Complete genome sequence of the plant commensal Pseudomonas fluorescens Pf-5. Nat Biotechnol, 2005, 23: 873–878
Kim W, Levy S B. Increased fitness of Pseudomonas fluorescens Pf0-1 leucine auxotrophs in soil. Appl Environ Microbiol, 2008, 74: 3644–3651
Qiu X, Gurkar A U, Lory S. Interstrain transfer of the large pathogenicity island (PAPI-1) of Pseudomonas aeruginosa. Proc Natl Acad Sci USA, 2006, 103: 19830–19835
van Passel M W, Luyf A C, van Kampen A H, et al. δρ-Web, an online tool to assess composition similarity of individual nucleic acid sequences. Bioinformatics, 2005, 21: 3053–3055
Marchler-Bauer A, Anderson J B, Chitsaz F, et al. CDD: Specific functional annotation with the Conserved Domain Database. Nucleic Acids Res, 2009, 37: D205–D210
Webb J S, Lau M, Kjelleberg S. Bacteriophage and phenotypic variation in Pseudomonas aeruginosa biofilm development. J Bacteriol, 2004, 186: 8066–8073
Grindley N D, Whiteson K L, Rice P A. Mechanisms of site-specific recombination. Annu Rev Biochem, 2006, 75: 567–605
Haren L, Ton-Hoang B, Chandler M. Integrating DNA: Transposases and retroviral integrases. Annu Rev Microbiol, 1999, 53: 245–281
Williams K P. Integration sites for genetic elements in prokaryotic tRNA and tmRNA genes: Sublocation preference of integrase subfamilies. Nucleic Acids Res, 2002, 30: 866–875
Song L, Zhang X H. Innovation for ascertaining genomic islands in PAO1 and PA14 of Pseudomonas aeruginosa. Chinese Sci Bull, 2009, 54: 3991–3999
Mantri Y, Williams K P. Islander: A database of integrative islands in prokaryotic genomes, the associated integrases and their DNA site specificities. Nucleic Acids Res, 2004, 32: D55–D58
Langille M G, Brinkman F S. IslandViewer: An integrated interface for computational identification and visualization of genomic islands. Bioinformatics, 2009, 25: 664–665
Mathee K, Narasimhan G, Valdes C, et al. Dynamics of Pseudomonas aeruginosa genome evolution. Proc Natl Acad Sci USA, 2008, 105: 3100–3105
Mavrodi D V, Loper J E, Paulsen I T, et al. Mobile genetic elements in the genome of the beneficial rhizobacterium Pseudomonas fluorescens Pf-5. BMC Microbiol, 2009, 9: 8
Silby M W, Cerdeño-Tárraga A M, Vernikos G S, et al. Genomic and genetic analyses of diversity and plant interactions of Pseudomonas fluorescens. Genome Biol, 2009, 10: R51
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at Springerlink.com
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Song, L., Zhang, X. Accurate localization and excision of genomic islands in four strains of Pseudomonas aeruginosa and Pseudomonas fluorescens . Chin. Sci. Bull. 56, 987–995 (2011). https://doi.org/10.1007/s11434-011-4410-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11434-011-4410-6