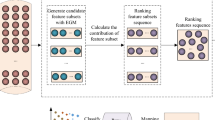
Abstract
The feature selection is an important challenge in many areas of machine learning because it plays a crucial role in the interpretations of machine-driven decisions. There are various approaches to the feature selection problem and methods based on the information theory comprise an important group. Here, the minimum redundancy maximum relevance (mRMR) feature selection is undoubtedly the most popular one with widespread application. In this paper, we prove in contrast to an existing finding that the mRMR is not equivalent to Max-Dependency criterion for first-order incremental feature selection. We present another form of equivalence leading to a generalization of mRMR feature selection. Additionally, we compare several feature selection methods based on mRMR, Max-Dependency, and feature ranking, employing different measures of dependency. The results on high-dimensional real-world datasets show that the distance correlation is the suitable measure for dependency-based feature selection methods. The results also indicate that the Max-Dependency incremental algorithm combined with distance correlation appears to be a promising feature selection approach.
Similar content being viewed by others
References
Li Y, Li T, Liu H. Recent advances in feature selection and its applications. Knowl Inf Syst, 2017, 53:551–577
Li J D, Liu H. Challenges of feature selection for big data analytics. IEEE Intell Syst, 2017, 32:9–15
Bolóon-Canedo V, Sáanchez-Maroñno N, Alonso-Betanzos A. Recent advances and emerging challenges of feature selection in the context of big data. Knowledge-Based Syst, 2015, 86:33–45
Ang J C, Mirzal A, Haron H, et al. Supervised, unsupervised, and semi-supervised feature selection:a review on gene selection. IEEE/ACM Trans Comput Biol Bioinf, 2016, 13:971–989
Li J D, Cheng K W, Wang S H, et al. Feature selection. ACM Comput Surv, 2018, 50:1–45
Battiti R. Using mutual information for selecting features in supervised neural net learning. IEEE Trans Neural Netw, 1994, 5:537–550
Kwak N, Choi C H. Input feature selection for classification problems. IEEE Trans Neural Netw, 2002, 13:143–159
Cai R C, Hao Z F, Yang X W, et al. An efficient gene selection algorithm based on mutual information. Neurocomputing, 2009, 72:991–999
Fleuret F. Fast binary feature selection with conditional mutual information. J Mach Learn Res, 2004, 5:1531–1555
Cheng H R, Qin Z G, Feng C S, et al. Conditional mutual information-based feature selection analyzing for synergy and redundancy. ETRI J, 2011, 33:210–218
Yang H H, Moody J. Data visualization and feature selection:new algorithms for nongaussian data. In: Proceedings of the 12th International Conference on Neural Information Processing Systems. Cambridge: MIT Press, 1999. 687–693
Vergara J R, Estéevez P A. A review of feature selection methods based on mutual information. Neural Comput Appl, 2014, 24:175–186
Brown G, Pocock A, M-Zhao J, et al. Conditional likelihood maximisation:a unifying framework for information theoretic feature selection. Mach J Learn Res, 2012, 13, 27–66
Peng H C, Long F H, Ding C. Feature selection based on mutual information:criteria of max-dependency, maxrelevance, and min-redundancy. IEEE Trans Pattern Anal Machine Intell, 2005, 27:1226–1238
Ding C, Peng H. Minimum redundancy feature selection from microarray gene expression data. J Bioinform Comput Biol, 2005, 03:185–205
Corredor G, Wang X, Zhou Y, et al. Spatial architecture and arrangement of tumor-infiltrating lymphocytes for predicting likelihood of recurrence in early-stage non-small cell lung cancer. Clin Cancer Res, 2019, 25:1526–1534
Toyoda A, Ogawa T, Haseyama M. Favorite video estimation based on multiview feature integration via KMvLFDA. IEEE Access, 2018, 6:63833–63842
Berrendero J R, Cuevas A, Torrecilla J L. The mRMR variable selection method:a comparative study for functional data. J Stat Comput Simul, 2016, 86:891–907
Guyon I, Elisseeff A. An introduction to variable and feature selection. Mach J Learn Res, 2003, 3:1157–1182
Golub T R, Slonim D K, Tamayo P, et al. Molecular classification of cancer:class discovery and class prediction by gene expression monitoring. Science, 1999, 286:531–537
Gordon G J G, Jensen R V R, Hsiao L-L L, et al. Translation of microarray data into clinically relevant cancer diagnostic tests using gene expression ratios in lung cancer and mesothelioma. Cancer Res, 2002, 62:4963–4967
Alon U, Barkai N, Notterman D A, et al. Broad patterns of gene expression revealed by clustering analysis of tumor and normal colon tissues probed by oligonucleotide arrays. Proc Natl Acad Sci USA, 1999, 96:6745–6750
Tian E, Zhan F, Walker R, et al. The role of the Wnt-signaling antagonist DKK1 in the development of osteolytic lesions in multiple myeloma. New Engl J Med, 2003, 349:2483–2494
Burczynski M E, Peterson R L, Twine N C, et al. Molecular classification of crohn’s disease and ulcerative colitis patients using transcriptional profiles in peripheral blood mononuclear cells. J Mol Diagn, 2006, 8:51–61
Pomeroy S L, Tamayo P, Gaasenbeek M, et al. Prediction of central nervous system embryonal tumour outcome based on gene expression. Nature, 2002, 415:436–442
Singh Y N, Singh S K, Ray A K. Bioelectrical signals as emerging biometrics:issues and challenges. ISRN Signal Process, 2012, 2012:136–151
Chowdary D, Lathrop J, Skelton J, et al. Prognostic gene expression signatures can be measured in tissues collected in RNAlater preservative. J Mol Diagn, 2006, 8:31–39
Széekely G J, Rizzo M L, Bakirov N K. Measuring and testing dependence by correlation of distances. Ann Statist, 2007, 35:2769–2794
Robnik-Šikonja M, Kononenko I. Theoretical and empirical analysis of ReliefF and RReliefF. Mach Learn, 2003, 53:23–69
Acknowledgements
This work was supported by Slovak Research and Development Agency (Grant No. APVV-16-0211).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bugata, P., Drotar, P. On some aspects of minimum redundancy maximum relevance feature selection. Sci. China Inf. Sci. 63, 112103 (2020). https://doi.org/10.1007/s11432-019-2633-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11432-019-2633-y