Abstract
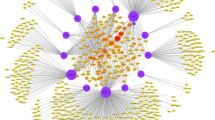
With the emergence of multi-dimensional data, we can more comprehensively analyze the pathogenic mechanisms of complex diseases, and thereby improve the diagnosis, treatment and prevention of these diseases. This study presents a novel multi-layer network-based strategy that integrates multi-dimensional data, and identifies disease-related modules and components of viral infections. We first propose a systematic method that constructs a virus-host interaction network with three layers: a viral protein layer, a host protein layer and a host gene layer. This method combines the data of high-throughput gene expression, viral protein interactions, virus-host interactions, protein-protein interactions and transcriptional regulatory relationships. To extract the underlying mechanisms of viral infections from the multi-layer networks, we identify the intra-layer and crosslayer modules, and investigate the conserved modules across multiple datasets. The essential components in the multi-layer networks are detected by singular-value decomposition. The identified conserved modules and essential components are combined into a functional enrichment analysis that reveals their contributions to influenza virus replication. By this analysis, we elucidate the common and specific mechanisms of the replication cycles of two influenza strains. By combining the different layers of information, we can comprehensively understand pathogenic mechanisms of complex diseases.
Similar content being viewed by others
References
Hsu N Y, Ilnytska O, Belov G, et al. Viral reorganization of the secretory pathway generates distinct organelles for RNA replication. Cell, 2010, 141: 799–811
Jin S Q, Zou X F. Construction of the influenza A virus infection-induced cell-specific inflammatory regulatory network based on mutual information and optimization. BMC Syst Biol, 2013, 7: 105
Jin S Q, Li Y Y, Pan R G, et al. Characterizing and controlling the inflammatory network during influenza A virus infection. Sci Rep, 2014, 4: 3799
Li Y Y, Jin S Q, Lei L, et al. Deciphering deterioration mechanisms of complex diseases based on the construction of dynamic networks and systems analysis. Sci Rep, 2015, 5: 9283
Barabasi A L, Gulbahce N, Loscalzo J. Network medicine: a network-based approach to human disease. Nat Rev Genet, 2011, 12: 56–68
Cantini L, Medico E, Fortunato S, et al. Detection of gene communities in multi-networks reveals cancer drivers. Sci Rep, 2015, 5: 17386
Li W, Dai C, Liu C C, et al. Algorithm to identify frequent coupled modules from two-layered network series: application to study transcription and splicing coupling. J Comput Biol, 2012, 19: 710–730
Shapira S D, Gat-Viks I, Shum B O, et al. A physical and regulatory map of host-influenza interactions reveals pathways in H1N1 infection. Cell, 2009, 139: 1255–1267
Watanabe T, Watanabe S, Kawaoka Y. Cellular networks involved in the influenza virus life cycle. Cell Host Microbe, 2010, 7: 427–439
Wang Y C, Chen B S. Integrated cellular network of transcription regulations and protein-protein interactions. BMC Syst Biol, 2010, 4: 20
Srivastava A, Kumar S, Ramaswamy R. Two-layer modular analysis of gene and protein networks in breast cancer. BMC Syst Biol, 2014, 8: 81
Barrett T, Suzek T O, Troup D B, et al. NCBI GEO: mining millions of expression profiles—database and tools. Nucl Acids Res, 2005, 33: D562–566
Hsu A C Y, Barr I, Hansbro P M, et al. Human influenza is more effective than avian influenza at antiviral suppression in airway cells. Amer J Resp Cell Mol Biol, 2011, 44: 906–913
Josset L, Zeng H, Kelly S M, et al. Transcriptomic characterization of the novel avian-origin influenza A (H7N9) virus: specific host response and responses intermediate between avian (H5N1 and H7N7) and human (H3N2) viruses and implications for treatment options. mBio, 2014, 5: e01102–01113
Ozawa M, Kawaoka Y. Taming influenza viruses. Virus Res, 2011, 162: 8–11
Chen W, Calvo P A, Malide D, et al. A novel influenza A virus mitochondrial protein that induces cell death. Nat Med, 2001, 7: 1306–1312
Wise H M, Foeglein A, Sun J, et al. A complicated message: identification of a novel PB1-related protein translated from influenza A virus segment 2 mRNA. J Virol, 2009, 83: 8021–8031
Lamesch P, Li N, Milstein S, et al. hORFeome v3.1: a resource of human open reading frames representing over 10,000 human genes. Genomics, 2007, 89: 307–315
Schaefer M H, Fontaine J F, Vinayagam A, et al. HIPPIE: integrating protein interaction networks with experiment based quality scores. PLoS ONE, 2012, 7: e31826
Zheng G, Qian Z, Yang Q, et al. The combination approach of SVM and ECOC for powerful identification and classification of transcription factor. BMC Bioinform, 2008, 9: 282
Zheng G, Tu K, Yang Q, et al. ITFP: an integrated platform of mammalian transcription factors. Bioinformatics, 2008, 24: 2416–2417
Sun N, Carroll R J, Zhao H. Bayesian error analysis model for reconstructing transcriptional regulatory networks. Proc Nat Acad Sci USA, 2006, 103: 7988–7993
Wang R S, Jin G, Zhang X S, et al. Modeling post-transcriptional regulation activity of small non-coding RNAs in Escherichia coli. BMC Bioinform, 2009, 10: S6
Geeven G, van Kesteren R E, Smit A B, et al. Identification of context-specific gene regulatory networks with GEMULA-gene expression modeling using LAsso. Bioinformatics, 2012, 28: 214–221
Saito S, Hirokawa T, Horimoto K. Discovery of chemical compound groups with common structures by a network analysis approach (affinity prediction method). J Chem Inf Model, 2011, 51: 61–68
Brunel H, Gallardo-Chacon J J, Buil A, et al. MISS: a non-linear methodology based on mutual information for genetic association studies in both population and sib-pairs analysis. Bioinformatics, 2010, 26: 1811–1818
Zhang X, Zhao X M, He K, et al. Inferring gene regulatory networks from gene expression data by path consistency algorithm based on conditional mutual information. Bioinformatics, 2012, 28: 98–104
Nepusz T, Yu H, Paccanaro A. Detecting overlapping protein complexes in protein-protein interaction networks. Nat Methods, 2012, 9: 471–472
Xiao X Y, Zhang W, Zou X F. A new asynchronous parallel algorithm for inferring large-scale gene regulatory networks. PLoS ONE, 2015, 10: e0119294
Zhang W, Zou X F. A new method for detecting protein complexes based on the three node cliques. IEEE/ACM Trans Comput Biol Bioinform, 2015, 12: 879–886
Huang D W, Sherman B T, Lempicki R A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc, 2009, 4: 44–57
Cao M M, Wei C D, Zhao L L, et al. DnaJA1/Hsp40 is co-opted by influenza A virus to enhance its viral RNA polymerase activity. J Virol, 2014, 88: 14078–14089
Momose F, Naito T, Yano K, et al. Identification of Hsp90 as a stimulatory host factor involved in influenza virus RNA synthesis. J Biol Chem, 2002, 277: 45306–45314
Menche J, Sharma A, Kitsak M, et al. Uncovering disease-disease relationships through the incomplete interactome. Science, 2015, 347: 1257601
de Domenico M, Sole-Ribalta A, Omodei E, et al. Ranking in interconnected multilayer networks reveals versatile nodes. Nat Commun, 2015, 6: 6868
Tan J Y, Zou X F. Complex dynamical analysis of a coupled network from innate immune responses. Int J Bifurcat Chaos, 2013, 23: 1350180
Tan J Y, Zou X F. Optimal control strategy for abnormal innate immune response. Comput Math Methods Med, 2015, 2015: 386235
Wang D J, Jin S Q, Wu F X, et al. Estimation of control energy and control strategies for complex networks. Adv Complex Syst, 2015, 18: 1550018
Zou X F, Niu L L, Jin S Q. The mathematical modeling and analysis for S1PR1-mediated cytokine signaling pathway. J Jiangxi Norm Univ (Nat Sci Ed), 2015, 39: 7–14
Jin S Q, Niu L L, Wang G, et al. Mathematical modeling and nonlinear dynamical analysis of cell growth in response to antibiotics. Int J Bifurcat Chaos, 2015, 25: 1540007
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, Y., Zou, X. Identifying disease modules and components of viral infections based on multi-layer networks. Sci. China Inf. Sci. 59, 070102 (2016). https://doi.org/10.1007/s11432-016-5580-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11432-016-5580-2