Abstract
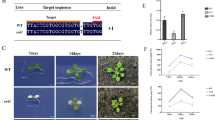
Male sterility is an essential trait in hybrid seed production, especially for monoclinous and autogamous food crops. Soybean male-sterile ms1 mutant has been known for more than 50 years and could be instrumental in making hybrid seeds. However, the gene responsible for the male-sterile phenotype has remained unknown. Here, we report the map-based cloning and characterization of the MS1 gene in soybean. MS1 encodes a kinesin protein and localizes to the nucleus, where it is required for the male meiotic cytokinesis after telophase II. We further substantiated that MS1 colocalizes with microtubules and is essential for cell plate formation in soybean male gametogenesis through immunostaining. Both ms1 and CRISPR/Cas9 knockout mutants show complete male sterility but are otherwise phenotypically normal, making them perfect tools for producing hybrid seeds. The identification of MS1 has the practical potential for assembling the sterility system and speeding up hybrid soybean breeding.
Similar content being viewed by others
Data availability
Data supporting the findings of this work are available within the paper and its Supplementary Information files. The RNA-seq data supporting this research have been deposited in the NCBI SRA with the accession number PRJNA726823.
References
Albertsen, M.C., and Palmer, R.G. (1979). A comparative light- and electron-microscopic study of microsporogenesis in male sterile (MS1) and male fertile soybeans (Glycine max (L.) Merr.). Am J Bot 66, 253–265.
Brim, C.A., and Young, M.F. (1971). Inheritance of a male-sterile character in soybeans. Crop Sci 11, 564–566.
Chang, F., Wang, Y., Wang, S., and Ma, H. (2011). Molecular control of microsporogenesis in Arabidopsis. Curr Opin Plant Biol 14, 66–73.
Chen, L., and Liu, Y.G. (2014). Male sterility and fertility restoration in crops. Annu Rev Plant Biol 65, 579–606.
Chen, S., Zhou, Y., Chen, Y., and Gu, J. (2018). fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890.
Davis, W.H. (1985). Route to hybrid soybean production. US Patent, 4545146.
Echlin, P., and Godwin, H. (1968). The ultrastructure and ontogeny of pollen in Helleborus Foetidus L. J Cell Sci 3, 175–186.
Fang, X., Zhao, Y., Ma, Q., Huang, Y., Wang, P., Zhang, J., Nian, H., and Yang, C. (2013). Identification and comparative analysis of cadmium tolerance-associated miRNAs and their targets in two soybean genotypes. PLoS ONE 8, e81471.
Fang, X., Han, Y., Liu, M., Jiang, J., Li, X., Lian, Q., Xie, X., Huang, Y., Ma, Q., Nian, H., et al. (2021). Modulation of evening complex activity enables north-to-south adaptation of soybean. Sci China Life Sci 64, 179–195.
Hackenberg, D., and Twell, D. (2019). The evolution and patterning of male gametophyte development. In: Grossniklaus, U., ed. Current Topics in Developmental Biology. New York: Academic Press. 257–298.
Ho, C.M.K., Hotta, T., Guo, F., Roberson, R.W., Lee, Y.R.J., and Liu, B. (2011). Interaction of antiparallel microtubules in the phragmoplast is mediated by the microtubule-associated protein MAP65–3 in Arabidopsis. Plant Cell 23, 2909–2923.
Huang, Y., Wang, H., Huang, X., Wang, Q., Wang, J., An, D., Li, J., Wang, W., and Wu, Y. (2019). Maize VKS1 regulates mitosis and cytokinesis during early endosperm development. Plant Cell 31, 1238–1256.
Jiang, B., Chen, L., Yang, C., Wu, T., Yuan, S., Wu, C., Zhang, M., Gai, J., Han, T., Hou, W., et al. (2021). The cloning and CRISPR/Cas9-mediated mutagenesis of a male sterility gene MS1 of soybean. Plant Biotechnol J 19, 1098–1100.
Kim, D., Langmead, B., and Salzberg, S.L. (2015). HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12, 357–360.
Kim, Y.J., and Zhang, D. (2018). Molecular control of male fertility for crop hybrid breeding. Trends Plant Sci 23, 53–65.
Lawrence, C.J., Dawe, R.K., Christie, K.R., Cleveland, D.W., Dawson, S. C., Endow, S.A., Goldstein, L.S.B., Goodson, H.V., Hirokawa, N., Howard, J., et al. (2004). A standardized kinesin nomenclature. J Cell Biol 167, 19–22.
Lewers, K.S., and Palmer, R.G. (1997). Recurrent selection in soybean. Plant Breed Rev 15, 275–313.
Li, H., and Durbin, R. (2009). Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760.
Li, H., Handsaker, B., Wysoker, A., Fennell, T., Ruan, J., Homer, N., Marth, G., Abecasis, G., and Durbin, R. (2009). The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079.
Liu, S., Zhang, M., Feng, F., and Tian, Z. (2020). Toward a “Green Revolution” for soybean. Mol Plant 13, 688–697.
Liu, W., Wang, C., Wang, G., Ma, Y., Tian, J., Yu, Y., Dong, L., and Kong, Z. (2019). Towards a better recording of microtubule cytoskeletal spatial organization and dynamics in plant cells. J Integr Plant Biol 61, 388–393.
Love, M.I., Huber, W., and Anders, S. (2014). Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15, 550.
Lu, P., Chai, M., Yang, J., Ning, G., Wang, G., and Ma, H. (2014). The Arabidopsis CALLOSE DEFECTIVE MICROSPORE1 gene is required for male fertility through regulating callose metabolism during microsporogenesis. Plant Physiol 164, 1893–1904.
Ma, H. (2005). Molecular genetic analyses of microsporogenesis and microgametogenesis in flowering plants. Annu Rev Plant Biol 56, 393–434.
Ma, X., Zhang, Q., Zhu, Q., Liu, W., Chen, Y., Qiu, R., Wang, B., Yang, Z., Li, H., Lin, Y., et al. (2015). A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol Plant 8, 1274–1284.
McKenna, A., Hanna, M., Banks, E., Sivachenko, A., Cibulskis, K., Kernytsky, A., Garimella, K., Altshuler, D., Gabriel, S., Daly, M., et al. (2010). The genome analysis toolkit: a mapreduce framework for analyzing next-generation DNA sequencing data. Genome Res 20, 1297–1303.
Nadeem, M., Chen, A., Hong, H., Li, D., Li, J., Zhao, D., Wang, W., Wang, X., and Qiu, L. (2021). GmMs1 encodes a kinesin-like protein essential for male fertility in soybean (Glycine max L.). J Integr Plant Biol 63, 1054–1064.
Naito, Y., Hino, K., Bono, H., and Ui-Tei, K. (2015). CRISPRdirect: software for designing CRISPR/Cas guide RNA with reduced off-target sites. Bioinformatics 31, 1120–1123.
Nebenführ, A., and Dixit, R. (2018). Kinesins and myosins: molecular motors that coordinate cellular functions in plants. Annu Rev Plant Biol 69, 329–361.
Nie, Z., Zhao, T., Liu, M., Dai, J., He, T., Lyu, D., Zhao, J., Yang, S., and Gai, J. (2019). Molecular mapping of a novel male-sterile gene msNJ in soybean [Glycine max (L.) Merr.]. Plant Reprod 32, 371–380.
Nishihama, R., Soyano, T., Ishikawa, M., Araki, S., Tanaka, H., Asada, T., Irie, K., Ito, M., Terada, M., Banno, H., et al. (2002). Expansion of the cell plate in plant cytokinesis requires a kinesin-like protein/MAPKKK complex. Cell 109, 87–99.
Palmer, R.G., Winger, C.L., and Albertsen, M.C. (1978). Four independent mutations at the ms1, locus in soybeans. Crop Sci 18, 727–729.
Pertea, M., Pertea, G.M., Antonescu, C.M., Chang, T.C., Mendell, J.T., and Salzberg, S.L. (2015). StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotechnol 33, 290–295.
Pertea, M., Kim, D., Pertea, G.M., Leek, J.T., and Salzberg, S.L. (2016). Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat Protoc 11, 1650–1667.
Rutley, N., and Twell, D. (2015). A decade ofpollen transcriptomics. Plant Reprod 28, 73–89.
Sasabe, M., and Machida, Y. (2012). Regulation of organization and function of microtubules by the mitogen-activated protein kinase cascade during plant cytokinesis. Cytoskeleton 69, 913–918.
Sasabe, M., Soyano, T., Takahashi, Y., Sonobe, S., Igarashi, H., Itoh, T.J., Hidaka, M., and Machida, Y. (2006). Phosphorylation of NtMAP65–1 by a MAP kinase down-regulates its activity of microtubule bundling and stimulates progression of cytokinesis of tobacco cells. Genes Dev 20, 1004–1014.
Spielman, M., Preuss, D., Li, F.L., Browne, W.E., Scott, R.J., and Dickinson, H.G. (1997). TETRASPORE is required for male meiotic cytokinesis in Arabidopsis thaliana. Development 124, 2645–2657.
Sun, H., Zhao, L., and Huang, M. (1993). A study on the cytoplasm-nuclear male sterile soybean line (in Chinese). Chin Sci Bull 38, 1535–1536.
Tanaka, H., Ishikawa, M., Kitamura, S., Takahashi, Y., Soyano, T., Machida, C., and Machida, Y. (2004). The AtNACK1/HINKEL and STUD/TETRASPORE/AtNACK2 genes, which encode functionally redundant kinesins, are essential for cytokinesis in Arabidopsis. Genes Cells 9, 1199–1211.
Tester, M., and Langridge, P. (2010). Breeding technologies to increase crop production in a changing world. Science 327, 818–822.
Thu, S.W., Rai, K.M., Sandhu, D., Rajangam, A., Balasubramanian, V.K., Palmer, R.G., and Mendu, V. (2019). Mutation in a PHD-finger protein MS4 causes male sterility in soybean. BMC Plant Biol 19, 378.
Wan, X., Wu, S., Li, Z., Dong, Z., An, X., Ma, B., Tian, Y., and Li, J. (2019). Maize genic male-sterility genes and their applications in hybrid breeding: progress and perspectives. Mol Plant 12, 321–342.
Wang, K., Li, M., and Hakonarson, H. (2010). ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res 38, e164.
Whitford, R., Fleury, D., Reif, J.C., Garcia, M., Okada, T., Korzun, V., and Langridge, P. (2013). Hybrid breeding in wheat: technologies to improve hybrid wheat seed production. J Exp Bot 64, 5411–5428.
Wilson, Z.A., and Zhang, D.B. (2009). From Arabidopsis to rice: pathways in pollen development. J Exp Bot 60, 1479–1492.
Wu, G., Fanzo, J., Miller, D.D., Pingali, P., Post, M., Steiner, J.L., and Thalacker-Mercer, A.E. (2014). Production and supply of high-quality food protein for human consumption: sustainability, challenges, and innovations. Ann NY Acad Sci 1321, 1–19.
Wu, Y., Fox, T.W., Trimnell, M.R., Wang, L., Xu, R.J., Cigan, A.M., Huffman, G.A., Garnaat, C.W., Hershey, H., and Albertsen, M.C. (2016). Development of a novel recessive genetic male sterility system for hybrid seed production in maize and other cross-pollinating crops. Plant Biotechnol J 14, 1046–1054.
Xu, X., Walter, W.J., Liu, Q., Machens, I., and Nick, P. (2018). A rice class-XIV kinesin enters the nucleus in response to cold. Sci Rep 8, 3588.
Yang, C.Y., Spielman, M., Coles, J.P., Li, Y., Ghelani, S., Bourdon, V., Brown, R.C., Lemmon, B.E., Scott, R.J., and Dickinson, H.G. (2003a). TETRASPORE encodes a kinesin required for male meiotic cytokinesis in Arabidopsis. Plant J 34, 229–240.
Yang, X., Makaroff, C.A., and Ma, H. (2003b). The ArabidopsisMALE MEIOCYTE DEATH1 gene encodes a PHD-finger protein that is required for male meiosis. Plant Cell 15, 1281–1295.
Yang, Y., Speth, B.D., Boonyoo, N., Baumert, E., Atkinson, T.R., Palmer, R.G., and Sandhu, D. (2014). Molecular mapping of three male-sterile, female-fertile mutants and generation of a comprehensive map of all known male sterility genes in soybean. Genome 57, 155–160.
Zhang, D., Luo, X., and Zhu, L. (2011). Cytological analysis and genetic control of rice anther development. J Genet Genomics 38, 379–390.
Zhao, L., Sun, H., Wang, S., Wang, Y., Huang, M., and Li, J. (2004). Breeding of hybrid soybean HybSoy1 (in Chinese). Chin J Oil Crop Sci 26, 15–17.
Zhu, C., and Dixit, R. (2012). Functions of the Arabidopsis kinesin superfamily of microtubule-based motor proteins. Protoplasma 249, 887–899.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (32072084, 31871648, and 31971969). We gratefully acknowledge Zhaosheng Kong from the Institute of Microbiology, Chinese Academy of Sciences, for sharing the pTUB6::mCherry-TUB6 transgenic Arabidopsis seeds and useful discussions. We would like to thank Dr. Ali Muhammad for editing and improving the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Compliance and ethics The author(s) declare that they have no conflict of interest.
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Fang, X., Sun, X., Yang, X. et al. MS1 is essential for male fertility by regulating the microsporocyte cell plate expansion in soybean. Sci. China Life Sci. 64, 1533–1545 (2021). https://doi.org/10.1007/s11427-021-1973-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-021-1973-0