Abstract
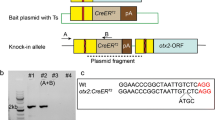
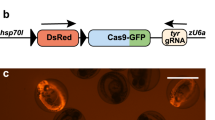
The zebrafish has become a popular vertebrate animal model in biomedical research. However, it is still challenging to make conditional gene knockout (CKO) models in zebrafish due to the low efficiency of homologous recombination (HR). Here we report an efficient non-HR-based method for generating zebrafish carrying a CKO and knockin (KI) switch (zCKOIS) coupled with dual-color fluorescent reporters. Using this strategy, we generated hey2zKOIS which served as a hey2 KI reporter with EGFP expression. Upon Cre induction in targeted cells, the hey2zCKOIS was switched to a non-functional CKO allele hey2zCKOIS-invassociated with TagRFP expression, enabling visualization of the CKO alleles. Thus, simplification of the design, and the visibility and combination of both CKO and KI alleles make our zCKOIS strategy an applicable CKO approach for zebrafish.
Similar content being viewed by others
References
Bertrand, J.Y., Chi, N.C., Santoso, B., Teng, S., Stainier, D.Y.R., and Traver, D. (2010). Haematopoietic stem cells derive directly from aortic endothelium during development. Nature 464, 108–111.
Chang, N., Sun, C., Gao, L., Zhu, D., Xu, X., Zhu, X., Xiong, J.W., and Xi, J.J. (2013). Genome editing with RNA-guided Cas9 nuclease in zebrafish embryos. Cell Res 23, 465–472.
Dai, J., Cui, X., Zhu, Z., and Hu, W. (2010). Non-homologous end joining plays a key role in transgene concatemer formation in transgenic zebrafish embryos. Int J Biol Sci 6, 756–768.
Gibb, N., Lazic, S., Yuan, X., Deshwar, A.R., Leslie, M., Wilson, M.D., and Scott, I.C. (2018). Hey2 regulates the size of the cardiac progenitor pool during vertebrate heart development. Development 145, dev167510.
Hoshijima, K., Jurynec, M.J., and Grunwald, D.J. (2016). Precise editing of the zebrafish genome made simple and efficient. Dev Cell 36, 654–667.
Jia, H., King, I.N., Chopra, S.S., Wan, H., Ni, T.T., Jiang, C., Guan, X., Wells, S., Srivastava, D., and Zhong, T.P. (2007). Vertebrate heart growth is regulated by functional antagonism between Gridlock and Gata5. Proc Natl Acad Sci USA 104, 14008–14013.
Korten, S., Brunssen, C., Poitz, D.M., Großklaus, S., Brux, M., Schnittler, H.J., Strasser, R.H., Bornstein, S.R., Morawietz, H., and Goettsch, W. (2013). Impact of Hey2 and COUP-TFII on genes involved in arteriovenous differentiation in primary human arterial and venous endothelial cells. Basic Res Cardiol 108, 362.
Li, J., Zhang, B., Ren, Y., Gu, S., Xiang, Y., Huang, C., and Du, J. (2015). Intron targeting-mediated and endogenous gene integrity-maintaining knockin in zebrafish using the CRISPR/Cas9 system. Cell Res 25, 634–637.
Li, W., Zhang, Y., Han, B., Li, L., Li, M., Lu, X., Chen, C., Lu, M., Zhang, Y., Jia, X., et al. (2019). One-step efficient generation of dual-function conditional knockout and geno-tagging alleles in zebrafish. eLife 8, e48081.
Liu, D., Wang, Z., Xiao, A., Zhang, Y., Li, W., Zu, Y., Yao, S., Lin, S., and Zhang, B. (2014). Efficient gene targeting in zebrafish mediated by a zebrafish-codon-optimized cas9 and evaluation of off-targeting effect. J Genet Genomics 41, 43–46.
Lovett-Barron, M., Chen, R., Bradbury, S., Andalman, A.S., Wagle, M., Guo, S., and Deisseroth, K. (2019). Multiple overlapping hypothalamus-brainstem circuits drive rapid threat avoidance. bioRxiv, doi: https://doi.org/10.1101/745075.
Mu, Y., Bennett, D.V., Rubinov, M., Narayan, S., Yang, C.T., Tanimoto, M., Mensh, B.D., Looger, L.L., and Ahrens, M.B. (2019). Glia accumulate evidence that actions are futile and suppress unsuccessful behavior. Cell 178, 27–43.e19.
Peterson, R.T., Shaw, S.Y., Peterson, T.A., Milan, D.J., Zhong, T.P., Schreiber, S.L., MacRae, C.A., and Fishman, M.C. (2004). Chemical suppression of a genetic mutation in a zebrafish model of aortic coarctation. Nat Biotechnol 22, 595–599.
Robles, V., Marti, M., and Izpisua Belmonte, J.C. (2011). Study of pluripotency markers in zebrafish embryos and transient embryonic stem cell cultures. Zebrafish 8, 57–63.
Rowlinson, J.M., and Gering, M. (2010). Hey2 acts upstream of Notch in hematopoietic stem cell specification in zebrafish embryos. Blood 116, 2046–2056.
Satow, T., Bae, S.K., Inoue, T., Inoue, C., Miyoshi, G., Tomita, K., Bessho, Y., Hashimoto, N., and Kageyama, R. (2001). The basic helix-loop-helix gene hesr2 promotes gliogenesis in mouse retina. J Neurosci 21, 1265–1273.
Sugimoto, K., Hui, S.P., Sheng, D.Z., and Kikuchi, K. (2017). Dissection of zebrafish shha function using site-specific targeting with a Cre-dependent genetic switch. eLife 6, e24635.
Sun, Y., and Zhu, Z. (2019). Designing future farmed fishes using genome editing. Sci China Life Sci 62, 420–422.
Suzuki, K., Yamamoto, M., Hernandez-Benitez, R., Li, Z., Wei, C., Soligalla, R.D., Aizawa, E., Hatanaka, F., Kurita, M., Reddy, P., et al. (2019). Precise in vivo genome editing via single homology arm donor mediated intron-targeting gene integration for genetic disease correction. Cell Res 29, 804–819.
Wang, Y., Wang, F., Wang, R., Zhao, P., and Xia, Q. (2015). 2A self-cleaving peptide-based multi-gene expression system in the silkworm Bombyx mori. Sci Rep 5, 16273.
Yu, Y., and Bradley, A. (2001). Engineering chromosomal rearrangements in mice. Nat Rev Genet 2, 780–790.
Zhong, T.P., Childs, S., Leu, J.P., and Fishman, M.C. (2001). Gridlock signalling pathway fashions the first embryonic artery. Nature 414, 216–220.
Zhong, T.P., Rosenberg, M., Mohideen, M.A., Weinstein, B., and Fishman, M.C. (2000). gridlock, an HLH gene required for assembly of the aorta in zebrafish. Science 287, 1820–1824.
Zu, Y., Tong, X., Wang, Z., Liu, D., Pan, R., Li, Z., Hu, Y., Luo, Z., Huang, P., Wu, Q., et al. (2013). TALEN-mediated precise genome modification by homologous recombination in zebrafish. Nat Methods 10, 329–331.
Acknowledgements
We are grateful to Drs. N. Lawson for providing the Tg(flk1:EGFP) line, D. Traver for providing the Tg(bactin2:loxP-STOP-loxP-DsRedEx) line and K. Kikuchi for providing the pZwitch+1 plasmid. This work was supported by the Young Scientists Fund of the National Natural Science Foundation of China (31500849), Shanghai Municipal Science and Technology Major Project (18JC1410100, 2018SHZDZX05), the Key Research Program of Frontier Sciences (QYZDY-SSW-SMC028), the Strategic Priority Research Program (XDB32010200) of Chinese Academy of Sciences, the International Partnership Program, Bureau of International Co-operation of Chinese Academy of Sciences (153D31KYSB20170059), China Wan-Ren Program, and Shanghai Leading Scientist Program.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Compliance and ethics The author(s) declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Li, J., Li, HY., Gu, SY. et al. One-step generation of zebrafish carrying a conditional knockout-knockin visible switch via CRISPR/Cas9-mediated intron targeting. Sci. China Life Sci. 63, 59–67 (2020). https://doi.org/10.1007/s11427-019-1607-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-019-1607-9