Abstract
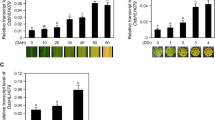
The class III homeodomain-leucine zipper (HD-Zip III) gene family plays important roles in plant growth and development, including regulation of apical embryo patterning, embryonic shoot meristem formation, leaf polarity, vascular development, and meristem function, with a particularly crucial function in leaf development. Although HD-Zip III members are highly conserved in land plants, previous studies, such as genetic analyses based on multiple mutants in Arabidopsis and other plants, suggest that various HD-Zip III family genes have evolved with distinct functions and pleiotropic effects on plant growth and development. In this study, we analyzed a HD-Zip III member, OsHox33, and demonstrated that it plays an important role in age-dependent leaf senescence in rice. We constructed two specific RNAi vectors derived from the 5′-end region and 3′-UTR of OsHox33 to knockdown its expression. Transgenic plants harboring either RNAi construct displayed similar phenotypes of precocious leaf senescence symptoms, suggesting that knockdown of OsHox33 accelerates leaf senescence in rice. pOsHox33::GUS fusion expression and RT-PCR revealed that OsHox33 is highly expressed in young organs, especially in young meristems such as shoot apical meristems, intercalary meristems, and young callus. In addition, real-time PCR indicated that OsHox33 was more highly expressed in young leaves than in old leaves. To further investigate OsHox33 function, we analyzed chloroplast ultrastructure in different-aged leaves of RNAi plants, and found that OsHox33 knockdown accelerated chloroplast degradation, which is consistent with RNAi phenotypes. Finally, real-time PCR studies showed that OsHox33 can regulate the expression of GS1 and GS2, two senescence-associated genes. Taken together, the work presented here provides new insights into the function of HD-Zip III members in plants.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Yoshida S. Molecular regulation of leaf senescence. Curr Opin Plant Biol, 2003, 6: 79–84
Himelblau E, Amasino M. Nutrients mobilized from leaves of Arabidopsis thaliana during leaf senescence. J Plant Physiol, 2001, 158: 1317–1323
Lohman K N, Gan S S, John M C, et al. Molecular analysis of natural senescence in Arabidopsis thaliana. Plant Physiol, 1994, 92: 322–328
van der Graaff E, Schwacke R, Schneider A, et al. Transcription analysis of Arabidopsis membrane transporters and hormone pathways during developmental and induced leaf senescence. Plant Physiol, 2006, 141: 776–792
Tanaka R, Hirashima M, Satoh S, et al. The Arabidopsis-accelerated cell death gene ACD1 is involved in oxygenation of pheophorbide a: Inhibition of the pheophorbide a oxygenase activity does not lead to the “stay-green” phenotype in Arabidopsis. Plant Cell Physiol, 2003, 44: 1266–1274
Ren G, An K, Liao Y, et al. Identification of a novel chloroplast protein AtNYE1 regulating chlorophyll degradation during leaf senescence in Arabidopsis. Plant Physiol, 2007, 144: 1429–1441
Kusaba M, Ito H, Morita R, et al. Rice NON-YELLOW COLORING1 is involved in light-harvesting complex II and grana degradation during leaf senescence. Plant Cell, 2007, 19: 1362–1375
Jiang H, Li M, Liang N, et al. Molecular cloning and function analysis of the stay green gene in rice. Plant J, 2007, 52: 197–209
Park S Y, Yu J W, Park J S, et al. The senescence-induced staygreen protein regulates chlorophyll degradation. Plant Cell, 2007, 19: 1649–1664
Morita R, Sato Y, Masuda Y, et al. Defect in NON YELLOW COLORING 3, an α/β hydrolase-fold family protein, causes a stay green phenotype during leaf senescence in rice. Plant J, 2009, 59: 940–952
Sato Y, Morita R, Katsuma S, et al. Two short-chain dehydrogenase/ reductases, NON-YELLOW COLORING 1 and NYC1-LIKE, are required for chlorophyll b and light-harvesting complex II degradation during senescence in rice. Plant J, 2009, 57: 120–131
Jiao B B, Wang J J, Zhu X D, et al. A novel protein RLS1 with NB-ARM domains is involved in chloroplast degradation during leaf senescence in rice. Mol Plant, 2012, 5: 205–217
Yamatani H, Sato Y, Masuda Y, et al. NYC4, the rice ortholog of Arabidopsis THF1, is involved in the degradation of chlorophyll-protein complexes during leaf senescence. Plant J, 2013, doi: 10. 1111/tpj.12154
Moore B, Zhou L, Rolland F, et al. Role of the Arabidopsis glucose sensor HXK1 in nutrient, light, and hormonal signaling. Science, 2003, 300: 332–336
Hinderhofer K, Zentgraf U. Identification of a transcription factor specifically expressed at the onset of leaf senescence. Planta, 2001, 213: 469–473
Woo H R, Chung K M, Park J H, et al. ORE9, an F-box protein that regulates leaf senescence in Arabidopsis. Plant Cell, 2001, 13: 1779–1790
Yoshida S, Ito M, Nishida I, et al. Identification of a novel gene HYS1/CPR5 that has a repressive role in the induction of leaf senescence and pathogen-defence responses in Arabidopsis thaliana. Plant J, 2002, 29: 427–437
Robatzek S, Somssich I E. Targets of AtWRKY6 regulation during plant senescence and pathogen defense. Genes Dev, 2002, 16: 1139–1149
Miao Y, Laun T, Zimmermann P, et al. Targets of the WRKY53 transcription factor and its role during leaf senescence in Arabidopsis. Plant Mol Biol, 2004, 55: 853–867
Guo Y, Gan S. AtNAP, a NAC family transcription factor, has an important role in leaf senescence. Plant J, 2006, 46: 601–612
Lim P O, Kim Y, Breeze E, et al. Overexpression of a chromatin architecture-controlling AT-hook protein extends leaf longevity and increases the post-harvest storage life of plants. Plant J, 2007, 52: 1140–1153
Kim J H, Woo H R, Kim J, et al. Trifurcate feed-forward regulation of age-dependent cell death involving miR164 in Arabidopsis. Science, 2009, 323: 1053–1057
Hou K, Wu W, Gan S S. SAUR36, a small auxin up RNA gene, is involved in the promotion of leaf senescence in Arabidopsis. Plant Physiol, 2013, 161: 1002–1009
Doelling J H, Walker J M, Friedman E M, et al. The APG8/12-activating enzyme APG7 is required for proper nutrient recycling and senescence in Arabidopsis thaliana. J Biol Chem, 2002, 277: 33105–33114
Hanaoka H, Noda T, Yoshimoto K, et al. Leaf senescence and starvation-induced chlorosis are accelerated by the disruption of an Arabidopsis autophagy gene. Plant Physiol, 2002, 129: 1181–1193
Xiong Y, Contento A L, Bassham D C. AtATG18a is required for the formation of autophagosomes during nutrient stress and senescence in Arabidopsis thaliana. Plant J, 2005, 42: 535–546
Raab S, Drechsel G, Zarepour M, et al. Identification of a novel E3 ubiquitin ligase that is required for suppression of premature senescence in Arabidopsis. Plant J, 2009, 59: 39–51
Besseau S, Li J, Palva E T. WRKY54 and WRKY70 co-operate as negative regulators of leaf senescence in Arabidopsis thaliana. J Exp Bot, 2012, 63: 2667–2679
Ramachandran S, Hiratsuka K, Chua N H. Transcription factors in plant growth and development. Curr Opin Genet Dev, 1994, 4: 642–646
Ariel F D, Manavella P A, Dezar C A, et al. The true story of the HD-Zip family. Trends Plant Sci, 2007, 12: 419–426
Kong Z, Li M, Yang W, et al. A novel nuclear-localized CCCH-type zinc finger protein, OsDOS, is involved in delaying leaf senescence in rice (Oryza sativa L.). Plant Physol, 2006, 141: 1376–1388
Jan A, Maruyama K, Todaka D, et al. OsTZF1, a CCCH-tandem zinc finger protein, confers delayed senescence and stress tolerance in rice by regulating stress-related genes. Plant Physiol, 2013, 161: 1202–1216
Söderman E, Mattsson J, Engstrom P. The Arabidopsis homeobox gene ATHB-7 is induced by water deficit and by abscisic acid. Plant J, 1996, 10: 375–381
Söderman E, Hjellström M, Fahleson J, et al. The HD-Zip gene ATHB6 in Arabidopsis is expressed in developing leaves, roots and carpels and up-regulated by water deficit conditions. Plant Mol Biol, 1999, 40: 1073–1083
Hanson J, Johannesson H, Engström P. Sugar-dependent alterations in cotyledon and leaf development in transgenic plants expressing the HDZhdip gene ATHB13. Plant Mol Biol, 2001, 45: 247–262
Rueda E C, Dezar C A, Gonzalez D H, et al. Hahb-10, a sunflower homeobox-leucine zipper gene, is regulated by light quality and quantity, and promotes early flowering when expressed in Arabidopsis. Plant Cell Physiol, 2005, 46: 1954–1963
Manavella P A, Arce A L, Dezar C A, et al. Cross-talk between ethylene and drought signalling pathways is mediated by the sunflower Hahb-4 transcription factor. Plant J, 2006, 48: 125–137
Steindler C, Matteucci A, Sessa G, et al. Shade avoidance responses are mediated by the ATHB-2 HD-zip protein, a negative regulator of gene expression. Development, 1999, 126: 4235–4245
Sawa S, Ohgishi M, Goda H, et al. The HAT2 gene, a member of the HD-Zip gene family, isolated as an auxin inducible gene by DNA microarray screening, affects auxin response in Arabidopsis. Plant J, 2002, 32: 1011–1022
Abe M, Katsumata H, Komeda Y, et al. Regulation of shoot epidermal cell differentiation by a pair of homeodomain proteins in Arabidopsis. Development, 2003, 130: 635–643
Ohashi-Ito K, Fukuda H. HD-zip III homeobox genes that include a novel member, ZeHB-13 (Zinnia)/ATHB-15 (Arabidopsis), are involved in procambium and xylem cell differentiation. Plant Cell Physiol, 2003, 44: 1350–1358
Nakamura A, Nakajima N, Goda H, et al. Arabidopsis Aux/IAA genes are involved in brassinosteroid-mediated growth responses in a manner dependent on organ type. Plant J, 2006, 45: 193–205
Baima S, Possenti M, Matteucci A, et al. The Arabidopsis ATHB-8 HD-zip protein acts as a differentiation-promoting transcription factor of the vascular meristems. Plant Physiol, 2001, 126: 643–655
McConnell J R, Emery J, Eshed Y, et al. Role of PHABULOSA and PHAVOLUTA in determining radial patterning in shoots. Nature, 2001, 411: 709–713
Emery J F, Floyd S K, Alvarez J, et al. Radial patterning of Arabidopsis shoots by class III HD-ZIP and KANADI genes. Curr Biol, 2003, 13: 1768–1774
Prigge M J, Otsuga D, Alonso J M, et al. Class III homeodomain-leucine zipper gene family members have overlapping, antagonistic, and distinct roles in Arabidopsis development. Plant Cell, 2005, 17: 61–76
Mallory A C, Reinhart B J, Jones-Rhoades M W, et al. microRNA control of PHABULOSA in leaf development: Importance of pairing to the microRNA 5′ region. EMBO J, 2004, 23: 3356–3364
Kim J, Jung J H, Reyes J L, et al. microRNA-directed cleavage of ATHB15 mRNA regulates vascular development in Arabidopsis inflorescence stems. Plant J, 2005, 42: 84–94
Williams L, Grigg S P, Xie M, et al. Regulation of Arabidopsis shoot apical meristem and lateral organ formation by microRNA miR166g and its AtHD-ZIP target genes. Development, 2005, 132: 3657–3668
Mallory A C, Vaucheret H. Functions of microRNAs and related small RNAs in plants. Nat Genet, 2006, 38: S31–S36
Smith Z R, Long J A. Control of Arabidopsis apical-basal embryo polarity by antagonistic transcription factors. Nature, 2010, 464: 423–426
Otsuga D, Deguzman B, Prigge M, et al. REVOLUTA regulates meristem initiation at lateral positions. Plant J, 2001, 25: 223–236
Prigge M J, Clark S E. Evolution of the class III HD-Zip gene family in land plants. Evol Dev, 2006, 8: 350–361
Green K A, Prigge M J, Katzman R B, et al. CORONA, a member of the class III homeodomain leucine zipper gene family in Arabidopsis, regulates stem cell specification and organogenesis. Plant Cell, 2005, 17: 691–704
Brandt R, Salla-Martret M, Bou-Torrent J, et al. Genome-wide binding-site analysis of REVOLUTA reveals a link between leaf patterning and light-mediated growth responses. Plant J, 2012, 72: 31–42
Robischon M, Du J, Miura E, et al. The Populus class III HD ZIP, popREVOLUTA, influences cambium initiation and patterning of woody stems. Plant Physio, 2011, 155: 1214–1225
Zhu Y, Song D, Sun J, et al. PtrHB7, a class III HD-Zip gene, plays a critical role in regulation of vascular cambium differentiation in Populus. Mol Plant, 2013, 6: 1331–1343
Itoh J, Hibara K, Sato Y, et al. Developmental role and auxin respon-siveness of Class III homeodomain leucine zipper gene family members in rice. Plant physiol, 2008, 147: 1960–1975
Xu F, Meng T, Li P, et al. A soybean dual-specificity kinase, GmSARK, and its Arabidopsis homolog, AtSARK, regulate leaf senescence through synergistic actions of auxin and ethylene. Plant Physiol, 2011, 157: 2131–2153
Olsson A S, Engström P, Söderman E. The homeobox genes ATHB12 and ATHB7 encode potential regulators of growth in response to water deficit in Arabidopsis. Plant Mol Biol, 2004, 55: 663–677
Hiei Y, Ohta S, Komari T, et al. Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J, 1994, 6: 271–282
Livak K J, Schmittgen T D. Analysis of relative gene expression data using real-time quantitative PCR and the {ie1123-1} method. Methods, 2001, 25: 402–408
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at Springerlink.com
Electronic supplementary material
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Luan, W., Shen, A., Jin, Z. et al. Knockdown of OsHox33, a member of the class III homeodomain-leucine zipper gene family, accelerates leaf senescence in rice. Sci. China Life Sci. 56, 1113–1123 (2013). https://doi.org/10.1007/s11427-013-4565-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-013-4565-2