Abstract
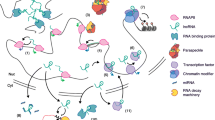
It is clear that RNA is more than just a messenger between gene and protein. The mammalian genome is pervasively transcribed, giving rise to tens of thousands of non-coding transcripts. Whether all of these transcripts are functional remains to be elucidated, but it is evident that there are many functional long non-coding RNAs (lncRNAs). Recent studies have set out to decode the regulatory role and functional diversity of lncRNAs. Here we organize these studies to highlight the significant involvements of lncRNAs in regulation of gene expression and human physiological and pathological processes, which are achieved by their interaction with DNA, RNA or protein.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Amaral P P, Dinger M E, Mercer T R, et al. The eukaryotic genome as an RNA machine. Science, 2008, 319: 1787–1789
Mattick J S. The genetic signatures of noncoding RNAs. PLoS Genet, 2009, 5: e1000459
Mercer T R, Dinger M E, Mattick J S. Long non-coding RNAs: Insights into functions. Nat Rev Genet, 2009, 10: 155–159
Carthew R W, Sontheimer E J. Origins and mechanisms of miRNAs and siRNAs. Cell, 2009, 136: 642–655
Guttman M, Amit I, Garber M, et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature, 2009, 458: 223–227
Ulitsky I, Shkumatava A, Jan C H, et al. Conserved function of lincRNAs in vertebrate embryonic development despite rapid sequence evolution. Cell, 2011, 147: 1537–1550
Spizzo R, Almeida M I, Colombatti A, et al. Long non-coding RNAs and cancer: A new frontier of translational research? Oncogene, 2012, 31: 4577–4587
Tsai M C, Spitale R C, Chang H Y. Long intergenic noncoding RNAs: New links in cancer progression. Cancer Res, 2011, 71: 3–7
McPherson R, Pertsemlidis A, Kavaslar N, et al. A common allele on chromosome 9 associated with coronary heart disease. Science, 2007, 316: 1488–1891
Johnson R. Long non-coding RNAs in Huntington’s disease neurodegeneration. Neurobiol Dis, 2012, 46: 245–254
Tan L, Yu J T, Hu N, et al. Non-coding RNAs in Alzheimer’s disease. Mol Neurobiol, 2013, 47: 382–393
Du Toit A. Non-coding RNA: RNA stability control by Pol II. Nat Rev Mol Cell Biol, 2013, 14: 128
Carninci P, Kasukawa T, Katayama S, et al. The transcriptional landscape of the mammalian genome. Science, 2005, 309: 1559–1563
Djebali S, Davis C A, Merkel A, et al. Landscape of transcription in human cells. Nature, 2012, 489: 101–108
Cheng J, Kapranov P, Drenkow, J, et al. Transcriptional maps of 10 human chromosomes at 5-nucleotide resolution. Science, 2005, 308: 1149–1154
Goodrich J A, Kugel J F. Dampening DNA binding: A common mechanism of transcriptional repression for both ncRNAs and protein domains. RNA Biol, 2010, 7: 305–309
Ravasi T, Ruzuki H, Pang K C, et al. Experimental validation of the regulated expression of large numbers of non-coding RNAs from the mouse genome. Genome Res, 2006, 16: 11–19
Batista P J, Chang H Y. Long noncoding RNAs: Cellular address codes in development and disease. Cell, 2013, 152: 1298–1307
Ponting C P, Oliver P L, Reik W. Evolution and functions of long noncoding RNAs. Cell, 2009, 136: 629–641
Prasanth K V, Spector D L. Eukaryotic regulatory RNAs: An answer to the “genome complexity” conundrum. Gene Dev, 2007, 21: 11–42
Wilusz J E, Sunwoo H, Spector D L. Long noncoding RNAs: Functional surprises from the RNA world. Genes Dev, 2009, 23: 1494–1504
Cremer T, Kupper K, Dietzel S, et al. Higher order chromatin architecture in cell nucleus: On the way from structure to function. Biol Cell, 2004, 96: 555–567
Fox A H, Lamond A I. Paraspeckles. Cold Spring Harb Perspect Biol, 2010, 2: a000687
Kapranov P, Cheng J, Dike S, et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science, 2007, 316: 1484–1488
Matera A G, lzaguire-Sierra M, Praveen K, et al. Nuclear bodies: Random aggregates of sticky proteins or crucibles of macromolecular assembly? Dev Cell, 2009, 17: 639–647
Spector D L. SnapShot: Cellular bodies. Cell, 2006, 127: 1071
Cautron-Herger M, Rippe K. Nuclear architecture by RNA. Curr Opin Genet Dev, 2012, 22: 179–187
Kino T, Hurt D E, Ichijo T, et al. Noncoding RNA gas5 is a growth arrest- and starvation-associated repressor of the glucocorticoid receptor. Sci Signal, 2010, 3: ra8
Martianov I, Ramadass A, Serra Barros A, et al. Repression of the human dihydrofolate reductase gene by a non-coding interfering transcript. Nature, 2007, 445: 666–670
Cesana M, Cacchiarelli D, Legnini I, et al. A long noncoding RNA controls muscle differetiation by functioning as a competing endogenous RNA. Cell, 2011, 147: 358–369
Keniry A, Oxley D, Monnier P, et al. The H19 lincRNA is a developmental reservoir of miR-675 that suppresses growth and lgf1r. Nat Cell Biol, 2012, 14: 659–665
Guil S, Esteller M. Cis-acting noncoding RNAs: Friends and foes. Nat Struct Mol Biol, 2012, 19: 1068–1075
Wang K C, Chang H Y. Molecular mechanisms of long noncoding RNAs. Mol Cell, 2011, 43: 904–914
Lee J T. Epigenetic regulation by long noncoding RNAs. Science, 2012, 338: 1435–1439
Rinn J L, Chang H Y. Genome regulation by long noncoding RNAs. Annu Rev Biochem, 2012, 81: 145–166
Kelley R L, Meller V H, Gordadze P R, et al. Epigenetic spreading of the Drosophila dosage compensation complex from roX RNA genes into flanking chromatin. Cell, 1999, 98: 513–522
Koziol M J, Rinn J L. RNA traffic control of chromatin complexes. Curr Opin Genet Dev, 2010, 20: 142–148
Mancini-Dinardo D, Steele S J, Levorse J M. Elongation of Kcnq1ot1 transcript is required for genomic imprinting of neighboring genes. Genes Dev, 2006, 20: 1268–1282
Fitzpatrick G V, Soloway P D, Higgins M J. Regional loss of imprinting and growth deficiency in mice with a targeted deletion of KvDMR1. Nat Genet, 2002, 32: 426–431
Pandey R R, Mondal T, Mohammad F, et al. Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol Cell, 2008, 32: 232–246
Rinn J L, Kertesz M, Wang J K, et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell, 2007, 129: 1311–1323
Nagano T, Mitchell J A, Sanz L A, et al. The Air noncoding RNA epigenetically silences transcription by targeting G9a to chromatin. Science, 2008, 322: 1717–1720
Tsai M C, Manor O, Wan Y, et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science, 2010, 329: 689–693
Gupta R A, Shah N, Wang K C, et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature, 2010, 464: 1071–1076
Mohammad F, Pandey G K, Mondal T, et al. Long noncoding RNA-mediated maintenance of DNA methylation and transcriptional gene silencing. Development, 2012, 139: 2792–2803
Wang X, Arai S, Song X, et al. Induced ncRNAs allosterically modify RNA-binding proteins in cis to inhibit transcription. Nature, 2008, 454: 126–130
Lanz R B, Razani B, Goldberg A D, et al. Distinct RNA motifs are important for coactivation of steroid hormone receptors by steroid receptor RNA activator (SRA). Proc Natl Acad Sci USA, 2002, 99: 16081–16086
Feng J, Bi C, Clark B S, et al. The Evf-2 noncoding RNA is transcribed from the Dlx-5/6 ultraconserved region and functions as a Dlx-2 transcriptional coactivator. Genes Dev, 2006, 20: 1470–1484
Kim T K, Hemberg M, Gray J M, et al. Widespread transcription at neuronal activity-regulated enhancers. Nature, 2010, 465: 182–187
Hah N, Danko C G, Core L, et al. A rapid, extensive, and transient transcriptional response to estrogen signaling in breast cancer cells. Cell, 2011, 145: 622–634
Wang D, Garcia-Bassets I, Benner C, et al. Reprogramming transcription by distinct classes of enhancers functionally defined by eRNA. Nature, 2011, 474: 390–394
Li W, Notani D, Ma Q, et al. Functional roles of enhancer RNAs for oestrogen-dependent transcriptional activation. Nature, 2013, 498: 516–520
Pan Q, Shai O, Lee L J, et al. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat Genet, 2008, 40: 1413–1415
Bernard D, Prasanth K V, Tripathi V, et al. A long nuclear-retained non-coding RNA regulates synaptogenesis by modulating gene expression. EMBO J, 2010, 29: 3082–3093
Tripathi V, Ellis J D, Shen Z, et al. The nuclear-retained noncoding RNA MALAT1 regulates alternative splicing by modulating SR splicing factor phosphorylation. Mol Cell, 2010, 39: 925–938
Sellier C, Rau F, Liu Y, et al. Sam68 sequestration and partial loss of function are associated with splicing alterations in FXTAS patients. EMBO J, 2010, 29: 1248–1261
Hansen T B, Jensen T I, Clausen B H, et al. Natural RNA circles function as efficient microRNA sponges. Nature, 2013, 495: 384–388
Franco-Zorrilla J M, Valli A, Todesco M, et al. Target mimicry provides a new mechanism for regulation of microRNA activity. Nat Genet, 2007, 39: 1033–1037
Poliseno L, Salmena L, Zhang J, et al. A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature, 2010, 465: 1033–1038
Karreth F A, Tay Y, Perna D, et al. In vivo identification of tumor-suppressive PTEN ceRNAs in an oncogenic BRAF-induced mouse model of melanoma. Cell, 2011, 147: 382–395
Tay Y, Kats L, Salmena L, et al. Coding-independent regulation of the tumor suppressor PTEN by competing endogenous mRNAs. Cell, 2011, 147: 344–357
Sumazin P, Yang X, Chiu H S, et al. An extensive microRNA-mediated network of RNA-RNA interactions regulates established oncogenic pathways in glioblastoma. Cell, 2011, 147: 370–381
Wang H, Iacoangeli A, Lin D, et al. Dendritic BC1 RNA in translational control mechanisms. J Cell Biol, 2005, 171: 811–821
Centonze D, Rossi S, Napoli I, et al. The brain cytoplasmic RNA BC1 regulates dopamine D2 receptor-mediated transmission in the striatum. J Neurosci, 2007, 27: 8885–8892
Lewejohann L, Skryabin B V, Sachser N, et al. Role of a neuronal small non-messenger RNA: Behavioural alterations in BC1 RNA-deleted mice. Behav Brain Res, 2004, 154: 273–289
Gong C, Maquat L E. lncRNAs transactivate stau1-mediated mRNA decay by duplexing with 3′ UTRs via Alu elements. Nature, 2011, 470: 284–288
Pibouin L, Villaudy J, Ferbus D, et al. Cloning of the mRNA of overexpression in colon carcinoma-1: A sequence overexpressed in a subset of colon carcinomas. Cancer Genet Cytogenet, 2002, 133: 55–60
Fu X, Ravindranath L, Tran N, et al. Regulation of apoptosis by a prostate-specific and prostate cancer-associated noncoding gene, PCGEM1. DNA Cell Biol, 2006, 25: 135–141
Ji P, Diederichs S, Wang W, et al. MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in earlystage non-small cell lung cancer. Oncogene, 2003, 22: 8031–8041
Galietta A, Gunby R H, Redaelli S, et al. NPM/ALK binds and phosphorylates the RNA/DNA-binding protein PSF in anaplastic large-cell lymphoma. Blood, 2007, 100: 2600–2609
Figueroa A, Fujita Y, Gorospe M. Hacking RNA: Hakai promotes tumorigenesis by enhancing the RNA-binding function of PSF. Cell Cycle, 2009, 8: 3648–3651
Song X, Wang B, Bromberg M, et al. Retroviral-mediated transmission of a mouse VL30 RNA to human melanoma cells promotes metastasis in an immunodeficient mouse model. Proc Natl Acad Sci USA, 2002, 99: 6269–6273
Song X, Sui A, Garen A. Binding of mouse VL30 retrotransposon RNA to PSF protein induces genes repressed by PSF: Effects on steroidogenesis and oncogenesis. Proc Natl Acad Sci USA, 2004, 101: 621–626
Song X, Sun Y, Garen A. Roles of PSF protein and VL30 RNA in reversible gene regulation. Proc Natl Acad Sci USA, 2005, 102: 12189–12193
Wang G, Cui Y, Zhang G, et al. Regulation of proto-oncogene transcription, cell proliferation, and tumorigenesis in mice by PSF protein and a VL30 noncoding RNA. Proc Natl Acad Sci USA, 2009, 106: 16794–16798
Li L, Feng T, Lian Y, et al. Role of human noncoding RNAs in the control of tumorigenesis. Proc Natl Acad Sci USA, 2009, 106: 16794–16798
Yap K L, Munoz-Cabello A M, Raguz S, et al. Molecular interplay of the noncoding RNA ANRIL and methylated histone H3 lysine 27 by polycomb CBX7 in transcriptional silencing of INK4a. Mol Cell, 2010, 38: 662–674
Lane D P, Fischer P M. Turning the key on p53. Nature, 2004, 427: 789–790
Lowe S W, Cepero E, Evan G. Intrinsic tumour suppression. Nature, 2004, 432: 307–315
Vogelstein B B, Lane D D, Levine A A. Surfing the p53 network. Nature, 2000, 408: 307–310
Yu J, Zhang L, Hwang P M, et al. Identification and classification of p53-regulated genes. Proc Natl Acad Sci USA, 1999, 96: 14517–14522
Zhao R, Gish K, Mruphy M, et al. Analysis of p53-regualted gene expression patterns using oligonucleotide arrays. Genes Dev, 2000, 14: 981–993
Brugarolas J, Chandrasekaran C, Gordon J I, et al. Radiation-induced cell cycle arrest compromised by p21 deficiency. Nature, 1995, 337: 552–557
Suzuki H I, Miyazono K. Dynamics of microRNA biogenesis: Crosstalk between p53 network and microRNA processing pathway. J Mol Med (Berl), 2010, 88: 1085–1094
Huarte M, Guttman M, Feldser D, et al. A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell, 2010, 142: 409–419
Hung T, Wang Y, Lin M F, et al. Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nat Genet, 2011, 43: 621–629
Zhang A, Zhou N, Huang J, et al. The human long non-coding RNA-RoR is a p53 repressor in response to DNA damage. Cell Res, 2013, 23: 340–350
Peng X, Gralinski L, Armour C D, et al. Unique signatures of long noncoding RNA expression in response to virus infection and altered innate immune signaling. MBio, 2010, 1: e00206–e00210
Collier S P, Collins P L, Williams C L, et al. Cutting edge: Influence of Tmevpg1, a long intergenic noncoding RNA, on the expression of lfng by Th1 cells. J Immunol, 2012, 189: 2084–2088
Gomez J A, Wapinski O L, Yang Y W, et al. The NeST long ncRNA controls microbial susceptibility and epigenetic activation of the interferon-γ locus. Cell, 2013, 152: 743–754
Medzhitov R, Horng T. Transcriptional control of the inflammatory response. Nat Rev Immunol, 2009, 9: 692–703
Carpenter S, Aiello D, Atianand M K, et al. A long noncoding RNA mediates both activation and repression of immune response genes. Science, 2013, 341: 789–792
Author information
Authors and Affiliations
Corresponding authors
Additional information
This article is published with open access at Springerlink.com
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Qi, W., Song, X. & Li, L. Long non-coding RNA-guided regulation in organisms. Sci. China Life Sci. 56, 891–896 (2013). https://doi.org/10.1007/s11427-013-4558-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-013-4558-1