Abstract
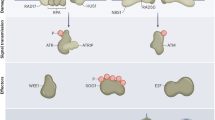
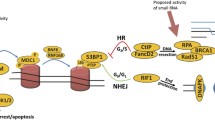
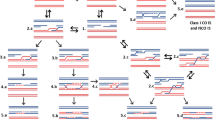
DNA double-strand break (DSB) is the most deleterious form of DNA damage and poses great threat to genome stability. Eukaryotes have evolved complex mechanisms to repair DSBs through coordinated actions of protein sensors, transducers, and effectors. DSB-induced small RNAs (diRNAs) or Dicer/Drosha-dependent RNAs (DDRNAs) have been recently discovered in plants and vertebrates, adding an unsuspected RNA component into the DSB repair pathway. DiRNAs/DDRNAs control DNA damage response (DDR) activation by affecting DDR foci formation and cell cycle checkpoint enforcement and are required for efficient DSB repair. Here, we summarize the findings of diRNAs/DDRNAs and discuss the possible mechanisms through which they act to facilitate DSB repair.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Jackson S P, Bartek J. The DNA-damage response in human biology and disease. Nature, 2009, 461: 1071–1078
Ciccia A, Elledge S J. The DNA damage response: Making it safe to play with knives. Mol Cell, 2010, 40: 179–204
Fillingham J, Keogh M C, Krogan N J. GammaH2AX and its role in DNA double-strand break repair. Biochem Cell Biol, 2006, 84: 568–577
Paull T T, Rogakou E P, Yamazaki V, et al. A critical role for histone H2AX in recruitment of repair factors to nuclear foci after DNA damage. Curr Biol, 2000, 10: 886–895
Podhorecka M, Skladanowski A, Bozko P. H2AX phosphorylation: Its role in DNA damage response and cancer therapy. J Nucleic Acids, 2010, 2010: 920161
Moynahan M E, Jasin M. Mitotic homologous recombination maintains genomic stability and suppresses tumorigenesis. Nat Rev Mol Cell Biol, 2010, 11: 196–207
San Filippo J, Sung P, Klein H. Mechanism of eukaryotic homologous recombination. Annu Rev Biochem, 2008, 77: 229–257
Lieber M R. The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu Rev Biochem, 2010, 79: 181–211
Carthew R W, Sontheimer E J. Origins and mechanisms of miRNAs and siRNAs. Cell, 2009, 136: 642–655
Ghildiyal M, Zamore P D. Small silencing RNAs: An expanding universe. Nat Rev Genet, 2009, 10: 94–108
Brennecke J, Aravin A A, Stark A, et al. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell, 2007, 128: 1089–1103
Houwing S, Kamminga L M, Berezikov E, et al. A role for Piwi and piRNAs in germ cell maintenance and transposon silencing in zebrafish. Cell, 2007, 129: 69–82
Vagin V V, Sigova A, Li C, et al. A distinct small RNA pathway silences selfish genetic elements in the germline. Science, 2006, 313: 320–324
Malone C D, Hannon G J. Small RNAs as guardians of the genome. Cell, 2009, 136: 656–668
Ishizu H, Siomi H, Siomi M C. Biology of Piwi-interacting RNAs: New insights into biogenesis and function inside and outside of germlines. Genes Dev, 2012, 26: 2361–2373
Havecker E R, Wallbridge L M, Hardcastle T J, et al. The Arabidopsis RNA-directed DNA methylation argonautes functionally diverge based on their expression and interaction with target loci. Plant Cell, 2010, 22: 321–334
Law J A, Jacobsen S E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat Rev Genet, 2010, 11: 204–220
Matzke M, Kanno T, Daxinger L, et al. RNA-mediated chromatin-based silencing in plants. Curr Opin Cell Biol, 2009, 21: 367–376
Wei W, Ba Z, Gao M, et al. A role for small RNAs in DNA double-strand break repair. Cell, 2012, 149: 101–112
Francia S, Michelini F, Saxena A, et al. Site-specific Dicer and Drosha RNA products control the DNA-damage response. Nature, 2012, 488: 231–235
Michalik K M, Bottcher R, Forstemann K. A small RNA response at DNA ends in Drosophila. Nucleic Acids Res, 2012, 40: 9596–9603
Stewart G S, Wang B, Bignell C R, et al. MDC1 is a mediator of the mammalian DNA damage checkpoint. Nature, 2003, 421: 961–966
Stucki M, Clapperton J A, Mohammad D, et al. MDC1 directly binds phosphorylated histone H2AX to regulate cellular responses to DNA double-strand breaks. Cell, 2005, 123: 1213–1226
Lou Z, Minter-Dykhouse K, Franco S, et al. MDC1 maintains genomic stability by participating in the amplification of ATM-dependent DNA damage signals. Mol Cell, 2006, 21: 187–200
Stucki M, Jackson S P. GammaH2AX and MDC1: Anchoring the DNA-damage-response machinery to broken chromosomes. DNA Repair (Amst), 2006, 5: 534–543
Bekker-Jensen S, Lukas C, Melander F, et al. Dynamic assembly and sustained retention of 53BP1 at the sites of DNA damage are controlled by Mdc1/NFBD1. J Cell Biol, 2005, 170: 201–211
Polo S E, Jackson S P. Dynamics of DNA damage response proteins at DNA breaks: A focus on protein modifications. Genes Dev, 2011, 25: 409–433
Lee H C, Chang S S, Choudhary S, et al. qiRNA is a new type of small interfering RNA induced by DNA damage. Nature, 2009, 459: 274–277
Heyer W D, Ehmsen K T, Liu J. Regulation of homologous recombination in eukaryotes. Annu Rev Genet, 2010, 44: 113–139
Krejci L, Altmannova V, Spirek M, et al. Homologous recombination and its regulation. Nucleic Acids Res, 2012, 40: 5795–5818
van Attikum H, Gasser S M. Crosstalk between histone modifications during the DNA damage response. Trends Cell Biol, 2009, 19: 207–217
Rossetto D, Truman A W, Kron S J, et al. Epigenetic modifications in double-strand break DNA damage signaling and repair. Clin Cancer Res, 2010, 16: 4543–4552
Sanders S L, Portoso M, Mata J, et al. Methylation of histone H4 lysine 20 controls recruitment of Crb2 to sites of DNA damage. Cell, 2004, 119: 603–614
Botuyan M V, Lee J, Ward I M, et al. Structural basis for the methylation state-specific recognition of histone H4-K20 by 53BP1 and Crb2 in DNA repair. Cell, 2006, 127: 1361–1373
Pei H, Zhang L, Luo K, et al. MMSET regulates histone H4K20 methylation and 53BP1 accumulation at DNA damage sites. Nature, 2011, 470: 124–128
Fnu S, Williamson E A, De Haro L P, et al. Methylation of histone H3 lysine 36 enhances DNA repair by nonhomologous end-joining. Proc Natl Acad Sci USA, 2010, 108: 540–545
Nguyen A T, Zhang Y. The diverse functions of Dot1 and H3K79 methylation. Genes Dev, 2011, 25: 1345–1358
Bird A W, Yu D Y, Pray-Grant M G, et al. Acetylation of histone H4 by ESA1 is required for DNA double-strand break repair. Nature, 2002, 419: 411–415
Qin S, Parthun M R. Histone H3 and the histone acetyltransferase Hat1p contribute to DNA double-strand break repair. Mol Cell Biol, 2002, 22: 8353–8365
Tamburini B A, Tyler J K. Localized histone acetylation and deacetylation triggered by the homologous recombination pathway of double-strand DNA repair. Mol Cell Biol, 2005, 25: 4903–4913
Murr R, Loizou J I, Yang Y G, et al. Histone acetylation by Trrap-Tip60 modulates loading of repair proteins and repair of DNA double-strand breaks. Nat Cell Biol, 2006, 8: 91–99
Huen M S, Grant R, Manke I, et al. RNF8 transduces the DNA-damage signal via histone ubiquitylation and checkpoint protein assembly. Cell, 2007, 131: 901–914
Kolas N K, Chapman J R, Nakada S, et al. Orchestration of the DNA-damage response by the RNF8 ubiquitin ligase. Science, 2007, 318: 1637–1640
Mailand N, Bekker-Jensen S, Faustrup H, et al. RNF8 ubiquitylates histones at DNA double-strand breaks and promotes assembly of repair proteins. Cell, 2007, 131: 887–900
Doil C, Mailand N, Bekker-Jensen S, et al. RNF168 binds and amplifies ubiquitin conjugates on damaged chromosomes to allow accumulation of repair proteins. Cell, 2009, 136: 435–446
Stewart G S, Panier S, Townsend K, et al. The riddle syndrome protein mediates a ubiquitin-dependent signaling cascade at sites of DNA damage. Cell, 2009, 136: 420–434
Messick T E, Greenberg R A. The ubiquitin landscape at DNA double-strand breaks. J Cell Biol, 2009, 187: 319–326
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at Springerlink.com
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Ba, Z., Qi, Y. Small RNAs: Emerging key players in DNA double-strand break repair. Sci. China Life Sci. 56, 933–936 (2013). https://doi.org/10.1007/s11427-013-4552-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-013-4552-7