Abstract
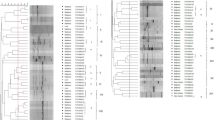
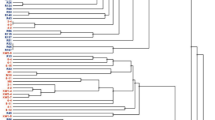
Diversity of 42 isolates from effective nodules of Pisum sativum in different geographical regions of China were studied using 16S rRNA gene RFLP patterns, 16S rRNA sequencing, 16S–23S rRNA intergenic spacer (IGS) region RFLP patterns and G-C rich random amplified polymorphic DNA (RAPD). The isolates were distributed in two groups on the basis of their 16S rRNA gene RFLP patterns. The 16S rRNA gene sequences of strains from 16S rRNA gene RFLP patterns group I were very closely related (identities higher than 99.5%) to Rhizobium leguminosarum USDA 2370. Group II consisting of WzP3 and WzP15 was closely related to Rhizobium etli CFN42. The analysis of the 16S-23S IGS RFLP patterns divided the isolates into 18 genotypes and four groups. Group I was clustered with R. leguminosarum USDA2370. Group II consisted of YcP2, YcP3 and CqP7. The strains of group III were distributed abroad. Group IV consisted of WzP3, WzP15 and R. etli CFN42. RAPD divided the isolates into nine clusters in which group IV only consisted of YcP2 and the strains of group V and IX were from Wenzhou and Xiantao, respectively. This assay demonstrated the geographical effect on genetic diversity of pea rhizobia.
Similar content being viewed by others
References
Yang C Y, Li Y G, Zhou J C, et al. The Function of Three Indigenous Plasmids in Mesorhizobium huakuii 2020 and its Symbiotic Interaction with Sym pJB5JI of Rhizobium leguminosarum. Sci China Ser C-Life Sci, 2008, 4: 35–361
Hadri A E, Bisseling T. Responses of the plant to Nod factors. In The Rhizobiaceae, ed. by Spaink H P, Kondorosi A, Hooykaas P J J. Dordrecht, The Netherlands: Kluwer Academic Publishers, 1998, pp: 403–416
Vance C P. Legume symbiotic nitrogen fixation: Agronomic aspects. In The Rhizobiaceae, ed. by Spaink H P, Kondorosi A, Hooykaas P J J. Dordrecht, The Netherlands: Kluwer Academic Publishers, 1998, pp: 509–530
Jourand P, Giraud E, Bena G, et al. Methylobacterium nodulans sp. nov., for a group of aerobic, facultatively methylotrophic, legume root-nodule-forming and nitrogen-Wxing bacteria. Int J Syst Evol Microbiol, 2004, 54: 2269–2273, 15545469, 10.1099/ijs.0.02902-0, 1:CAS:528:DC%2BD2cXhtFGjsLzM
Rivas R, Velázquez E, Willems A, et al. A new species of Devosia that forms a unique nitrogen-Wxing root-nodule symbiosis with the aquatic legume Neptunia natans (L.F.) Druce. Appl Environ Microbiol, 2002, 68: 5217–5222, 12406707, 1:CAS:528:DC%2BD38Xos1Wqur8%3D, 10.1128/AEM.68.11.5217-5222.2002
van Berkum P, Eardly B D. The aquatic budding bacterium Blastobacter denitriWcans is a nitrogen-Wxing symbiont of Aeschynomene indica. Appl Environ Microbiol, 2002, 68: 1132–1136, 11872460, 10.1128/AEM.68.3.1132-1136.2002, 1:CAS:528:DC%2BD38XitV2qu7o%3D
Moulin L, Munive A, Dreyfus B, et al. Nodulation of legumes by members of the beta-subclass of Proteobacteria. Nature, 2001, 411:948–950, 11418858, 10.1038/35082070, 1:CAS:528:DC%2BD3MXkvVWhu7Y%3D
Chen W M, Moulin L, Bontemps C, et al. Legume symbiotic nitrogen Wxation by β-proteobacteria is widespread in nature. J Bacteriol, 2003, 185: 7266–7272, 14645288, 10.1128/JB.185.24.7266-7272.2003, 1:CAS:528:DC%2BD3sXpvVaqtro%3D
Chen W M, James E K, Chou J H, et al. β-rhizobia from Mimosa pigra, a newly discovered invasive plant in Taiwan. New Phytol, 2005, 168:661–675, 16313648, 10.1111/j.1469-8137.2005.01533.x, 1:CAS:528:DC%2BD2MXhtlWgs7%2FE
Trujillo M E, Willems A, Abril A, et al. Nodulation of Lupinus albus by Strains of Ochrobactrum lupini sp. inov. Appl Environ Microbiol, 2005, 71: 1318–1327, 15746334, 10.1128/AEM.71.3.1318-1327.2005, 1:CAS:528:DC%2BD2MXisVOis70%3D
Valverde A, Velázquez E, Fernández-Santos F, et al. Phyllobacterium trifolii sp. nov., nodulating Trifolium and Lupinus in Spanish soils. Int J Syst Evol Microbiol, 2005, 55: 1985–1989, 16166699, 10.1099/ijs.0.63551-0, 1:CAS:528:DC%2BD2MXhtFCku7bN
Rivas R, Willems A, Palomo L J, et al. Bradyrhizobium betae sp. nov., isolated from roots of Beta vulgaris aVected by tumour-like deformations. Int J Syst Bacteriol, 2004, 54: 1271–1275, 1:CAS:528:DC%2BD2cXnt1Gitrw%3D
Benhizia Y, Benhizia H, Benguedouar A, et al. Gamma proteobacteria can nodulate legumes of the genus Hedysarum. Syst Appl Microbiol, 2004, 27: 462–468, 15368852, 10.1078/0723202041438527, 1:CAS:528:DC%2BD2cXos1enurk%3D
Aguilar O M, Lo’pez M V, Riccillo P M. The diversity of rhizobia nodulating beans in Northwest Argentina as a source of more efficient inoculant strains. J Biotechnol, 2001, 91: 181–188, 11566389, 10.1016/S0168-1656(01)00336-4, 1:CAS:528:DC%2BD3MXmvVKltLY%3D
Palmer K M, Young J P W. Higher Diversity of Rhizobium leguminosarum Biovar viciae Populations in Arable Soils than in Grass Soils. Appl Environ Microbiol, 2000, 6: 2445–2450, 10.1128/AEM.66.6.2445-2450.2000
Vincent J M. A manual for the practical study of root-nodule bacteria. International biological programme handbook. Oxford: Blackwell, 1970. 73–97
Terefework Z, Kaijalainen S, Lindström K. AFLP fingerprint as a tool to study the genetic diversity of Rhizobium galegae isolated from Galega orientalis and Galega offcinalis. J Biotechnol, 2001, 91:169–180, 11566388, 10.1016/S0168-1656(01)00338-8, 1:CAS:528:DC%2BD3MXmvVKltLk%3D
Tan Z Y, Xu X D, Wang E T, et al. Phylogenetic and genetic relationships of Mesorhizobium tianshanense and related rhizobia. Int J Syst Bacteriol, 1997, 47: 874–879, 9226921, 1:CAS:528:DyaK2sXkvFChur0%3D
Deya A A M, Odelson D A, Hickey R F, et al. Bacterial community fingerprinting of amplified 16S and 16S–23S ribosomal DNA gene sequences and restriction endonuclease analysis (ARDRA). Mol Microb Ecol Manual, 1995, pp: 1–8
Kimura M. A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol, 1980, 16: 111–120, 7463489, 10.1007/BF01731581, 1:CAS:528:DyaL3MXmtFSktg%3D%3D
Zuckerkandl E, Pauling L. Molecules as documents of evolutionary history. J Theor Biol, 1965, 8: 357–366, 5876245, 10.1016/0022-5193(65)90083-4, 1:STN:280:DyaF2s7jvFyhtw%3D%3D
Xu L M, Ge C, Cui Z, et al. Bradyrhizobium liaoningense sp. nov., isolated from the root nodules of soybeans. Int J Syst Bacteriol, 1995, 45: 706–711, 7547289, 1:STN:280:DyaK28%2Fhs1OhsQ%3D%3D, 10.1099/00207713-45-4-706
Peng G X, Tan Z Y, Wang E T, et al. Identification of isolates from soybean nodules in Xinjiang Region as Sinorhizobium xinjiangense and genetic differentiation of S. xinjiangense from Sinorhizobium fredii. Int J Syst Evol Microbiol, 2002, 52: 457–462, 11931157, 1:CAS:528:DC%2BD38XivV2rtbs%3D
Krasova-Wade T, Ndoye I, Braconnier S, et al. Diversity of indigeneous bradyrhizobia associated with three cowpea cultivars (Vigna unguiculata (L.) Walp.) grown under limited and favorable water conditions in Senegal (West Africa). African J Biotech, 2003, 2:13–22, 1:CAS:528:DC%2BD3sXpslWgsA%3D%3D
Yang S S, Bellogin R A, Buendia A, et al. Effect of pH and soybean cultivars on the quantitative analyses of soybean rhizobia populations. J Biotechnol, 2001, 91: 243–255, 11566395, 10.1016/S0168-1656(01)00340-6, 1:CAS:528:DC%2BD3MXmvVKltbo%3D
Author information
Authors and Affiliations
Corresponding author
Additional information
Supported by the Chinese Microbe Resource Project (Grant No. 2005DKA21208-6), Chinese High-tech Developing Program (Grant No. 2007AA05Z417), and the Opening Foundation of State Key Laboratory of Agricultural Microbiology, HAU, China
Rights and permissions
About this article
Cite this article
Yang, C., Yang, J., Li, Y. et al. Genetic diversity of root-nodulating bacteria isolated from pea (Pisum sativum) in subtropical regions of China. SCI CHINA SER C 51, 854–862 (2008). https://doi.org/10.1007/s11427-008-0104-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-008-0104-y