Abstract
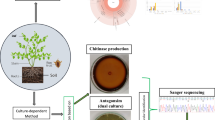
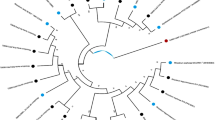
The traditional culture-dependent plate counting and culture-independent small-subunit-ribosomal RNA gene-targeted molecular techniques, Single-Strand Conformation Polymorphism (SSCP) and terminal Restriction Fragment Length Polymorphism (tRFLP) combined with 16S rDNA clone library were adopted to investigate the impacts of secretion from Camptotheca acuminata (abbreviated to Ca) roots on the quantities and structure of eukaryotic microbes and bacteria in the rhizosphere, and the possibility that Ca controls exotic invasive plant Eupatorium adenophorum (Ea). The counting results indicated that the number of bacteria increased in turn in rhizospheres of Ea, Ca-Ea mixed culture and Ca, while that of eukaryotic microbes decreased. PCR-SSCP profiles showed eukaryotic microbial bands (corresponding to biodiversity) in rhizosphere of Ea were more complex than those of Ca and CE. Meristolohmannia sp., Termitomyces sp. and Rhodophyllus sp. were the dominant populations in the rhizosphere of Ca. Bacterial terminal restriction fragments (TRFs) profiles showed no difference among three kinds of rhizospheres, and the sequences of the 16S rDNA clone library from Ca rhizospheres were distributed in 10 known phyla, in which phylum Proteobacteria were the absolute dominant group and accounted for 24.71% of the cloned sequences (δ-Proteobacteria accounted for up to 17.65%), and phyla Acidobacteria and Bacteroidetes accounted for 16.47% and 10.59% of the cloned sequences, respectively. In addition, high performance liquid chromatography detected a trace amount of camptothecin and hydroxycamptothecin in the rhizospheric soil of Ca and CE, but examined neither camptothecin nor hydroxycamptothecin in rhizospheric soil of Ea. Therefore, invasion and diffusion of Ea evidently depended on distinguishing the eukaryotic community structure, but not on that of the bacterial pattern. Ca was able to alter the eukaryotic community structure of invasive Ea by secreting camptothecin and hydroxycamptothecin into rhizospheres, and may benefit the control of overspread of Ea. This study provided theoretical evidence for rhizospheric microbial aspects on substituting Ca for Ea.
Similar content being viewed by others
References
Staker B L, Feese M D, Cushman M, et al. Structures of three classes of anticancer agents bound to the human topoisomerase I-DNA covalent complex. J Med Chem, 2005, 48(7): 2336–2345
Slesarev, A I, Lake J A, Stetter K O, et al. Purification and characterization of DNA topoisomerase V. An enzyme from the hyperthermophilic prokaryote Methanopyrus kandleri that resembles eukaryotic topoisomerase I. J Biol Chem, 1994, 269(5): 3295–3303
Flores H E, Weber C, Puffett J. Underground metabolism: The biosynthetic potential of roots, In: Waisel Y, et al., eds. Roots: The Hidden Half. 2nd ed. New York: Marcel Dekker, 1996. 931–956
Callaway R M, Newingham B, Zabinski C A, et al. Compensatory growth and competitive ability of an invasive weed are enhanced by soil fungi and native neighbors. Ecol Lett, 2001, 4(5): 429–433
Callaway R M, Thelen G C, Rodriguez A, et al. Soil biota and exotic plant invasion. Nature, 2004, 427: 731–733
Kourtev P S, Ehrenfeld J G, Haggblom M. Exotic plant species alter the microbial community structure and function in the soil. Ecology, 2002, 83(11): 3152–3166
Kourtev P S, Ehrenfeld J G, Haggblom M. Experimental analysis of the effect of exotic and native plant species on the structure and function of soil microbial communities. Soil Biol Biochem, 2003, 35: 895–905
Yu X J, Yu D, Lu Z J, et al. A new mechanism of invader success: Exotic plant inhibits natural vegetation restoration by changing soil microbe community. Chin Sci Bull, 2005, 50(11): 1105–1112
Yang C H, Crowley D E, Menge J A. 16S rDNA fingerprinting of rhizosphere bacterial communities associated with healthy and Phytophthora infected avocado roots. FEMS Microbiol Ecol, 2001, 35(2): 129–136
Schloter M, Bach H-J, Metz S, et al. Influence of precision farming on the microbial community structure and functions in nitrogen turnover. Agric Ecosyst Environ, 2003, 98: 295–304
Liu W T, Marsh T L, Cheng H, et al. Characterization of microbial diversity by determining terminal restriction fragment length polymorphisms of genes encoding 16S rRNA. Appl Environ Microbiol, 1997, 63: 4516–4522
Schwieger F, Tebbe C C. A new approach to utilize PCR-single-strand-conformation polymorphism for 16S rRNA gene-based microbial community analysis. Appl Environ Microbiol, 1998, 64: 4870–4876
Giovannoni S J, Britschgi T B, Moyer C L, et al. Genetic diversity in Sargasso Sea bacterioplankton. Nature, 1990, 345(6270): 60–63
State Environmental Protection Administration of China. Water and Wastewater Monitor Methods. 3rd ed. Beijing: Chinese Environmental Science Press, 1997. 216–220
Zu Y G, Zhao C J, Fu Y J, et al. Quantitative determination of camptothecin and hydroxycamptothecin in seeds of Camptotheca acuminata decaisne by wavelength detection in high performance liquid chromatography. Chinese J Anal Chem, 2004, 32(11): 1441–1444
Zhou D Q. Microbiology Experimental Manual. Shanghai: Shanghai Science and Technology Press, 1986
Peters S, Koschinsky S, Schwieger F, et al. Succession of microbial communities during hot composting as detected by PCR single strand conformation polymorphism based genetic profiles of small subunit rRNA genes. Appl Environ Microbiol, 2000, 66: 930–936
Subramanian K, Rutvisuttinunt W, Scott W, et al. The enzymatic basis of processivity in λ exonuclease. Nucleic Acids Res, 2003, 31(6): 1585–1596
Bassam B J, Caetano-Anolles G, Gresshoff P M. Fast and sensitive silver staining of DNA in polyacrylamide gels. Anal Biochem, 1991, 196(1): 80–83
Wang M, Ahrne S, Antonsson M, et al. T-RFLP combined with principal component analysis and 16S rRNA gene sequencing: An effective strategy for comparison of fecal microbiota in infants of different ages. J Microbiol Methods, 2004, 59(1): 53–69
Sandhu G S, Precup J W, Kline B C. Rapid one-step characterization of recombinant vectors by direct analysis of transformed Escherichia coli colonies. Biotechniques, 1989, 7(7): 689–690
Altschul S F, Madden T L, Schaffer A A, et al. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res, 1997, 25: 3389–3402
Neefs J M, Van de Peer Y, De Rijk P, et al. Compilation of small ribosomal subunit RNA structures. Nucleic Acids Res, 1993, 21(13): 3025–3049
Thompson J D, Higgins D G, Gibson T J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res, 1994, 22: 4673–4680
Felsenstein J. PHYLIP (Phylogeny Inference Package) version 3.6. Distributed by the author. Department of Genome Sciences, University of Washington, Seattle. 2005
Duda J J, Freeman D C, Emlen J M, et al. Differences in native soil ecology associated with invasion of the exotic annual chenopod, Halogeton glomeratus. Biol Fertil Soils, 2003, 38: 72–77
Miethling R, Ahrends K, Tebbe C. Structural differences in the rhizosphere communities of legumes are not equally reflected in community-level physiological profiles. Soil Biol Biochem, 2003, 35(10): 1405–1410
Suding K N, LeJeune K D, Seastedt T R. Competitive impacts and responses of an invasive weed: Dependencies on nitrogen and phosphorus availability. Oecologia, 2004, 141: 526–535
Ashton I W, Hyatt L A, Howe, K M, et al. Invasive species accelerate decomposition and litter nitrogen loss in a mixed deciduous forest. Ecol Appl, 2005, 15: 1263–1272
Callaway R M, Thelen G C, Barth S, et al. Soil fungi alter interactions between the invader Centaurea maculosa and north American Natives. Ecology, 2004, 85(4): 1062–1071
Ren N Q, Zhao Y G, Wang A J, et al. The effect of decreasing alkalinity on microbial community dynamics in a sulfate-reducing bioreactor as analyzed by PCR-SSCP. Sci China Ser C-Life Sci, 2006, 49(4): 370–378
Quaiser A, Ochsenreiter T, Lanz C, et al. Acidobacteria form a coherent but highly diverse group within the bacterial domain: Evidence from environmental genomics. Mol Microbiol, 2003, 50(2): 563–575
O’sullivan L A, Rinna J, Humphreys G, et al. Culturable phylogenetic diversity of the phylum ‘Bacteroidetes’ from river epilithon and coastal water and description of novel members of the family Flavobacteriaceae: Epilithonimonas tenax gen. nov., sp. nov. and Persicivirga xylanidelens gen. nov., sp. nov. Int J Syst Evol Microbiol, 2006, 56(1): 169–180
Cytryn E, Galfend I, Barak Y, et al. Diversity of microbial communities correlated to physiochemical parameters in a digestion basin of a zero-discharge mariculture system. Environ Microbiol, 2003, 5(1): 55–63
Filion M, Hamelin R C, Benier L, et al. Molecular profiling of rhizosphere microbial communities associated with healthy and diseased black spruce (Picea mariana) seedlings grown in a nursery. Appl Environ Microbiol, 2004, 70(6): 3541–3551
Dai X, Wang B J, Hang Y, et al. Bacterial diversity in the sediments of Taihu Lake by using traditional nutrient medium and dilution nutrient medium. Acta Microbiol Sin, 2005, 45(2): 161–165
Author information
Authors and Affiliations
Corresponding author
Additional information
Supported by the Excellent Young Teacher’s Innovation Foundation of Northeast Forestry University to Yang FengJian, the Key Research Fund of Ministry of Education of China (Grant No. 104191) and the Forestry Noxious Plant Investigation Fund of State Forestry Administration of China to Zu YuanGang
Rights and permissions
About this article
Cite this article
Zu, Y., Gao, C., Wang, W. et al. Characteristics of the microbial community in rhizosphere of Camptotheca acuminata cultured with exotic invasive plant Eupatorium adenophorum . SCI CHINA SER C 50, 22–30 (2007). https://doi.org/10.1007/s11427-007-0012-6
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s11427-007-0012-6