Abstract
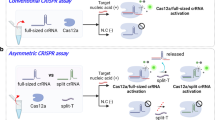
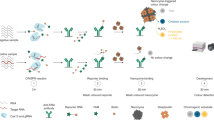
The efficient signal amplification capacity of several class 2 CRISPR-Cas systems with trans-cleavage activity has exhibited great value in molecular diagnostics, but its potential application for non-nucleic-acid targets is yet underdeveloped. Here, we deploy CRISPR-Cas system for the ultrasensitive detection of protease biomarkers by the coupling of proteolysis-triggered transcription. In this strategy, a protease-activatable RNA polymerase is adopted for the conversion of each protease-catalyzed proteolysis event into the output of multiple programable RNA sequences by in vitro transcription, and the transcribed RNA subsequently serves as the guide RNA of Cas12a proteins with trans-cleavage activity. The rational design of the transcribed RNA efficiently couples the signal conversion and amplification of proteolysis-triggered transcription and the self-signal amplification of CRISPR-Cas12a, resulting in a two-stage amplified detection of target protease. The versatility of this strategy has been demonstrated in the detection of protease biomarkers including MMP-2 and thrombin with femtomolar sensitivity, which is 5–6 orders of magnitude lower than that of the standard peptide-based methods. Moreover, the proposed method has been further applied in the analysis of MMP-2 secreted by different cancer cell lines as well the assessment of MMP-2 activity in clinical serum samples, providing a generic method for the ultrasensitive detection of protease biomarkers in biochemical research and clinical diagnosis.

Similar content being viewed by others
References
Siegel RL, Miller KD, Jemal A. CA-A Cancer J Clin, 2016, 66: 7–30
Wu L, Qu X. Chem Soc Rev, 2015, 44: 2963–2997
Sawyers CL. Nature, 2008, 452: 548–552
Zen K, Zhang CY. Med Res Rev, 2012, 32: 326–348
López-Otín C, Bond JS. J Biol Chem, 2008, 283: 30433–30437
Abudayyeh OO, Gootenberg JS, Konermann S, Joung J, Slaymaker IM, Cox DBT, Shmakov S, Makarova KS, Semenova E, Minakhin L, Severinov K, Regev A, Lander ES, Koonin EV, Zhang F. Science, 2016, 353: aaf5573
Harrington LB, Burstein D, Chen JS, Paez-Espino D, Ma E, Witte IP, Cofsky JC, Kyrpides NC, Banfield JF, Doudna JA. Science, 2018, 362: 839–842
Yang H, Gao P, Rajashankar KR, Patel DJ. Cell, 2016, 167: 1814–1828.e12
Li SY, Cheng QX, Liu JK, Nie XQ, Zhao GP, Wang J. Cell Res, 2018, 28: 491–493
Li SY, Cheng QX, Wang JM, Li XY, Zhang ZL, Gao S, Cao RB, Zhao GP, Wang J. Cell Discov, 2019, 5: 17
Chen JS, Ma E, Harrington LB, Da Costa M, Tian X, Palefsky JM, Doudna JA. Science, 2018, 360: 436–439
Gootenberg JS, Abudayyeh OO, Kellner MJ, Joung J, Collins JJ, Zhang F. Science, 2018, 360: 439–444
Gootenberg JS, Abudayyeh OO, Lee JW, Essletzbichler P, Dy AJ, Joung J, Verdine V, Donghia N, Daringer NM, Freije CA, Myhrvold C, Bhattacharyya RP, Livny J, Regev A, Koonin EV, Hung DT, Sabeti PC, Collins JJ, Zhang F. Science, 2017, 356: 438–442
Myhrvold C, Freije CA, Gootenberg JS, Abudayyeh OO, Metsky HC, Durbin AF, Kellner MJ, Tan AL, Paul LM, Parham LA, Garcia KF, Barnes KG, Chak B, Mondini A, Nogueira ML, Isern S, Michael SF, Lorenzana I, Yozwiak NL, MacInnis BL, Bosch I, Gehrke L, Zhang F, Sabeti PC. Science, 2018, 360: 444–448
Teng F, Guo L, Cui T, Wang XG, Xu K, Gao Q, Zhou Q, Li W. Genome Biol, 2019, 20: 132
Wang B, Wang R, Wang D, Wu J, Li J, Wang J, Liu H, Wang Y. Anal Chem, 2019, 91: 12156–12161
Hu M, Yuan C, Tian T, Wang X, Sun J, Xiong E, Zhou X. J Am Chem Soc, 2020, 142: 7506–7513
Dai Y, Somoza RA, Wang L, Welter JF, Li Y, Caplan AI, Liu CC. Angew Chem Int Ed, 2019, 58: 17399–17405
Liang M, Li Z, Wang W, Liu J, Liu L, Zhu G, Karthik L, Wang M, Wang KF, Wang Z, Yu J, Shuai Y, Yu J, Zhang L, Yang Z, Li C, Zhang Q, Shi T, Zhou L, Xie F, Dai H, Liu X, Zhang J, Liu G, Zhuo Y, Zhang B, Liu C, Li S, Xia X, Tong Y, Liu Y, Alterovitz G, Tan GY, Zhang LX. Nat Commun, 2019, 10: 3672
Xiong Y, Zhang J, Yang Z, Mou Q, Ma Y, Xiong Y, Lu Y. J Am Chem Soc, 2020, 142: 207–213
Bond JS. J Biol Chem, 2019, 294: 1643–1651
Lee M, Fridman R, Mobashery S. Chem Soc Rev, 2004, 33: 401–409
Vizovisek M, Vidmar R, Drag M, Fonovic M, Salvesen GS, Turk B. Trends Biochem Sci, 2018, 43: 829–844
Chang C, Werb Z. Trends Cell Biol, 2001, 11: S37–S43
Deryugina EI, Quigley JP. Cancer Metast Rev, 2006, 25: 9–34
Wang X, Khalil R A. Adv Pharmacol, 2018, 81: 241–330
Roy R, Yang J, Moses MA. JCO, 2009, 27: 5287–5297
Qin Y, Yin Y, Liu YK, Feng JJ, Jin LF, Pu Y, Hua D, Qi XW. Cell Biochem Biophys, 2015, 73: 253–259
Martínez D, Munera M, Cantillo JF, Wortmann J, Zakzuk J, Keller W, Caraballo L, Puerta L. Int J Mol Sci, 2019, 20: 3025
Mori S, Pawankar R, Ozu C, Nonaka M, Yagi T, Okubo K. Allergy Asthma Immunol Res, 2012, 4: 231–239
Cheng H, Li SY, Zheng HR, Li CX, Xie BR, Chen KW, Li B, Zhang XZ. Anal Chem, 2017, 89: 4349–4354
Gao X, Guo W, Jiang L, Hu B, Liu X, Xu K, Tang B. Sci China Chem, 2020, 63: 135–140
Fischbach M, Resch-Genger U, Seitz O. Angew Chem Int Ed, 2014, 53: 11955–11959
Ma T, Hou Y, Zeng J, Liu C, Zhang P, Jing L, Shangguan D, Gao M. J Am Chem Soc, 2018, 140: 211–218
Fonfara I, Richter H, Bratovic M, Le Rhun A, Charpentier E. Nature, 2016, 532: 517–521
Gao P, Yang H, Rajashankar KR, Huang Z, Patel DJ. Cell Res, 2016, 26: 901–913
Kundert K, Lucas JE, Watters KE, Fellmann C, Ng AH, Heineike BM, Fitzsimmons CM, Oakes BL, Qu J, Prasad N, Rosenberg OS, Savage DF, El-Samad H, Doudna JA, Kortemme T. Nat Commun, 2019, 10: 2127
Liu F, Yang M, Song W, Luo X, Tang R, Duan Z, Kang W, Xie S, Liu Q, Lei C, Huang Y, Nie Z, Yao S. Chem Sci, 2020, 11: 2993–2998
Pu J, Chronis I, Ahn D, Dickinson BC. J Am Chem Soc, 2015, 137: 15996–15999
English MA, Soenksen LR, Gayet RV, de Puig H, Angenent-Mari NM, Mao AS, Nguyen PQ, Collins JJ. Science, 2019, 365: 780–785
Dong D, Ren K, Qiu X, Zheng J, Guo M, Guan X, Liu H, Li N, Zhang B, Yang D, Ma C, Wang S, Wu D, Ma Y, Fan S, Wang J, Gao N, Huang Z. Nature, 2016, 532: 522–526
Swarts DC, van der Oost J, Jinek M. Mol Cell, 2017, 66: 221–233.e4
Yamano T, Nishimasu H, Zetsche B, Hirano H, Slaymaker IM, Li Y, Fedorova I, Nakane T, Makarova KS, Koonin EV, Ishitani R, Zhang F, Nureki O. Cell, 2016, 165: 949–962
Sheen-Chen SM, Chen HS, Eng HL, Sheen CC, Chen WJ. Cancer Lett, 2001, 173: 79–82
Murnane MJ, Cai J, Shuja S, McAneny D, Klepeis V, Willett JB. Int J Cancer, 2009, 125: 2893–2902
Garbisa S, Scagliotti G, Masiero L, Francesco C D, Caenazzo C, Onisto M, Micela M, Stetler-Stevenson W G, Liotta L A. Cancer Res, 1992, 52: 4548–4549
Li L, Lin H, Lei C, Nie Z, Huang Y, Yao S. Biosens Bioelectron, 2014, 54: 42–47
Maragoudakis ME, Tsopanoglou NE, Andriopoulou P, Maragoudakis MEM. Matrix Biol, 2000, 19: 345–351
Acknowledgements
This work was supported by the National Natural Science Foundation of China (21974038, 21725503) and the Fundamental Research Funds for the Central Universities.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
The authors declare no conflict of interest.
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Yang, M., Shi, K., Liu, F. et al. Coupling of proteolysis-triggered transcription and CRISPR-Cas12a for ultrasensitive protease detection. Sci. China Chem. 64, 330–336 (2021). https://doi.org/10.1007/s11426-020-9863-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11426-020-9863-y