Abstract
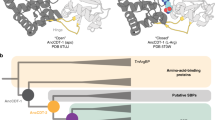
The origin of the catalytic power of enzymes with a meta-stable native state, e.g. molten globular state, is an unsolved challenging issue in biochemistry. To help understand the possible differences between this special class of enzymes and the typical ones, we report here computer simulations of the catalysis of both the well-folded wild-type and the molten globular mutant of chorismate mutase. Using the ab initio quantum mechanical/molecular mechanical minimum free-energy path method, we determined the height of reaction barriers that are in good agreement with experimental measurements. Enzyme-substrate interactions were analyzed in detail to identify factors contributing to catalysis. Computed angular order parameters of backbone N-H bonds and side-chain methyl groups suggested site-specific, non-uniform rigidity changes of the enzymes during catalysis. The change of conformational entropy from the ground state to the transition state revealed distinctly contrasting entropy/enthalpy compensations in the dimeric wild-type enzyme and its molten globular monomeric variant. A unique catalytic strategy was suggested for enzymes that are natively molten globules: some may possess large conformational flexibility to provide strong electrostatic interactions to stabilize the transition state of the substrate and compensate for the entropy loss in the transition state. The equilibrium conformational dynamics in the reactant state were analyzed to quantify their contributions to the structural transitions enzymes needed to reach the transition states. The results suggest that large-scale conformational dynamics make important catalytic contributions to sampling conformational regions in favor of binding the transition state of substrate.
Similar content being viewed by others
References
Wolfenden R. Degrees of difficulty of water-consuming reactions in the absence of enzymes. Chem Rev, 2006, 106(8): 3379–3396
Kamerlin SCL, Warshel A. At the dawn of the 21st century: Is dynamics the missing link for understanding enzyme catalysis? Proteins: Struct Func & Bioinfo, 2010, 78(6): 1339–1375
Hammes-Schiffer S, Benkovic SJ. Relating protein motion to catalysis. Annu Rev Biochem, 2006, 75: 519–541
Benkovic SJ, Hammes GG, Hammes-Schiffer S. Free-energy landscape of enzyme catalysis. Biochemistry, 2008, 47(11): 3317–3321
Watt ED, Shimada H, Kovrigin EL, Loria JP. The mechanism of rate-limiting motions in enzyme function. Proc Natl Acad Sci USA, 2007, 104(29): 11981–11986
Warshel A. Computer simulations of enzyme catalysis: Methods, progress, and insights. Annu Rev Biophys Biomol Struct, 2003, 32: 425–443
Warshel A, Sharma PK, Kato M. Electrostatic basis for enzyme catalysis. Chem Rev, 2006, 106(8): 3210–3235
Kamerlin SCL, Warshel A. Multiscale modeling of biological functions. Phys Chem Chem Phys, 2011, 13(22): 10401–10411
Gao JL, Truhlar DG. Quantum mechanical methods for enzyme kinetics. Annu Rev Phys Chem, 2002, 53: 467–505
Hu H, Yang WT. Free energies of chemical reactions in solution and in enzymes with ab initio quantum mechanics/molecular mechanics methods. Annu Rev Phys Chem, 2008, 59: 573–601
Garcia-Viloca M, Gao J, Karplus M, Truhlar DG. How enzymes work: Analysis by modern rate theory and computer simulations. Science, 2004, 303(5655): 186–195
Senn HM, Thiel W. Qm/mm methods for biomolecular systems. Angew Chem Int Ed, 2009, 48(7): 1198–1229
Nam K, Prat-Resina X, Garcia-Viloca M, Devi-Kesavan LS, Gao J. Dynamics of an enzymatic substitution reaction in haloalkane dehalogenase. J Am Chem Soc, 2004, 126(5): 1369–1376
Gao JL. Catalysis by enzyme conformational change as illustrated by orotidine 5′-monophosphate decarboxylase. Curr Opinion Struct Biol, 2003, 13(2): 184–192
Benkovic SJ, Hammes-Schiffer S. Biochemistry-enzyme motions inside and out. Science, 2006, 312(5771): 208–209
Villa J, Strajbl M, Glennon TM, Sham YY, Chu ZT, Warshel A. How important are entropic contributions to enzyme catalysis? Proc Natl Acad Sci USA, 2000, 97(22): 11899–11904
Pisliakov AV, Cao J, Kamerlin SCL, Warshel A. Enzyme millisecond conformational dynamics do not catalyze the chemical step. Proc Natl Acad Sci, 2009, 106(41): 17359–17364
Adamczyk AJ, Cao J, Kamerlin SCL, Warshel Arieh. Catalysis by dihydrofolate reductase and other enzymes arises from electrostatic preorganization, not conformational motions. Proc Natl Acad Sci USA, 2011, 108(34): 14115–14120
Olsson MHM, Parson WW, Warshel A. Dynamical contributions to enzyme catalysis: Critical tests of a popular hypothesis. Chem Rev, 2006, 106(5): 1737–1756
Roca M, Messer B, Hilvert D, Warshel A. On the relationship between folding and chemical landscapes in enzyme catalysis. Proc Natl Acad Sci USA, 2008, 105(37): 13877–13882
Wolfenden R, Snider MJ. The depth of chemical time and the power of enzymes as catalysts. Acc Chem Res, 2001, 34(12): 938–945
Cooper A, Dryden DTF. Allostery without conformational change-A plausible model. Eur Biophys J, 1984, 11(2): 103–109
Ota N, Agard DA. Intramolecular signaling pathways revealed by modeling anisotropic thermal diffusion. J Mol Biol, 2005, 351(2): 345–354
Szefczyk B, Mulholland AJ, Ranaghan KE, Ranaghan, Sokalski WA Differential transition-state stabilization in enzyme catalysis: Quantum chemical analysis of interactions in the chorismate mutase reaction and prediction of the optimal catalytic field. J Am Chem Soc, 2004, 126(49): 16148–16159
Strajbl M, Shurki A, Kato M, Warshel A. Apparent nac effect in chorismate mutase reflects electrostatic transition state stabilization. J Am Chem Soc, 2003, 125(34): 10228–10237
Hur S, Bruice TC. Just a near attack conformer for catalysis (chorismate to prephenate rearrangements in water, antibody, enzy mes, and their mutants). J Am Chem Soc, 2003, 125(35): 10540–10542
Guo H, Cui Q, Lipscomb WN, Karplus M. Substrate conformational transitions in the active site of chorismate mutase: Their role in the catalytic mechanism. Proc Natl Acad Sci USA, 2001, 98(16): 9032–9037
Marti S, Andres J, Moliner V, Silla E, Tuñón I, Bertrán J, Field MJ. A hybrid potential reaction path and free energy study of the chorismate mutase reaction. J Am Chem Soc, 2001, 123(8): 1709–1712
Vamvaca K, Vogeli B, Kast P, Pervushin K, Hilvert D. An enzymatic molten globule: Efficient coupling of folding and catalysis. Proc Natl Acad Sci USA, 2004, 101(35): 12860–12864
Pervushin K, Vamvaca K, Vogeli B, Hilvert D. Structure and dynamics of a molten globular enzyme. Nat Struct Mol Biol, 2007, 14: 1202–1206
Vamvaca K, Jelesarov I, Hilvert D. Kinetics and thermodynamics of ligand binding to a molten globular enzyme and its native counterpart. J Mol Biol, 2008, 382(4): 971–977
Woycechowsky KJ, Choutko A, Vamvaca K, Hilvert D. Relative tolerance of an enzymatic molten globule and its thermostable counterpart to point mutation. Biochemistry, 2008, 47(51): 13489–13496
Roca M, Vardi-Kilshtain A, Warshel A. Toward accurate screening in computer-aided enzyme design. Biochemistry, 2009, 48(14): 3046–3056
Hu H, Lu ZY, Yang WT. QM/MM minimum free-energy path: Methodology and application to triosephosphate isomerase. J Chem Theory Comput, 2007, 3(2): 390–406
Hu H, Lu ZY, Parks JM, Burger SK, Yang W. Quantum mechanics/ molecular mechanics minimum free-energy path for accurate reaction energetics in solution and enzymes: Sequential sampling and optimization on the potential of mean force surface. J Chem Phys, 2008, 128(3): 034105
Lee AY, Karplus PA, Ganem B, Clardy Jon. Atomic-structure of the buried catalytic pocket of escherichia-coli chorismate mutase. J Am Chem Soc, 199 Jon, (12): 3627–3628
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML. Comparison of simple potential functions for simulating liquid water. J Chem Phys, 1983, 79(2): 926–935
Essmann U, Perera L, Berkowitz M, Darden Tom, Lee H and Pedersen LG. J Chem Phys, 1995, 103: 8577–8593
MacKerell Jr. AD, Bashford D, Dunbrack Jr RL, Evanseck JD, Field MJ, Fischer S, Gao J, Guo H, Ha S, Joseph-McCarthy D,. Kuchnir L, Kuczera K, Lau FTK, Mattos C, Michnick S, Ngo T, Nguyen D T, Prodhom B, Reiher III WE, Roux B, Schlenkrich M, Smith JC, Stote R, Straub J, Watanabe M, Wiórkiewicz-Kuczera J, Yin D, Karplus M. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J Phys Chem B, 1998, 102: 3586–3616
Berendsen HJC, Postma JPM, van Gunsteren WF, DiNola A, Haak JR. Molecular dynamics with coupling to an external bath. J Chem Phys, 1984, 81: 3684–3690
Hu H, Yang WT. Development and application of ab initio qm/mm methods for mechanistic simulation of reactions in solution and in enzymes. J Mol Struct-Theochem, 2009, 898(1-3): 17–30
Ayala PY, Schlegel HB. A combined method for determining reaction paths, minima, and transition state geometries. J Chem Phys, 1997, 107(2): 375–384
Frisch MJTGW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Montgomery Jr JA, Vreven T, Kudin KN, Burant JC, Millam JM, Iyengar SS. Tomasi J. Barone V, Mennucci B, Cossi M, Scalmani G, Rega N, Petersson GAm, Nakatsuji H, Hada M, Ehara, M, Toyota K, Fukuda R.; Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Klene M, Li X, Knox JE, Hratchian HP, Cross JB, Bakken V, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski, JW, Ayala PY, Morokuma K, Voth, G. A, Salvador, P, Dannenberg, J. J, Zakrzewski, V. G, Dapprich, S, Daniels, A. D, Strain, M. C, Farkas O, Malick DK, Rabuck AD, Raghavachari K, Foresman JB, Ortiz J V, Cui Q, Baboul AG, Clifford S, Cioslowski J, Stefanov BB, Liu G, Liashenko A, Piskorz P, Komaromi I, Martin RL, Fox DJ, Keith T, Al-Laham MA, Peng CY, Nanayakkara A, Challacombe M, Gill PMW, Johnson B, Chen W, Wong MW, Gonzalez C, Pople JA. Gaussian 03. Secondary. 2004
Choy WY, Shortle D, Kay LE. Side chain dynamics in unfolded protein states: An nmr based 2h spin relaxation study of d131d. J Am Chem Soc, 2003, 125(7): 1748–1758
Eisenmesser EZ, Millet O, Labeikovsky W, Korzhnev DM, Wolf-Watz M, Bosco DA, Skalicky JJ, Kay LE, Kern D. Intrinsic dynamics of an enzyme underlies catalysis. Nature, 2005, 438(7064): 117–121
Lee AL, Kinnear SA, Wand AJ. Redistribution and loss of side chain entropy upon formation of a calmodulin-peptide complex. Nat Struct Biol, 2000, 7(1): 72–77
Lee AL, Wand AJ. Microscopic origins of entropy, heat capacity and the glass transition in proteins. Nature, 2001, 411(6836): 501–504
Fuentes EJ, Der CJ, Lee AL. Ligand-dependent dynamics and intramolecular signaling in a pdz domain. J Mol Biol, 2004, 335(4): 1105–1115
Hu H, Clarkson MW, Hermans J, Lee AL. Increased rigidity of eglinc at acidic pH: Evidence from nmr spin relaxation and md simulations. Biochemistry, 2003, 42(47): 13856–13868
Karplus M, Kushick JN. Methods for estimating the configurational entropy of macromolecules. Macromolecules, 1981, 14(2): 325–332
Andricioaei I, Karplus M. On the calculation of entropy from covariance matrices of the atomic fluctuations. J Chem Phys, 2001, 115(14): 6289–6292
Fan Y, Cembran A, Ma S, Gao J. Connecting protein conformational dynamics with catalytic function as illustrated in dihydrofolate reductase. Biochemistry, 2013, 52(12): 2036–2049
Strater N, Hakansson K, Schnappauf G, Braus G, Lipscomb WN. Crystal structure of the T state of allosteric yeast chorismate mutase and comparison with the R state. Proc Natl Acad Sci USA, 1996, 93(8): 3330–3334
Strajbl M, Sham YY, Villa J, Chu ZT, Warshel A. Calculations of activation entropies of chemical reactions in solution. J Phys Chem B, 2000, 104(18): 4578–4584
Shurki A, Strajbl M, Villa J, Warshel A. How much do enzymes really gain by restraining their reacting fragments? J Am Chem Soc, 2002, 124(15): 4097–4107
Galopin CC, Zhang S, Wilson DB, Bruce Ganem. On the mechanism of chorismate mutases: Clues from wild-type E-coli enzyme and a site-directed mutant related to yeast chorismate mutase. Tetra Lett, 1996, 37(48): 8675–8678
Galopin CC, Zhang S, Wilson DB, Bruce Ganem. On the mechanism of chorismate mutases: Clues from wild-type E-coli enzyme and a site-directed mutant related to yeast chorismate mutase (vol 37, pg 8675, 1996). Tetra Lett, 1997, 38(9): 1467–1467
Warshel A. Electrostatic origin of the catalytic power of enzymes and the role of preorganized active sites. J Biol Chem, 1998, 273(42): 27035–27038
Li L, Weinreb V, Francklyn C, Carter CWJ. Histidyl-trna synthetase urzymes class i and ii aminoacyl tRNA synthetase urzymes have comparable catalytic activities for cognate amino acid activation. J Biol Chem, 2011, 286(12): 10387–10395
Pham Y, Kuhlman B, Butterfoss GL, Hu H, Weinreb V, Carter CW. Tryptophanyl-trna synthetase urzyme: A model to recapitulate molecular evolution and investigate intramolecular complementation. J Biol Chem, 2010, 285(49): 38590–38601
Daggett V, Levitt M. A model of the molten globule state from molecular-dynamics simulations. Proc Natl Acad Sci USA, 1992, 89(11): 5142–5146
Mark AE, Van Gunsteren WF. Simulation of the thermal-denaturation of hen egg-white lysozyme-trapping the molten globule state. Biochemistry, 1992, 31(34): 7745–7748
Zhang S, Pohnert G, Kongsaeree P, Wilson DB, Clardy J, Ganem B. Chorismate mutase-prephenate dehydratase from escherichia colistudy of catalytic and regulatory domains using genetically engineered proteins. J Biol Chem, 1998, 273(11): 6248–6253
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Hu, H. Wild-type and molten globular chorismate mutase achieve comparable catalytic rates using very different enthalpy/entropy compensations. Sci. China Chem. 57, 156–164 (2014). https://doi.org/10.1007/s11426-013-5021-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11426-013-5021-7