Abstract
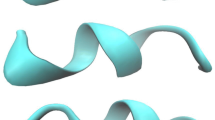
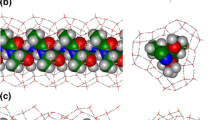
We have performed molecular dynamics simulations on the reversible folding/unfolding of small α-helix (short Ala based peptide Ala5) in explicit water solvent in terms of ABEEMσπ/MM. A dynamics analysis shows that the α-helical turn can be preserved up to a period of about 2 ns at 300 K, which supports the conclusions of Margulis et al. The time trajectory of the root mean square deviation between the heavy atoms of the backbone and the helical reference structure indicate that “helix melting and formation occurs rapidly on a time scale of 0.1 ns at 300 K” is not a felicitous conclusion. We first quantificationally concluded that the helix nucleation can maintain 2 ns, 1–1.5 ns and 0.8 ns for Ala5 at 300 K, 400 K and 500 K, respectively. Furthermore, increasing temperature dose not alter the pathway of folding/unfolding, but change the rate. An analysis of structures in a “transition-state ensemble” shows that helix-to-coil transitions occurs predominantly through breaking of hydrogen bonds at the helix ends (92%), particularly at the C-terminus(50%). Hydrogen bonds’ breaking and formation occurs on a time scale of 0.1 ns.
Similar content being viewed by others
References
Zhou R. Trp-cage. Folding free energy landscape in explicit water. Proc Natl Acad Sci USA 2003, 100: 13280–13285
Jiao Y, Yang P. Metal-amyloid-β peptide interactions: a preliminary investigation of molecular mechanisms for Alzheimer’s disease. Sci China Ser B-Chem, 2007, 50: 453–467
Jiao Y, Han D X, Yang P. Molecular modeling of the inhibitory mechanism of copper(II) on aggregation of amyloid β-peptide. Sci China Ser B-Chem, 2005, 48: 580–590
Snow C D, Nguyen H, Pande V S, Gruebele M. Absolute comparison of simulated and experimental protein-folding dynamics. Nature, 2002, 420: 102–106
Daggett V. Protein Folding Simulation. Chem Rev, 2006, 106: 1898–1916
Hummer G, Garcia A E, Garde S. Helix nucleation kinetics from molecular simulations in explicit solvent. PROTEINS: Struc, Funct, Genet, 2001, 42: 77–84
Brooks C L. Helix Coil Kinetics: Folding time III scales for helical peptides from a sequential kinetic model. J Phys Chem, 1996, 100: 2546–2549
Elmer S, Pande V S. A new twist on the helix-coil transition: A nonbiological helix with protein-like intermediates and traps. J Phys Chem B, 2001, 105: 482–485
Buchete N V, Straub J E. Mean first-passage time calculations for the coil-to-helix transition: The active helix ising model. J Phys Chem B, 2001, 105: 6684–6697
Shea J E, Brooks C L. From folding theories to folding proteins: A review and assessment of simulation studies of protein folding and unfolding. Annu Rev Phys Chem, 2001, 52: 499–535
Thompson P A, Eaton W A, Hofrichter J. Laser temperature jump study of the helix coil kinetics of an alanine peptide interpreted with a ‘Kinetic Zipper’ model. Biochemistry, 1997, 36: 9200–9210
Thompson P A, Muñoz V, Jas G S, Henry E R, Eaton W A, Hofrichter J. The helix-coil kinetics of a heteropeptide. J Phys Chem B, 2000, 104: 378–389
William S, Causgrove T P, Gilmanshin R, Fang K S, Callender R H, Woodruff W H, Dyer R B. Fast events in protein folding: helix melting and formation in a small peptide. Biochemistry, 1996, 35: 691–697
Lednev I K, Karnoup A S, Sparrow M C, Asher S A. Transient UV Raman spectroscopy finds no crossing barrier between the peptide α-helix and fully random coil conformation. J Am Chem Soc, 2001, 123: 2388–2392
Werner J H, Dyer R B, Fesinmeyer R M, Andersen N H. Dynamics of the primary processes of protein folding: Helix nucleation. J Phys Chem B, 2002, 106: 487–494
Jas G S, Eaton W A, Hofrichter J. Effect of viscosity on the kinetics of α-helix and β-hairpin formation. J Phys Chem B, 2001, 105: 261–272
Huang C, Klemke J W, Getahum Z, DeGrado W F, Gai F. Temperature-dependent helix coil transition of an alanine based peptide. J Am Chem Soc, 2001, 123: 9235–9238
Clarke D T, Doig A J, Stapley B J, Jones G R. The a-helix folds on the millisecond time scale. Proc Natl Acad Sci USA, 1999, 96: 7232–7237
Padmanabhan S, Marqusee S, Ridgeway T, Laue T L, Baldwin R L. Relative helix-forming tendencies of nonpolar amino acids. Nature, 1990, 344: 268–270
Padmanabhan S, Baldwin R L. Helix-stabilizing interaction between tyrosine and leucine or valine when the Spacing is i, i+4. J Mol Biol, 1994, 3: 706–713
Baldwin R L. α-Helix formation by peptides of defined sequence. Biophys Chem, 1995, 55: 127–135
Rohl C A, Fiori R L, Baldwin R L. Alanine is helix-stabilizing in both template-nucleated and standard peptide helices. Proc Natl Acad Sci USA, 1999, 96: 3682–3687
Brasseur R. Simulating the folding of small proteins by use of the local minimum energy and the free solvation energy yields native-like structures. J Mol Graph, 1995, 13: 312–322
Daggett V, Levitt M. Molecular dynamics simulations of helix denaturation. J Mol Biol, 1992, 223: 1121–1138
Margulis C J, Stern H A, Berne B J. Helix unfolding and intramolecular hydrogen bond dynamics in small α-helices in explicit solvent. J Phys Chem B, 2002, 106: 10748–10752
Hansmann U H, Masuya M, Okamoto Y. Characteristic temperatures of folding of a small peptide. Proc Natl Acad Sci USA, 1997, 94: 10652–10656
Nakazawa T, Okamoto Y. Electrostatic effects on the [alpha]-helix and [beta]-strand formation of BPTI(16–36) studied by Monte Carlo simulated annealing. J Pept Res, 1999, 54: 230–236
Hansmann U H, Okamoto Y, Onuchic J N. The folding funnel landscape for the peptide met-enkephalin. Proteins, 1999, 34: 472–483
Jacchieri S G, Richards N G J. Probing the influence of sequence-dependent interactions upon -helix stability in alanine-based linear peptides. Biopolymers, 1993, 33: 971–984
Aleman C, Roca R, Luque F J, Orozco M. Helical preferences of alanine, glycine, and aminoisobutyric homopeptides. Proteins, 1997, 28: 83–93
Sung S S. Folding simulations of alanine-based peptides with lysine residues. Biophys J, 1995, 68: 826–834
Klein C T, Mayer B, Köhler G, Wolschann P. Influence of solvation on helix formation of poly-alanine studied by multiple annealing simulations. J Mol Struct (Theochem), 1996, 370: 33–43
Klein C T, Mayer B, Köhler G, Wolschann P. Systematic stepsize variation: Efficient method for searching conformational space of polypeptides. J Comput Chem, 1998, 19: 1470–1481
Sham Y Y, Ma B, Tsai C J, Nussinov R. Thermal unfolding molecular dynamics simulation of Escherichia coli. Proteins, 2002, 46: 308–320
Pande V S, Rokhsar D S. Molecular dynamics simulations of unfolding and refolding of a beta-hairpin fragment of protein G. Proc Natl Acad Sci USA, 1999, 96: 9062–9067
Wong K B, Clarke J, Bond C J, Neira J I, Freund S. Towards a complete description of the structural and dynamic properties of the denatured state of barnase and the role of residual structure in folding. J Mol Biol, 2000, 296: 1257–1282
Ma B, Nussinov R. Molecular dynamics simulations of the unfolding of beta(2)-microglobulin. Protein Eng, 2003, 16: 561–575
Jemth P, Gianni S, Day R, Li B, Johnson C M, Daggett V, Fersht A R. Demonstration of a low-energy on-pathway intermediate in a fast-folding protein by kinetics, protein engineering, and simulation. Proc Natl Acad Sci USA, 2004, 101: 6450–6455
Day R, Bennion B J, Ham S, Daggett V. Increasing temperature accelerates protein unfolding without changing the pathway of unfolding. J Mol Biol, 2002, 322: 189–203
Day R, Daggett V. Ensemble versus single-molecule protein unfolding. Proc Natl Acad Sci USA, 2005, 102: 13445–13450
Petrovich M, Jonsson A L, Ferguson N, Daggett V, Fersht A R. Phi-analysis at the experimental limits: mechanism of beta-hairpin formation. J Mol Biol, 2006, 360: 865–881
Beck D A C, Daggett V. Methods for molecular dynamics simulations of protein folding/unfolding. Methods, 2004, 34: 112–120
Wang T, Wade R C. On the use of elevated temperature in simulations to study protein unfolding mechanisms. J Chem Theory Comput, 2007, 3(4): 1476–1483
Wu Y, Yang Z Z. Atom-bond electronegativity equalization method fused into molecular mechanics. II. A seven-site fluctuating charge and flexible body water potential function for liquid water. J Phys Chem A, 2004, 108: 7563–7576
Yang Z Z, Wang C S. Atom-bond electronegativity equalization method. 1. Calculation of the charge distribution in large molecules. J Phys Chem A, 1997, 101: 6315–6321
Yang Z Z, Wu Y, Zhao D X. Atom-bond electronegativity equalization method fused into molecular mechanics. I. A seven-site fluctuating charge and flexible body water potential function for water clusters. J Chem Phys, 2004, 120: 2541–2557
Yang Z Z, Zhang Q. Study of peptide conformation in terms of the ABEEM/MM method. J Comput Chem, 2006, 27: 1–10
Cong Y, Yang Z Z. General atom-bond electronegativity equalization method and its application in prediction of charge distributions in polypeptide. Chem Phys Lett, 2000, 316: 324–329
Cong Y, Yang Z Z, Wang C S, Liu X C, Bao X H. Investigation of the regio- and stereoselectivity of Diels-Alder reactions by newly developed ABEEMσπ model on the basis of local HSAB principle and maximum hardness principle. Chem Phys Lett, 2002, 357: 59–64
Li X, Yang Z Z. Study of lithium cation in water clusters: Based on atom-bond electronegativity equalization method fused into molecular mechanics. J Phys Chem A, 2005, 109: 4102–4111
Li X, Yang Z Z. Hydration of Li+-ion in atom-bond electronegativity equalization method-7P water: A molecular dynamics simulation study. J Chem Phys, 2005, 122: 084514
Guan Q M, Yang Z Z. Study on complexes of trypsin and its inhibitors by means of atom-bond electronegativity equalization method fused into molecular mechanics. J Theor Comput Chem, 2007, 6: 731–744
Yang Z Z, Cui B Q. Atomic charge calculation of metallobiomolecules in terms of the ABEEM method. J Chem Theory Comput, 2007, 3: 1561
Yang Z Z, Qian P. A study of N-methylacetamide in water clusters: Based on atom-bond electronegativity equalization method fused into molecular mechanics. J Chem Phys, 2006, 125: 064311
Liu C, Zhao D X, Yang Z Z. ABEEMσπ fluctuating charge force field applied to alanine dipeptide and alanine dipeptide-water systems. J Theor Comput Chem, 2009, in press.
Mayer B, Klein C T. Influence of solvation on the helix-forming tendency of nonpolar amino acids. J Mol Struct (Theochem), 2000, 532: 213–226
Kabsch W, Sander C. Dictionary of protein secondary structure: Pattern recognition of hydrogen-bonded and geometrical features. Biopolymers, 1983, 22: 2577–2637
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liu, C., Yang, Z. Reversible folding/unfolding of small a-helix in explicit solvent investigated by ABEEMσπ/MM. Sci. China Ser. B-Chem. 52, 1917–1924 (2009). https://doi.org/10.1007/s11426-009-0257-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11426-009-0257-y