Abstract
Influenza is an acute transmissible respiratory infectious disease in humans and animals with high morbidity and mortality. It was reported that luteolin, extracted from Chinese herbs, could potently inhibit influenza virus replication in vitro. To assess the effect and explore the fundamental mechanism of luteolin, we infected several cell lines with two subtypes of influenza A virus (IAV), including A/Jiangxi/312/2006 (H3N2) and A/Fort Monmouth/1/1947 (H1N1) and demonstrated that luteolin suppressed the replication of IAV by cytopathic effect reduction method, qRT-PCR, immunofluorescence and Western blot assays. A time-of-addition assay indicated that this compound interfered with viral replication at the early stage of infection. In addition, we found that luteolin suppressed coat protein I complex expression, which was related to influenza virus entry and endocytic pathway. Overall, our findings demonstrated the antiviral effect of luteolin against IAV and its novel antiviral mechanism.
Similar content being viewed by others
Introduction
Influenza can cause acute and highly transmissible infectious respiratory disease in humans and animals with high morbidity and mortality. In 1918, the Spanish influenza outbreak killed an estimated 50 million people worldwide [1]. The H1N1 influenza pandemic in 2009 and the emergency of H5N1, H7N9 influenza viruses had also raised significant public health concerning about the emergence of a potential highly pathogenic, pandemic influenza virus [2].
Two types of antiviral drugs were used in clinic: neuraminidase inhibitors (NAIs), such as oseltamivir, peramivir, and the M2 ion-channel blocker (amantadine and rimantadine). Although these drugs are effective against IAV replication, their efficacy can decrease rapidly once the emergence of drug-resistant viral mutants occurs [3, 4]. Hence, there is an urgent need to develop new antiviral drugs.
Influenza virus enters the cell by cell surface glycoproteins or glycolipid-mediated endocytosis [5]. Upon pH-mediated fusion, the virus is uncoated, and the vRNPs are released into the cytoplasm and subsequently transported into the nucleus to initiate virus replication and protein translation [6]. The progeny virions are produced by assembling viral proteins and vRNP complexes at the plasma membrane. Then, cell surface sialic acids are cleaved by viral NA to allow assembled virions to be released from the infected cell [7].
During viral infection, several important host proteins are involved in influenza virus trafficking. For example, Rab, coat protein I (COPI) complex, histone deacetylase 8 and cullin participate in the transport of influenza viruses among the endocytic bodies and from the endocytic to the perinuclear space [8,9,10]. Among them, COPI components contribute to influenza virus entry and endocytic pathway. Sun et al. found that influenza virus particles trafficking in early endosome-to-intermediate/late endosome exhibited defects when the COPI gene was depleted in the infected cells [11]. Moreover, the inhibitor [golgicide A(GCA)], which can disrupt the COPI complexes function, directly repressed influenza viral membrane protein expression and the production of progeny influenza virus [11]. These findings suggest that small molecules targeting COPI protein might be potential anti-influenza drugs.
Flavonoids extracted from plants possess multiple biological activities, including antiviral activities. Luteolin (3,4,5,7-tetrahydroxyflavone, LUT, PubChem CID: 5280445, Fig. 1a) is a natural flavonoid that has shown multiple biological benefits, including immune-regulation, anti-inflammation, antioxidant, anticancer, and antiviral activities [12,13,14]. Liu et al. reported that luteolin isolated from herbs inhibited the activities of NA in vitro, implying its potential as an anti-IAV agent [15]. In the study, we showed that luteolin had a potent antiviral activity against influenza virus in vitro. In addition, we demonstrated that luteolin inhibited influenza virus replication, at least in part, by targeting the COPI subunit, β-COP. Our findings not only determined the antiviral effect of luteolin in vitro, but also provided a novel antiviral mechanism.
Chemical structure of luteolin and the cytotoxicity of luteolin. a The chemical structure of luteolin. b The effect of luteolin on cell viability. MDCK, Vero and Calu-3 cells were seeded in 96-well plates. After 18 h, the cells were treated with various concentrations of luteolin. Cell viability was measured by CCK assay after 48 h of post-treatment. Values and error bars represent the average and standard deviations from three independent experiments
Materials and methods
Reagents and antibody
Luteolin was purchased from J&K Scientific (CAS:491-70-3, Beijing, China), and oseltamivir carboxylate (OC) was from Medchem Express (CAS: 187227-45-8, NJ, USA). Luteolin and OC were dissolved in DMSO (2 mM stock). Golgicide A was purchased from Selleck (CAS:1139889-93-2, Shanghai, China). Pitstop2-100 was obtained from Abcam Biochemicals (Cambridge, MA, USA). The following antibodies were used for Western blot: β-actin (1:5000) (Cell Signaling Technology), β-COP (1:1000) (abcam), and IAV nonstructural protein (NS1, 1:400) (Santa Cruz), respectively.
Cells and virus
Madin-Darby canine kidney (MDCK) cells and Vero cells (African green monkey kidney) were grown at 37 °C in minimum essential medium (MEM) supplemented with 10% fetal bovine serum and 1% penicillin–streptomycin (10,000 U/mL). Extra 1% MEM nonessential amino acids solution was added to MDCK cells.
Influenza virus A/Fort Monmouth/1/1947 (H1N1) was purchased from the America Type Culture Collection (ATCC, VR-97™). Influenza virus A/Jiangxi/312/2006 (H3N2) was kindly donated by Professor Yuelong Shu from the Institute for Viral Disease Control and Prevention, China Centers for Disease Control and Prevention, Beijing, China.
Cell viability test
To measure luteolin cytotoxicity, MDCK (2.5 × 104), Vero (3 × 104) and Calu-3 (2 × 104) cells were grown in 96-well plates. After being cultured for overnight, cells were incubated with various concentrations of luteolin that ranged from 3.75 to 240 μM for an additional 48 h. Cell viability was measured using a CCK-8 assay kit (TransGen, Beijing, China) following the manufacturer's instructions. The absorbance of each well was recorded at 450 nm using a Multiskan MK3 microplate reader.
Cytopathic effect (CPE) assays
For CPE assays, MDCK cells in 96-well plates were infected with influenza virus at an MOI of 0.001 for 2 h, and then incubated with a maintenance medium supplemented with or without luteolin. After 48 h, the CPE was recorded. Then, we calculated the 50% CPE inhibition concentration (IC50) values of luteolin. For the antiviral assay, MDCK and Vero cells were infected with two IAV strains. After 24 h, we collected cells and supernatants by freezing and thawing thrice, and then determined virus titer by TCID50 assay for 48 h.
Time-of-addition assay
A time-of-addition experiment was conducted as previously described with some modifications [16]. Briefly, MDCK cells were inoculated with 0.01 MOI of virus at 37 °C for 2 h. The media with or without luteolin 15 µM were added during the periods of – 2 to 8 h, 0–8 h, 2–8 h, 4–8 h and 6–8 h. After each incubation period, the collected cells were washed with PBS, and then determined the viral mRNA through qRT-PCR.
Hemagglutination inhibition (HI) assay
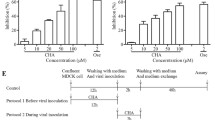
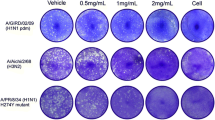
The effect of luteolin on the agglutination of chicken RBCs by influenza virus was determined by an HI assay [17]. Briefly, 50 µL of twofold dilutions of the compound in normal saline were prepared and mixed with 50 µL influenza virus solution (equivalent to 4 HA units) in a U-bottom 96-well plate. The plate was kept at 4 °C for 30 min, and then 100 µL 1.2% chicken erythrocyte suspension was added to each well. Lastly, cell agglutination in the well was checked visually after incubation for 40 min at room temperature.
Indirect immunofluorescence assay
MDCK cells were washed twice with PBS and incubated for 2 h with influenza virus (0.001 MOI) at 37 °C in 5% CO2 atmosphere for adsorption. After 2 h, inoculum was decanted and infected cells were supplemented with maintenance medium with or without luteolin. After 18 h, MDCK cells were fixed for 10 min with 4% paraformaldehyde at room temperature, and washed three times with PBS to remove the fixation buffer. The cells were permeabilized in 0.5% Triton X-100 in PBS for 15 min. The samples were then incubated with 3% BSA for 1 h at room temperature and further incubated with the mouse anti-influenza virus M2 antibody overnight at 4 °C. After washing three times with TBST, cells were incubated with Alexa Fluor 488 anti-mouse IgG (TransGen) for 1 h. The nucleus was stained with Hoechst 33342 (Beyotime) for 5 min. Photos were taken with an Olympus TH4-200 microscope.
For co-localization studies with β-COP and GM130, fixed Vero cells were permeabilized in 0.5% Triton X-100 for 15 min and blocked with 3% BSA for 1 h at room temperature. The cells were incubated with rabbit anti-β-COP and mouse anti-GM130 antibody overnight at 4°C, rinsed with TBST, stained with Alexa Fluor 488 anti-rabbit IgG (TransGen) and Alexa Fluor 594 anti-mouse IgG (Cell Signaling Technology) for 1 h. The nuclei were stained with DAPI (ZSBIO, Beijing). Images were acquired with a Zeiss LSM 710 microscope.
Western blotting
Protein extracts (10 µg) were subjected to electrophoresis with 10% polyacrylamide gel and were blotted onto PVDF membranes (Millipore Corp.). The membrane was incubated with primary antibody overnight at 4 °C, followed by incubation with HRP-conjugated rabbit anti-mouse or goat anti-rabbit IgG. The signal was detected with an ECL kit.
Quantitative real-time RT-PCR
For intracellular viral RNA quantification, total RNA was extracted from the cells using RNeasy Mini Kit (Qiagen) according to the manufacturer’s instructions. QRT-PCR analysis was performed as described as follows. Each qRT-PCR reaction mixture (15 μL) contained 7.5 μL of 2xTransStart Tip Green qPCR SuperMix, 0.3 μL of forward and reverse primers (10 μmol/L), 2 μL RNA Template, 0.3 μL of one step enzyme Mix, 0.3 μL of passive reference dye and 4.3 μL RNase-free Water (TransGen, Beijing, China). Oligonucleotide primer pairs were (1) forward 5′-GACCRATCCTGTCACCTCTGAC-3′ and reverse 5′-GGGCATTYTGGACAAAKCGTCTACG-3′ for influenza virus M2 (2) forward 5′-AGTCAAGGCTGAGAACGGGAAACT-3′ and reverse 5′-TCCACAACATACTCAGCACCAGCA-3′ for dog glyceraldehyd-3-phosphate dehydrogenase (GAPDH). Triplicate qRT-PCRs were prepared for each sample. The CT method was applied using the GAPDH cycle threshold for normalization to evaluate influenza virus M2 mRNA expression.
Statistical analysis
All data were analyzed by the SPSS 19.0 software using one-way ANOVA, and the two independent groups were analyzed via Student’s t test. Values and error bars represent the average and standard deviations from three independent experiments. Statistical differences were considered significant at *p < 0.05; and **p < 0.01.
Results
Cytotoxic and antiviral activity of luteolin in vitro
The cytotoxicity of luteolin against MDCK, Vero and Calu-3 cells were determined using a CCK assay kit (TransGen, Beijing, China). As shown in Fig. 1b, the CC50 (50% cytotoxic concentration of antiviral drugs) value of luteolin was 164.02 μM, 148.43 μM and greater than 240 μM in Vero, MDCK and Calu-3 cells, respectively.
The antiviral activity of luteolin was first measured by the CPE assay in MDCK cells [18]. Results indicated that luteolin efficiently inhibited the CPE induced by two IAV strains, A/Jiangxi/312/2006 (H3N2) and A/Fort Monmouth/1/1947 (H1N1). The IC50 values of luteolin were 6.89 μM and 7.15 μM, respectively. Coincidently, we observed a dose-dependent reduction in virus titers when the MDCK and Vero cells were treated with luteolin after infection (Fig. 2a–d). These results indicated that luteolin inhibited viral replication.
Luteolin dose-dependently inhibits the yield of IAV. a–d A/Jiangxi/312/2006 (H3N2) and A/Fort Monmouth/1/1947 (H1N1) infected MDCK (0.001 MOI) and Vero (0.01 MOI) cells, and luteolin was added after viral infection. OC serves as the positive control. At 24 h post infection, cells were frozen and thawed, and virus from the cells and supernatant was harvested. CPE reduction assay were quantified and the results were expressed as − log10 values of the mean viral load ± standard deviations. e, f MDCK cells in 12-well plates were infected with IAV (H1N1 and H3N2) at 0.001 MOI and mRNA was harvested at 12 h post infection. The viral M2 mRNA level was detected using qRT-PCR. *p < 0.05, **p < 0.01 statistically significant difference from the Con group. Values and error bars represent the average and standard deviations from three independent experiments
Viral mRNA was measured by qRT-PCR and the results showed that luteolin significantly reduced the IAV M2 mRNA level in MDCK cells in a dose-dependent manner (Fig. 2e, f). To further explore whether luteolin inhibits viral protein synthesis, the expression of viral M2 protein was also analyzed through an indirect immunofluorescence assay. As shown in Fig. 3a, b, luteolin exhibited a dose-dependent inhibition of M2 protein expression in MDCK cells. Additionally, positive drug OC considerably inhibited the replication of the influenza virus. Collectively, luteolin demonstrated a potent antiviral activity against influenza virus.
Luteolin dose-dependently inhibits the replication of IAV. MDCK cells were infected with influenza virus (0.001 MOI) for 2 h. MDCK cells were then treated with luteolin for 18 h. Then the cells were stained with an anti-M2 antibody and imaged by immunofluorescence microscopy. M2 staining is shown in green. Nuclei stained by Hoechst33342 are shown in blue. Photos were captured and merged using an Olympus TH4-200 microscope. Bars, 100 μm
Luteolin mainly inhibits replication of influenza virus at early stages of its lifecycle
To investigate which stage was involved in anti-influenza virus activity of luteolin, a time-of-addition assay was carried out. It had been reported that progeny viruses can be detected after 8 h inoculation of the influenza virus [16]. Results showed that luteolin had antiviral activity from − 2 to 4 h post infection by detecting M2 mRNA level using QRT-PCR assay (Fig. 4). This result indicated that luteolin primarily inhibited the influenza virus during the early stages of the life cycle. Luteolin showed low antiviral activity when applied at the late stage, with the inhibitory rate only about 20%. The results suggested that luteolin prevented virus replication, either by blocking virion absorption to the cell surface, or blocking the receptor-binding site of viral hemagglutinin, or blocking the fusion of the viral envelope with the endosome membrane.
Luteolin mainly stops the event of early stage in viral replication. MDCK cells infected with IAV (A/Fort Monmouth/1/1947) at 0.01 MOI were treated with 15 μM luteolin either at − 2 h, 0, 2, 4 or 6 h p.i., and virus M2 mRNA level were detected by qRT-PCR assay at 8 h p.i..*p < 0.05, **p < 0.01, statistically significant difference from the Con group. Values and error bars represent the average and standard deviations from three independent experiments
Luteolin affects the entry and internalization steps of influenza virus
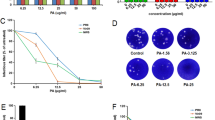
To identify whether luteolin can inhibit the entry and internalizaiton steps of influenza virus, we incubated with IAV at 4 °C for 1 h with luteolin and determined the mRNA level of the IAV. Result showed that the mRNA level of M2 with luteolin decreased 28.3% (Fig. 5a). Subsequently we tested the effect of luteolin on influenza virus internalization by quantifying the fraction of internalized IAV in the MDCK cells. Compared to the control cells, the suppression ratio of luteolin and pitstop2-100 were 20.4% and 56.8%, respectively (Fig. 5b). In addition, we performed a hemagglutination inhibition examination to assess the inhibition ability of luteolin for binding of hemagglutinin. The result showed that neither luteolin nor OC can inhibit hemagglutination by the influenza virus (Fig. 5c). Furthermore, we tested whether luteolin inactivated the infectivity of influenza virus directly. IAV and luteolin were incubated outside of cells and virion particles were then recovered by ultracentrifugation. Cells were next cultured with the virion and the result showed that incubation of the virus treated with luteolin did not reduce its infectivity in MDCK cells (Fig. 5d). Collectively, our results showed that luteolin did not target hemagglutination and influenza virus itself, but blocked influenza virus absorption and internalization, which in part contributed to the clarification that the luteolin inhibited the influenza virus during the early stages of infection.
Effect of luteolin on viral absorption and internalization. a MDCK cells were incubated with IAV (A/Fort Monmouth/1/1947, 0.01 MOI) at 4 °C for 1 h in the presence of DMSO or luteolin. The viral particles binding to the cell surface were measured by qRT-PCR. b The MDCK cells were incubated with IAV (A/Fort Monmouth/1/1947, 0.01 MOI) at 4 °C for 1 h. After incubation with fresh medium in the presence of DMSO, luteolin or pitstop2-100 (10 µM) for 30 min, the cells were washed with pH 3.0 PBS to remove the viral particles binding to the cell surface. The viral particles enter to the cell endosomes were measured by qRT-PCR. *p < 0.05, **p < 0.01, statistically significant difference from the Con group. Values and error bars represent the average and standard deviations from three independent experiments. c Effect of luteolin on hemagglutination inhibition. The concentration of luteolin in row 1 was 15 μM, whereas in the following rows is a serial of twofold dilution. d IAV stock (0.001 MOI) was treated with luteolin (15 μM) or DMSO control for 1 h at 4 °C. Group 1 was ultracentrifuged at 20,000×g for 2.5 h at 4 °C and the pellet was resuspended with maintenance medium; Group 2 was not centrifuged and served as control. MDCK cells were infected with above treated virus, the intracellular IAV NS1 protein was detected after 24 h
Luteolin inhibits IAV replication by down-regulating β-COP protein expression, but does not alter the localization of COPI
Considering that luteolin affected the entry and internalization steps of influenza virus and COPI complex is a critical host protein mediating influenza virus entry and endocytic pathway, we sought to determine whether luteolin disturbed the COPI during the replication of the influenza virus [19]. Viral protein NS1 and COPI were detected at 18 h after infection using the Western blot assay. Results showed that luteolin significantly inhibited influenza virus nonstructural protein NS1 and COPI expression in MDCK, Vero and Calu-3 cells, respectively, infected with A/Jiangxi/312/2006 (H3N2) and A/Fort Monmouth/1/1947 (H1N1) influenza virus (Fig. 6a–f).
Luteolin inhibits COPI expression. MDCK (0.001 MOI), Vero (0.01 MOI) and Calu-3 (0.01 MOI) cells were infected with H1N1 or H3N2 influenza virus for 2 h. The cell lysates were harvested for immunoblotting analysis. The protein levels of IAV NS1 and β-COP were analyzed by Western blot. The experiments were performed in triplicate, each value represents mean ± SD. *p < 0.05; **p < 0.01 versus Con group
In addition to the defect in viral entry and internalization upon inhibiting COPI protein expression, rapid disruption of COPI complex function with golgicide A (GCA) decreased viral membrane protein expression and infectious progeny virion production [11, 20]. Because luteolin could inhibit the expression of influenza virus M2 (data not shown) and progeny virions production, we further test whether luteolion treatment could affect the function of COPI complex. First, we investigated the effect of luteolin on β-COP in mock- and IAV-infected cells. GM130 is a resident Golgi protein, and β-COP, a component of COPI vesicles, is currently at high steady state concentrations in cis-Golgi membranes. In uninfected cells, GCA (10 μM) can disperse the distribution of GM130 and β-COP compared with untreated cells (Fig. 7a). However, luteolin treatment did not alter the intracellular distribution of GM130 and β-COP (Fig. 7a). The result in Fig. 7b shows that IAV can markedly disassemble the complex of GM130 and β-COP, with GM130 fluorescence showing a discrete punctate pattern and β-COP exhibiting a generally diffused pattern. After treatment of luteolin, the pattern of Golgi (GM130 and β-COP) staining was similar to that of uninfected cells. Collectively, our results showed that luteolin inhibits the expression of COPI protein without disturbing the localization of COPI, which may partly contribute to the reduction of the viral protein.
Luteolin did not disturb the assembly of the complex comprising GM130 and β-COP. a Vero cells were untreated or treated with GCA (10 μM) and luteolin (15 μM) for 1 h. b Vero cells infected with influenza virus at 0.01 MOI for 2 h, and then treated with luteolin for 18 h. The cells were stained with anti-GM130, and anti-β-COP antibody and were imaged by a Zeiss LSM 710 confocal microscope. GM130 staining is shown in red, whereas β-COP is shown in green. Nuclei were stained by DAPI are shown in blue. Bar, 10 μm
Discussion
Along with the wide use of IAV drugs in clinics, IAV strains resistant to available antiviral drugs have gradually led to the need for the development of novel anti-IAV agents [4, 21, 22]. Luteolin, a member of the flavonoid family, has been found to have multiple beneficial properties [12,13,14]. Moreover, a previous study revealed that luteolin had an inhibitory activity on influenza virus in vitro [23]. In this study, we found that luteolin inhibited the replication of two subtypes of influenza virus in MDCK, Vero and Calu-3 cells using varied methods including CPE reduction assay, qRT-PCR, Western blot and immunofluorescence assays. Our results indicated that luteolin exhibited an antiviral activity against IAV infection. The time-of-addition assay provided evidence that luteolin inhibited the whole process of influenza virus replication and played more important role at the early stages. Consistent with time-of-addition assay, we found that luteolin displayed 42.1% of inhibitory activity against the NA at a concentration of 20 μM (data not shown); however, the inhibition rate of luteolin on IAV replication is approximate 80%, which indicated that its antiviral effect was most likely due to inhibition of some other replication events of IAV. Furthermore, we determined that luteolin blocked influenza virus absorption and internalization, and the cumulative inhibition rate of absorption and internalization can reach 50% which was consistent with the time-of-addition result.
The COPI complex contains nine different COPI complex subunits. β-COP is one of the nine subunits of the COPI complex and is an essential component of the coatomer complex. The COPI complex was shown to be involved in transporting contents between the Golgi apparatus and the endoplasmic reticulum, and it has also been reported to mediate the influenza virus entry and endocytic pathway [24, 25]. That is to say, if these complexes were perturbed, the function of endosomal sorting, multivesicular body formation, and/or membrane trafficking will be missing [26,27,28]. Additionally, König et al. reported that vRNP nuclear import can be inhibited by knocking down δ-COP (ARCN1) expression [29]. The knockdown of β-COP by siRNA causes a significant reduction in the replication of the IAV [30]. Specifically, in COPI knockdown cells, viral infection was primarily blocked at the stage of virus internalization and early endosome-to-intermediate/late endosome trafficking [11].
In this study, we demonstrated that luteolin reduced β-COP protein expression, which contributed to the inhibition of the absorption and internalization of IAV. In mammalian cells, GBF1 (Golgi BFA resistance factor 1) can assist in the recruitment of COPI [31]. The GCA compound is the inhibitor of GBF1 function, resulting in rapid dissociation of COPI from Golgi membranes, and decomposition of the Golgi [20]. Furthermore, the influenza virus M2, NA and NP expression was decreased moderately in GCA-treated cells [11]. It showed that disturbing of COPI had anti-IAV activity. In this study, we found that luteolin did not disturb the function of COPI, but played a role of antiviral activity by inhibiting the expression of COPI protein.
Conclusions
We provided evidence that luteolin inhibited the replication of the IAV. We next confirmed that luteolin can interfere with virus at the early stages of its lifecycle and moderately block influenza virus absorption and internalization. Interestingly, we revealed that luteolin exhibited antiviral activity, at least in part, by targeting the host protein COPI with the reduction of its expression level. Our study showed that luteolin is a potential anti-IAV agent that can suppress IAV replication through host protein modulation, thereby providing new opportunities for designing anti-influenza virus drugs.
References
Webby RJ, Webster RG (2003) Are we ready for pandemic influenza? Science 302:1519–1522
Zhu H, Wang D, Kelvin DJ, Li L, Zheng Z, Yoon SW et al (2013) Infectivity, transmission, and pathology of human-isolated H7N9 influenza virus in ferrets and pigs. Science 341:183–186
Krug RM, Aramini JM (2009) Emerging antiviral targets for influenza A virus. Trends Pharmacol Sci 30:269–277
Moscona A (2009) Global transmission of oseltamivir-resistant influenza. N Engl J Med 360:953–956
Ohmit SE, Petrie JG, Cross RT, Johnson E, Monto AS (2011) Influenza hemagglutination-inhibition antibody titer as a correlate of vaccine-induced protection. J Infect Dis 204:1879–1885
Hutchinson E, Fodor E (2013) Transport of the influenza virus genome from nucleus to nucleus. Viruses 5:2424–2446
Liu G, Xiang Y, Guo C et al (2014) Cofilin-1 is involved in regulation of actin reorganization during influenza A virus assembly and budding. Biochem Biophys Res Commun 453:821–825
Müller KH, Kakkola L, Nagaraj AS, Cheltsov AV, Anastasina M, Kainov DE (2012) Emerging cellular targets for influenza antiviral agents. Trends Pharmacol Sci 33:89–99
Yamauchi Y, Boukari H, Banerjee I, Sbalzarini IF, Horvath P, Helenius A (2011) Histone deacetylase 8 is required for centrosome cohesion and influenza A virus entry. PLoS Pathog 7:e1002316
Huotari J, Meyer-Schaller N, Hubner M, Stauffera S, Kathedera N, Horvath P et al (2012) Cullin-3 regulates late endosome maturation. Proc Natl Acad Sci USA 109:823–828
Sun E, He J, Zhuang X (2013) Dissecting the role of COPI complexes in influenza virus infection. J Virol 87:2673–2685
Hytti M, Piippo N, Korhonen E, Honkakoski P, Kaarniranta K, Kauppinen A (2015) Fisetin and luteolin protect human retinal pigment epithelial cells fromoxidative stress- induced cell death and regulate inflammation. Sci Rep 5:17645
Bai L, Nong Y, Shi Y, Liu M, Yan L, Shang J et al (2016) Luteolin inhibits hepatitis B virus replication through extra cellular signal regulated kinase mediated down-regulation of hepatocyte nuclear factor 4α expression. Mol Pharm 13:568–577
Yan J, Wang Q, Zheng X, Sun H, Zhou Y, Li D et al (2012) Luteolin enhances TNF-related apoptosis-inducing ligand’s anticancer activity in a lung cancer xenograft mouse model. Biochem Biophys Res Commun 417:842–846
Liu AL, Liu B, Qin HL, Lee SM, Wang YT, Du GH (2008) Anti-influenza virus activities of flavonoids from the medicinal plant Elsholtzia rugulosa. Planta Med 74:847–851
Furuta Y, Takahashi K, Kuno MM, Sangawa H, Uehara S, Kozaki K et al (2005) Mechanism of action of T-705 against influenza virus. Antimicrob Agents Chemother 49:981–986
Kwon HJ, Kim HH, Yoon SY, Ryu YB, Chang JS, Cho KO et al (2010) In vitro inhibitory activity of Alpinia katsumadai extracts against influenza virus infection and hemagglutination. Virol J 7:307–315
Ma LL, Wang HQ, Wu P, Hu J, Yin JQ, Wu S et al (2016) Rupestonic acid derivative YZH-106 suppresses influenza virus replication by activation of heme oxygenase-1-mediated interferon response. Free Radic Biol Med 96:347–361
Aniento F, Gu F, Parton RG et al (1996) An endosomal beta COP is involved in the pH-dependent formation of transport vesicles destined for late endosomes. J Cell Biol 133:29–41
Sáenz JB, Sun WJ, Chang JW, Li J, Bursulaya B, Gray NS et al (2009) Golgicide A reveals essential roles for GBF1 in Golgi assembly and function. Nat Chem Biol 5:157–165
Wathen MW, Barro M, Bright RA (2013) Antivirals in seasonal and pandemic influenza—future perspectives. Influenza Other Respir Viruses 7:76–80
Lehnert R, Pletz M, Reuss A et al (2016) Antiviral medications in seasonal and pandemic influenza. Deutsches Arzteblatt Int 113:799
Sithisarn P, Michaelis M, Schubert-Zsilavecz M, Cinatl J Jr (2013) Differential antiviral and anti-inflammatory mechanisms of the flavonoids biochanin A and baicalein in H5N1 influenza A virus-infected cells. Antivir Res 97:41–48
Beck R, Rawet M, Wieland FT, Cassel D (2009) The COPI system: molecular mechanisms and function. FEBS Lett 5839:2701–2709
Popoff V, Adolf F, Brügger B, Wieland F (2011) COPI budding within the Golgi stack. Cold Spring Harb Perspect Biol 3:a005231
Gabriely G, Kama R, Gerst JE (2007) Involvement of specific COPI subunits in protein sorting from the late endosome to the vacuole in yeast. Mol Cell Biol 27:526–540
Gu F, Aniento F, Parton RG, Gruenberg J (1997) Functional dissection of COP-I subunits in the biogenesis of multivesicular endosomes. J Cell Biol 139:1183–1195
Cureton DK, Burdeinick-Kerr R, Whelan SPJ (2012) Genetic inactivation of COPI coatomer separately inhibits vesicular stomatitis virus entry and gene expression. J Virol 86:655–666
König R, Stertz S, Zhou Y, Inoue A, Hoffmann HH, Bhattacharyya S et al (2010) Human host factors required for influenza virus replication. Nature 463:813–817
Karlas A, Machuy N, Shin Y, Pleissner KP, Artarini A, Heuer D et al (2010) Genome-wide RNAi screen identifies human host factors crucial for influenza virus replication. Nature 463:818–822
Szul T, Grabski R, Lyons S et al (2007) Dissecting the role of the ARF guanine nucleotide exchange factor GBF1 in Golgi biogenesis and protein trafficking. J Cell Sci 120:3229–3240
Acknowledgements
This work was supported by the National Nature Science Foundation of China(NSFC) (Grant number: 81621064 and U1303224); The National Science and Technology Major Projects for "Major New Drugs Innovation and Development" (2018ZX09711003) and CAMS Innovation Fund for Medical Sciences (CIFMS) (No. 2017-I2M-3-010).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Yan, H., Ma, L., Wang, H. et al. Luteolin decreases the yield of influenza A virus in vitro by interfering with the coat protein I complex expression. J Nat Med 73, 487–496 (2019). https://doi.org/10.1007/s11418-019-01287-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11418-019-01287-7