Abstract
Purpose
Microorganisms have a multidimensional influence on plants, and they play pivotal roles in the soil-root interface mechanism. However, little is known about microorganisms for plants in the riparian environment regarding the endosphere, rhizosphere, and bulk soil. Therefore, this study assessed rhizodeposition, the structure of bacterial communities, and the assembly processes among the endosphere, rhizosphere, and bulk soil.
Methods
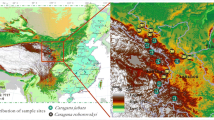
Salix matsudana, Taxodium distichum, Cynodon dactylon, and Hemarthria altissima were investigated in the riparian zone of the Ruxi River, China. We collected rhizospheres, bulk soil, and roots for these plants and analyzed the soil’s physico-chemical properties, enzyme activities, bacterial community complexity, and assembly processes.
Results
The results showed different abundant bacterial communities in the rhizosphere, bulk soils, and roots. Rhizodeposition of nitrogen and phosphorus had the greatest impact on bacterial diversity. Bacteria had more positive relationships in the endosphere than in soil compartments, indicating a relatively complex and stable community structure. The stochastic processes shape the microbial geographic variation of the endosphere, rhizosphere, and bulk soil for selected plants, with the proportional contribution in the rhizosphere and bulk soil being significantly less than that in the endosphere. Different microhabitats, via the assembly process, shape bacterial community structures.
Conclusions
We discovered that the bacterial community in the endosphere was less affected by environmental factors and more stable than the bacterial community in the soil compartment. This will help us find the best ways to keep soil and water safe in the newly formed riparian zone and understand the ecological evolution of microbes for vegetation restoration.









Similar content being viewed by others
References
An XC, Wang ZF, Teng XM, Zhou, RR, Wang XX, Xu M, Lian B (2022) Rhizosphere bacterial diversity and environmental function prediction of wild salt-tolerant plants in coastal silt soil. Ecol Indic 134:108503. https://doi.org/10.1016/j.ecolind.2021.108503
Arif M, Changxiao L (2022) Impacts of environmental literacy on ecological networks in the Three Gorges Reservoir, China. Ecol Indic 145:109571. https://doi.org/10.1016/j.ecolind.2022.109571
Arif M, Jiajia L, Dongdong D, Xinrui H, Qianwen G, Fan Y, Songlin Z, Changxiao L (2022) Effect of topographical features on hydrologically connected riparian landscapes across different land-use patterns in colossal dams and reservoirs. Sci Total Environ 851:158131. https://doi.org/10.1016/j.scitotenv.2022.158131
Asmaâ A, Etienne Y (2021) Engineering the plant microbiota in the context of the theory of ecological communities. Curr Opin Biotechnol 70:220–225. https://doi.org/10.1016/j.copbio.2021.06.009
Bárbara D, Liliana R, Eva L, Fernanda C, Cláudia P, Karl C (2021) Priority effects of stream eutrophication and assembly history on beta diversity across aquatic consumers, decomposers and producers. Sci Total Environ 797:149106. https://doi.org/10.1016/j.scitotenv.2021.149106
Barber SA (1995) Soil nutrient bioavailability: a mechanistic approach. John Wiley & Sons, New York
Bastian M, Heymann S, Jacomy M (2009) Gephi: an open source software for exploring and manipulating networks. In: International AAAI Conference on Weblog and Social Media: San Jose, California
Behzad H M, Muhammad A, Duan S, Kavousi A, Cao M, Liu J, Jiang Y (2022). Seasonal variations in water uptake and transpiration for plants in a karst critical zone in China. Sci Total Environ 160424. https://doi.org/10.1016/j.scitotenv.2022.160424
Chen J, Wu J, Liu M, Li L, Zhang W, Wang D, Ma T (2021) Bacterial community structure in the surface sediments of different habitats of Baiyangdian Lake, Northern China: effects of nutrient conditions. J Soils Sediments 21:1866–1874. https://doi.org/10.1007/s11368-021-02901-6
Chen Q, Yang F, Cheng XL (2022a) Effects of land use change type on soil microbial attributes and their controls: Data synthesis. Ecol Indic 138:108852. https://doi.org/10.1016/j.ecolind.2022a.108852
Chen Z, Song H, Muhammad A, Li C (2022b) Effects of hydrological regime on Taxodium ascendens plant decomposition and nutrient dynamics in the Three Gorges Reservoir riparian zone. Front Environ Sci 10:990485. https://doi.org/10.3389/fenvs.2022b.990485
Cong W, Yu J, Feng K, Deng Y, Zhang Y (2021) The coexistence relationship between plants and soil bacteria based on interdomain ecological network analysis. Front Microbiol 12:3664. https://doi.org/10.3389/fmicb.2021.745582
Ding D, Arif M, Liu M, Li J, Hu X, Geng Q, Yin F, Li C (2022) Plant-soil interactions and C:N:P stoichiometric homeostasis of plant organs in riparian plantation. Front Plant Sci 13:979023. https://doi.org/10.3389/fpls.2022.979023
Edwards J, Johnson C, Santos-Medellín C, Lurie E, Podishetty NK, Bhatnagar S, Bhatnagar S, Eisen A, J, Sundaresan V, (2015) Structure, variation, and assembly of the root-associated microbiomes of rice. Proc Natl Acad Sci USA 112:E911–E920. https://doi.org/10.1073/pnas.1414592112
Freilich M, Wieters E, Broitman B, Marquet P, Navarrete S (2018) Species co-occurrence networks: can they reveal trophic and non-trophic interactions in ecological communities? Ecology 99:690–699. https://doi.org/10.1002/ecy.2142
Gottel NR, Castro HF, Kerley M, Yang Z, Pelletier DA, Podar M, Karpinets T, Uberbacher E, Tuskan AG, Vilgalys R, Doktycz MJ, Schadt WC (2011) Distinct microbial communities within the endosphere and rhizosphere of Populus deltoides roots across contrasting soil types. Appl Environ Microbiol 77:5934–5944. https://doi.org/10.1128/aem.05255-11
Ha J, Gao Y, Zhang R, Ke Li, Zhang Y, Niu X, Xin C, Kai L, Yinhua C (2021) Diversity of the bacterial microbiome associated with the endosphere and rhizosphere of different cassava (Manihot esculenta Crantz) genotypes. Front Microbiol 12:2821. https://doi.org/10.3389/fmicb.2021.729022
Hale KRS, Valdovinos FS, Martinez ND (2021) Mutualism increases diversity, stability, and function of multiplex networks that integrate pollinators into food webs. Nat Commun 11:2182. https://doi.org/10.1038/s41467-020-15688-w
Hamza IS, Zouari AB, Akrout F, Keskes FA, Hamza A, Hassen M B (2021) Diversity and abundance of diazotrophic cyanobacteria in the central coastal area of the Gulf of Gabès (South-eastern Tunisia). Reg Stud Mar Sci 42:101653. https://doi.org/10.1016/j.rsma.2021.101653
Hira A, Muhammad A, Zarif N, Gul Z, Liu X, Cao Y (2022) Impacts of stressors on riparian health indicators in the upper and lower Indus River Basins in Pakistan. Int J Environ Res Public Health 19(20):13239. https://doi.org/10.3390/ijerph192013239
Hu X, Arif M, Ding D, Li J, He X, Li C (2022) Invasive plants and species richness impact litter decomposition in riparian zones. Front Plant Sci 13:955656. https://doi.org/10.3389/fpls.2022.955656
Jianbo W, Na L, Xiaodan W (2020) Nitrogen deposition strengthens the relationship between plants and the soil fungal community in alpine steppe, North Tibet. Appl Soil Ecol 147:103441. https://doi.org/10.1016/j.apsoil.2019.103441
Jin Z, Congcong J, Dayong Z, Rui H, Xinyi C, Zhongbo Y, Qinglong LW (2019) Patterns and assembly processes of planktonic and sedimentary bacterial community differ along a trophic gradient in freshwater lakes. Ecol Indic 106:105491. https://doi.org/10.1016/j.ecolind.2019.105491
Huot H, Joyner J, Córdoba A, Shaw RK, Wilson MA, Walker R, Muth TR, Cheng Z (2017) Characterizing urban soils in New York City: profile properties and bacterial communities. J Soils Sediments 17:393–407. https://doi.org/10.1007/s11368-016-1552-9
Khanna K, Kohli SK, Ohri P, Bhardwaj R (2021) Plants-nematodes-microbes crosstalk within soil: a trade-off among friends or foes. Microbiol Res 248:126755. https://doi.org/10.1016/j.micres.2021.126755
Kong W, Su F, Zhang Q, Ishii S, Michael JS, Banerjee S, Shao M, Qiu L, Wei X (2022) Erosion and deposition divergently affect the structure of soil bacterial communities and functionality. Catena 209:105805. https://doi.org/10.1016/j.catena.2021.105805
Kuang W, Li J, Zhang S, Long L (2015) Diversity and distribution of Actinobacteria associated with reef coral Porites lutea. Front Microbiol 6:1094. https://doi.org/10.3389/fmicb.2015.01094
Li X, Zhang Q, Ma J, Yang Y, Wang Y, Fu C (2020) Flooding irrigation weakens the molecular ecological network complexity of soil microbes during the process of dryland-to-paddy conversion. In J Environ Res Public Health 17:561. https://doi.org/10.3390/ijerph17020561
Liao L, Wang J, Lei S, Zhang L, Ye Z, Liu G, Zhang C (2022) Differential effects of nitrogen addition on the organic carbon fractions of rhizosphere and bulk soil based on a pot experiment. J Soils Sediments. https://doi.org/10.1007/s11368-022-03311-y
Liting D, Mingming Z, Ruixin B, Ayodej B, Ugochi UE, Jizhou Z, Shanshan L, Yanhui C, Yue H, Yu S, Xiuhong X (2021) Insight into the influence of biochar on nitrification based on multi-level and multi-aspect analyses of ammonia-oxidizing microorganisms during cattle manure composting. Bioresour Technol 339:125515. https://doi.org/10.1016/j.biortech.2021.125515
Liu C, Dong Y, Hou L, Deng N, Jiao R (2017) Acidobacteria community responses to nitrogen dose and form in Chinese fir plantations in southern China. Curr Microbiol 74:396–403. https://doi.org/10.1007/s00284-016-1192-8
Liu L, Kong J, Cui H, Zhang J, Wang F, Cai Z, Huang X (2016) Relationships of decomposability and C/N ratio in different types of organic matter with suppression of Fusarium oxysporum and microbial communities during reductive soil disinfestation. Biol Control 101:103–113. https://doi.org/10.1016/j.biocontrol.2016.06.011
Liu N, Hu H, Ma W, Deng Y, Wang Q, Luo A, Meng J, Feng X, Wang Z (2021) Relative importance of deterministic and stochastic processes on soil microbial community assembly in temperate grasslands. Microorganisms 9:1929. https://doi.org/10.3390/microorganisms9091929
Liyan Z, Jonathan MA, Marc D, Yuntao L, Yu S, Dan H (2019) Distinct methanotrophic communities exist in habitats with different soil water contents. Soil Biol Biochem 132:143–152. https://doi.org/10.1016/j.soilbio.2019.02.007
Maltseva IA, Maltsev YI (2021) Diversity of cyanobacteria and algae in dependence to forest-forming tree species and properties rocks of dump. Int J Environ Sci Technol 18:545–560. https://doi.org/10.1007/s13762-020-02868-w
Mapelli F, Vergani L, Terzagh E, Zecchin S, Raspa G, Marasco R, Rolli E, Zanardini E, Morosini C, Anelli S, Nastasio P, Sale VM, Armiraglio S, Di GA, Borin S (2022) Pollution and edaphic factors shape bacterial community structure and functionality in historically contaminated soils. Microbiol Res 263:127144. https://doi.org/10.1016/j.micres.2022.127144
Masaru N, Kohei O, Kyoko T, Yuichi A, Shinichi Y, Hisabumi T, Tsubasa S, Kazufumi Y, Akifumi S (2021) Tomato roots secrete tomatine to modulate the bacterial assemblage of the rhizosphere. Plant Physiol 186:270–284. https://doi.org/10.1093/plphys/kiab069
Muhammad A, Behzad HM, Tahir M, Li C (2022a) The impact of ecotourism on ecosystem functioning along main rivers and tributaries: implications for management and policy changes. J Environ Manage 320:115849. https://doi.org/10.1016/j.jenvman.2022a.115849
Muhammad A, Jiajia L, Tahir M, Jie Z, Changxiao L (2022b) Environmental literacy scenarios lead to land degradation and changes in riparian zones: implications for policy in China. Land Degrad Dev 1–17. https://doi.org/10.1002/ldr.4450
Ning D, Deng Y, Tiedje JM, Jizhong Z (2019) A general framework for quantitatively assessing ecological stochasticity. Proc Natl Acad Sci U S A 116:16892–16898. https://doi.org/10.1073/pnas
Promme J, Walker TWN, Wanek W, Braun J, Zezula D, Hu Y, Hofhansl F, Richter A (2020) Increased microbial growth, biomass, and turnover drive soil organic carbon accumulation at higher plant diversity. GCB Bioenergy 26:669–681. https://doi.org/10.1111/gcb.14777
R Development Core Team (2008) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria 3–900051–07–0. http://www.r-project.org
Ren Q, Song H, Yuan Z, Ni X, Li C (2018) Changes in soil enzyme activities and microbial biomass after revegetation in the Three Gorges Reservoir. China Forests 9:249. https://doi.org/10.3390/f9050249
Riley D, Barber SA (1970) Salt accumulation at the soybean (Glycine max. (L.) Merr.) root-soil interface. Soil Sci Soc Am J 34:154–155. https://doi.org/10.2136/sssaj1970.03615995003400010042x
Rosenberg E, Zilber-Rosenberg I (2018) The hologenome concept of evolution after 10 years. Microbiome 6:78. https://doi.org/10.1186/s40168-018-0457-9
Samuel SO, Suzuki K, Asiloglu R, Harada N (2021) Soil-root interface influences the assembly of the endophytic bacterial community in rice plants. Biol Fertil Soils 57. https://doi.org/10.1007/s00374-021-01611-y
Scharf BE, Hynes MF, Alexandre GM (2016) Chemotaxis signaling systems in model beneficial plant–bacteria associations. Plant Mol Biol 90:549–559. https://doi.org/10.1007/s11103-016-0432-4
Shakya M, Gottel N, Castro H, Yang ZK, Gunter L, Labbé J, Muchero W, Bonito G, Vilgalys R, Tuskan G, Podar M, Schad W (2013) A multifactor analysis of fungal and bacterial community structure in the root microbiome of mature Populus deltoides trees. PLoS ONE 8:e76382. https://doi.org/10.1371/journal.pone.0076382
Suzanne M, Fleishman HW, Bock DM, Eissenstat MC (2021) Undervine groundcover substantially increases shallow but not deep soil carbon in a temperate vineyard. Agric. Ecosyst Environ 313:107362. https://doi.org/10.1016/j.agee.2021.107362
Tripath BM, Stegen KM, Dong K, Adams J, Lee Y (2018) Soil pH mediates the balance between stochastic and de-terministic assembly of bacteria. ISME J 12:1072–1083. https://doi.org/10.1038/s41396-018-0082-4
Valdivia N, Garcés-V GI, Gómez I, Huovinen NN, Macaya EC, Pardo LM (2021) Beta diversity of Antarctic and Sub-Antarctic benthic communities reveals a major role of stochastic assembly processes. Front Mar Sci 8:1816. https://doi.org/10.3389/fmars.2021.780268
Wang H, Sun L (2018) Comparative metagenomic analysis of the microbial communities in the surroundings of Iheya north and Iheya ridge hydrothermal fields reveals insights into the survival strategy of microorganisms in deep-sea environments. J Mar Syst 180:102–111. https://doi.org/10.1016/j.jmarsys.2016.10.006
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microb 73(16):5261–5267. https://doi.org/10.1128/AEM.00062-07
Wang X, Li W, Xiao Y, Cheng A, Shen T, Zhu M, Yu L (2021) Abundance and diversity of carbon-fixing bacterial communities in karst wetland soil ecosystems. Catena 204:105418. https://doi.org/10.1016/j.catena.2021.105418
Wilkins L, Leray M, O'Dea A, Yuen B, Peixoto R, Pereira T, Bik H, Coil D, Duffy J, Herre E, Lessios H, Lucey N, Mejia L, Rashe D, Sharp K, Sogi E, Robert W, Thacker R, Wcisl W, Wilbank E, Eisen J (2019) Host-associated microbiomes drive structure and function of marine ecosystems. PLoS Biol 17:e3000533. https://doi.org/10.1371/journal.pbio.3000533
Wu H, Huang BH, Huang CL, Li G, Liao PC (2018) The aboveground vegetation type and underground soil property mediate the divergence of soil microbiomes and the biological interactions. Microb Ecol 75:434–446. https://doi.org/10.1007/s00248-017-1050-7
Xiao D, Xiao S, Ye Y, Zhang W, He X, Wang K (2019) Microbial biomass, metabolic functional diversity, and activity are affected differently by tillage disturbance and maize planting in a typical karst calcareous soil. J Soils Sediments 19:809–821. https://doi.org/10.1007/s11368-018-2101-5
Xu HW, Wang XK, Qu Q, Yang ZY, Wang MG, Liu GB, Xue S (2021a) Variations and factors characterizing ecological niches of species in a stable grassland plant community. Ecol Indic 128:107846. https://doi.org/10.1016/j.ecolind.2021.107846
Xu J, Gao W, Zhao B, Chen M, Ma L, Jia Z, Zhang J (2021b) Bacterial community composition and assembly along a natural sodicity/salinity gradient in surface and subsurface soils. Appl. Soil Ecol. 157, 103731. https://doi.org/10.1016/j.apsoil.2020.103731
Xu J, Zhang Y, Zhang P, Trivedi P, Riera N, Wang Y, Liu X, Fan G, Tang J, Helvécio D, Jaime C, Deng X, Ancona V, Lu Z, Zhong B, Roper M, Capote N, Catara V, Pietersen G, Vernière C, Abdullah M, Li L, Yang F, Xu X, Wang J, Yang H, Jin T, Wang N (2018) The structure and function of the global citrus rhizosphere microbiome. Nat Commun 9:4894. https://doi.org/10.1038/s41467-018-07343-2
Yan K, Dong Y, Gong Y, Zhu Q, Wang Y (2021) Climatic and edaphic factors affecting soil bacterial community biodiversity in different forests of China. Catena 207:105675. https://doi.org/10.1016/j.catena.2021.105675
Yang Z, Qing L, Qigen D, Yijun K (2019) Microbial mechanism underlying high and stable methane oxidation rates during mudflat reclamation with long-term rice cultivation: illumina high-throughput sequencing-based data analysis. J Hazard Mater 5:332–341. https://doi.org/10.1016/j.jhazmat.2019.03.032
Ye F, Ma MH, Wu SJ, Jiang Y, Zhu GB, Zhang H, Wang Y (2019) Soil properties and distribution in the riparian zone: the effects of fluctuations in water and anthropogenic disturbances. Eur J Soil Sci 70:664–673. https://doi.org/10.1111/ejss.12756
Yen NC, Daniel P (2022) Schachtman, Root exudates impact plant performance under abiotic stress. Trends Plant Sci 27:80–91. https://doi.org/10.1016/j.tplants.2021.08.003
Yuan H, Mei R, Liao J, Liu W (2019) Nexus of stochastic and deterministic processes on microbial community assembly in biological systems. Front Microbiol 10:1536. https://doi.org/10.3389/fmicb.2019.01536
Yuan MM, Guo X, Wu L, Zhang Y, Xiao N, Ning D, Shi Z, Zhou X, Wu L, Yang Y, Tiedje J, Zhou J (2021) Climate warming enhances microbial network complexity and stability. Nat Clim Change 11:343–348. https://doi.org/10.1038/s41558-021-00989-9
Yuancai Q, Arif M, Dong Z, Ting W, Qin Y, Bo P, Peng W, Wei H (2022) The effect of hydrological regimes on the concentrations of nonstructural carbohydrates and organic acids in the roots of Salix matsudana in the Three Gorges Reservoir, China. Ecol Indic 142:109176. https://doi.org/10.1016/j.ecolind.2022.109176
Zhu R, Liu J, Wang J, Wang J, Sun D (2020) Comparison of soil microbial community between reseeding grassland and natural grassland in Songnen Meadow. Sci Rep 10:16884. https://doi.org/10.1038/s41598-020-74023-x
Zuccarini P, Asensio D, Sardans J, Ogaya R, Peñuelas J (2021) Changes in soil enzymatic activity in a P-limited Mediterranean shrubland subject to experimental nitrogen deposition. Appl Soil Ecol 168:104159. https://doi.org/10.1016/j.apsoil.2021.104159
Funding
This research has been supported by the International Young Talents Program (No. QN2022168001L), the Chongqing Municipality Key Forestry Research Project (No. 2021–9), Chongqing Municipality Housing and Urban Construction Committee (No. Chengkezi 2019–1-4–2), Forestry Extension Project of China Central Finance (No. Yulinketui 2020–2), the Science Foundation of College of Life Sciences of Southwest University (No. 20212005406201), Ningxia Key Research and Development Project (No. 2020BFG03006), and Ningxia Natural Science Foundation Project (No. 2020AAC03107).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Responsible editor: Yuan Ge
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jiajia, L., Lijuan, L., Arif, M. et al. Newly formed riparian microhabitats simplify bacterial community structure and diversity. J Soils Sediments 23, 1927–1943 (2023). https://doi.org/10.1007/s11368-023-03454-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11368-023-03454-6