Abstract
Purpose
Reliable and effective techniques for removing contaminants from soil are highly desirable. However, metolachlor residue bioremediation and soil fertility improvement by Rhodospirillum rubrum (R. rubrum) in effluent after wastewater treatment have not yet been investigated. The aims of this study were to investigate the feasibility of bioremediation of metolachlor residues in soil and soil fertility improvement by R. rubrum in effluent and to explain the mechanism that R. rubrum in effluent was induced to express the regulatory gene.
Materials and methods
Soybean processing wastewater was obtained from Harbin Soybean Products Machining Factory. Soil samples were the surface soil (0–30 cm) from campus (1.77 g/kg total N, 4.15 g/kg total P, 1.58 g/kg total K, 17 g/kg SOM, 0.07 g/kg SMBC). Cytochrome P450 monooxygenase regulatory gene, MAPKKKs gene, was measured by RT-PCR.
Results and discussion
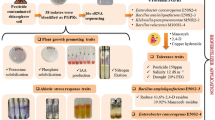
Compared to control treatment, metolachlor was removed efficiently and soil fertility was remediated by effluent containing R. rubrum. The removal in concentrations reached 2.97 mg/L (99%). Soil organic matter (SOM) and SMBC were enhanced 42 times. Molecular analysis revealed that metolachlor induced cpm gene expression to synthesize cytochrome P450 monooxygenase through activating MAPKKKs gene in MAPK signal transduction pathway.
Conclusions
Bioremediation of metolachlor in soil and improvement of soil fertility using R. rubrum in effluent were feasible. Metolachlor, as environmental pressure, induced cpm gene expression to synthesize cytochrome P450 monooxygenase and to remove metolachlor through activating MAPKKKs, MAPKKs, MAPKs genes in MAPK signal transduction pathway.



Similar content being viewed by others
Change history
13 December 2020
This article has been retracted. Please see the Retraction Notice for more detail: https://doi.org/10.1007/s11368-020-02853-3.
References
Boopathy R (2017) Anaerobic degradation of atrazine. Int Biodeterior Biodegrad 119:626–630
Carles L, Joly M, Bonnemoy F, Leremboure M, Donnadieu F, Batisson I, Besse-Hoggan P (2018) Biodegradation and toxicity of a maize herbicide mixture: mesotrione, nicosulfuron and S-metolachlor. J Hazardous Materials 354:42–53
El-Nahhal Y, Nir S, Polubesova T, Margulies L, Rubin B (1999) Movement of metolachlor in soil: effect of new organo-clay formulations. Pestic Sci 55:857–864
Fenga C, Gangemi S, Giambò F, Tsitsimpikou (2016) Low-dose occupational exposure to benzene and signal transduction pathways involved in the regulation of cellular response to oxidative stress. Life Sci 147(15):67–70
Giotta L, Agostiano A, Italiano F, Milano F, Trotta M (2006) Heavy metal ion influence on the photosynthetic growth of Rhodobacter sphaeroides. Chemosphere 62:1490–1499
Gunasekaran V, Donmez E, Girhard M, Urlacher VB, Constantí M (2014) Biodegradation of fuel oxygenates and their effect on the expression of a newly identified cytochrome P450 gene in Achromobacter xylosoxidans MCM2/2/1. Process Biochem 49(1):124–129
Hendrix S, Schröder P, Keunen E, Huber C, Cuypers A (2017) Chapter six: molecular and cellular aspects of contaminant toxicity in plants: the importance of sulphur and associated signalling pathways. Adv Bot Res 83:223–276
Huang L, Gao X, Liu M, Du G, Guo JS, Ntakirutimana T (2012) Correlation among soil microorganisms, soil enzyme activities, and removal rates of pollutants in three constructed wetlands purifying micro-polluted river water. Ecol Eng 46(3):98–106
Italiano F, Buccolieri A, Giotta L, Agostiano A, Valli L, Milano F, Trotta M (2009) Response of the carotenoidless mutant Rhodobacter sphaeroides growing cells to cobalt and nickel exposure. Int Biodeterior Biodegrad 63:948–957
Kaurin A, Cernilogar Z, Lestan D (2018) Revitalisation of metal-contaminated, EDTA-washed soil by addition of unpolluted soil, compost and biochar: effects on soil enzyme activity, microbial community composition and abundance. Chemosphere 193:726–736
Klein C, Schneider RJ, Meyer MT, Aga DS (2006) Enantiomeric separation of metolachlor and its metabolites using LC–MS and CZE. Chemosphere 62:1591–1599
Kong FY, Wang AJ, Ren HY, Huang LP, Xu MY, Tao HC (2014) Improved dechlorination and mineralization of 4-chlorophenol in a sequential biocathode-bioanode bioelectrochemical system with mixed photosynthetic bacteria. Bioresour Technol 158:32–38
Li LJ, Xia ZB, Ye RZ, Doane TA, Horwath WR (2018) Soil microbial biomass size and soil carbon influence the priming effect from carbon inputs depending on nitrogen availability. Soil Biol Biochem 119:41–49
Liu H, Xiong M (2009) Comparative toxicity of racemic metolachlor and S-metolachlor to Chlorella pyrenoidosa. Aquat Toxicol 93:100–106
Liu XM, Xu X, Li BH, Yao XX, Han YJ (2018a) Genomic and transcriptomic insights into cytochrome P450 monooxygenase genes involved in metolachlor tolerance in maize (Zea mays L.). J Integr Agric 17(8):1790–1799
Liu J, Zhang XY, Wu HH, Gao Y, Silver K, Ma E, Zhang JZ, Zhu KY (2018b) Comparisons of microsomal cytochrome P450 content and enzymatic activity towards selected model substrates and insecticides in different tissues from the migratory locust (Locusta migratoria). Chemosphere 208:366–373
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta DeltaC(T)). Methods 25:402–408
López-Aizpún M, Arango-Mora C, Santamaría C, Lasheras E, Elustondo D (2018) Atmospheric ammonia concentration modulates soil enzyme and microbial activity in an oak forest affecting soil microbial biomass. Soil Biol Biochem 116:378–387
Luo A, Wu YR, Xu Y, Kan J, Qiao J, Liang L, Huang TW, Hu Z (2016) Characterization of a cytochrome P450 monooxygenase capable of high molecular weight PAHs oxidization from Rhodococcus sp. P14. Process Biochem 51(12):2127–2133
Mai H, Gonzalez P, Pardon P, Tapie N, Budzinski H, Cachot J, Morin B (2014) Comparative responses of sperm cells and embryos of Pacific oyster (Crassostrea gigas) to exposure to metolachlor and its degradation products. Aquat Toxicol 147:48–56
Mujahid MD, Sasikala CH, Ramana CHV (2011) Production of indole-3-acetic acid and related indole derivatives from L-tryptophan by Rubrivivax benzoatilyticus JA2. Appl Microbiol Biotechnol 89:1001–1008
Munoz A, Koskinen WC, Cox L, Sadowsky MJ (2011) Biodegradation and Mineralization of Metolachlor and Alachlor by Candida xestobii. J Agric Food Chem 59(2):619–627
Ouchane S, Picaud M, Astier C (1995) A new mutation in the pufL gene responsible for the terbutryn resistance phenotype in Rubrivivax gelatinosus. FEBS Lett 374(1):130–134
Qin W, Deng T, Cui HY, Zhang Q, Liu XD, Yang X, Chen MQ (2018) Exposure to diisodecyl phthalate exacerbated Th2 and Th17-mediated asthma through aggravating oxidative stress and the activation of p38 MAPK. Food Chem Toxicol 114:78–87
Raja V, Majeed U, Kang HS, Andrabi KI, John R (2017) Abiotic stress: interplay between ROS, hormones and MAPKs. Environ Exp Bot 137:142–157
Rose CE, Coupe RH, Capel PD, Webb RMT (2018) Holistic assessment of occurrence and fate of metolachlor within environmental compartments of agricultural watersheds. Sci Total Environ 15:708–719
Sakkas VA, Arabatzis IM, Konstantinou IK, Dimou AD, Albanis TA, Falaras P (2004) Metolachlor photocatalytic degradation using TiO2 photocatalysts. Appl Catal B Environ 49:195–205
Sanchez-Hernandez JC, Pino JND, Capowiez Y, Mazzia C, Rault M (2018) Soil enzyme dynamics in chlorpyrifos-treated soils under the influence of earthworms. Sci Total Environ 612:1407–1416
Sasikala C, Ramana CV (1997) Biodegradation and metabolism of unusual carbon compounds by anoxygenic phototrophic bacteria. Adv Microb Physiol 39:339–377
Schimel J, Becerra CA, Blankinship J (2017) Estimating decay dynamics for enzyme activity in soils from different ecosystems. Soil Biol Biochem 114:5–11
Singh A, Ghoshal N (2013) Impact of herbicide and various soil amendments on soil enzymes activities in a tropical rainfed agroecosystem. Eur J Soil Biol 54:56–62
Sun Y, Zhao LX, Li XJ, Hao YQ, Xu HJ, Weng LP, Li YT (2019) Stimulation of earthworms (Eisenia fetida) on soil microbial communities to promote metolachlor degradation. Environ Pollut 248:219–228
Vance E, Brookes P, Jenkinson D (1987) An extraction method for measuring soil microbial biomass C. Soil Biol Biochem 19:703–707
Wang XG, Huang G, Hao Q, Ran S, Wu YQ, Cui P, Yang J, Jiang CX, Yang QF (2018) Insecticide resistance and enhanced cytochrome P450 monooxygenase activity in field populations of Spodoptera litura from Sichuan, China. Crop Prot 106:110–116
White PM, Potter TL, Culbreath AK (2010) Fungicide dissipation and impact on metolachlor aerobic soil degradation and soil microbial dynamics. Sci Total Environ 408:1393–1402
Wu P, Li JZ, Wang YL, Liu XS, Du C, Tong QY, Li N (2014) Promoting the growth of Rubrivivax gelatinosus in sewage purification by addition of magnesium ions. Biochem Eng J 91:66–71
Wu P, Zhang Y, Chen ZB, Wang YL, Zhu FF, Cao B, Wu Y, Li N (2019a). The organophosphorus pesticides in soil was degradated by Rhodobacter sphaeroides after wastewater treatment. Biochem Eng J 141:247–251
Wu P, Chen ZB, Zhang Y, Wang YL, Zhu FF, Cao B, Wu Y, Li N (2019b). Rhodopseudomonas palustris wastewater treatment: cyhalofop-butyl removal, biochemicals production and mathematical model establishment. Bioresource Tech 282:390–397
Yang AQ, Zhang GM, Yang G, Wang HY, Meng F, Wang HC, Meng Peng M (2017) Denitrification of aging biogas slurry from livestock farm by photosynthetic bacteria. Bioresour Technol 232:408–411
Yu HQ, Wilson F, Tay J (1998) Kinetic analysis of an anaerobic filter treating starch sewage. Water Res 32:3341–3352
Funding
The authors received financial support and basic scientific research service fees from the National Nature Science Fund for Distinguished Young Scholars of China (Grant No. 41625002), the National Natural Science Foundation Young of China (Grant No. 31700432), the National Natural Science Foundation of China (Grant No. 31700432; 31470550; 81500493; 31400386; 51778114), Natural Science Foundation of Guangdong Province (Grant No. 2015A030313098), Application Technology Research and Development Projects of Harbin (Grant No. 2016RAXXJ103), the MOA Modern Agricultural Talents Support Project of the Central University and Dalian Nationalities University (0113-20000101).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Responsible editor: Yanzheng Gao
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 198 kb)
About this article
Cite this article
Wu, P., Shi, J., Zhang, Y. et al. RETRACTED ARTICLE: The bioremediation of metolachlor in soil using Rhodospirillum rubrum after wastewater treatment. J Soils Sediments 19, 3534–3544 (2019). https://doi.org/10.1007/s11368-019-02279-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11368-019-02279-6