Abstract
Purpose
The aim of this study was to evaluate polychlorinated biphenyl (PCB) removal in relation to the associated bacterial community composition in Ohio River sediments (USA) using field and laboratory approaches.
Materials and methods
PCB removal was studied in different sediment layers applying dialysis equilibrators in the field for 4 months. In parallel, dissolved chemical constituents and bacterial community composition using the 16S rDNA cloning–sequencing approach were determined within the appropriate sediment layers. The effects of several bioremediation treatments on PCB removal and bacterial community composition were evaluated in laboratory experiments. Sediments spiked with 245-2’4’5’ hexachlorobiphenyl were incubated under different conditions for up to 250 days, including alternating anaerobic–aerobic conditions, amendments with electron donors, FeSO4, and incubations at different temperatures (10°C, 25°C, 40°C).
Results and discussion
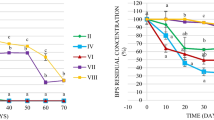
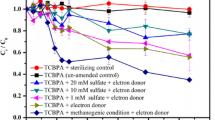
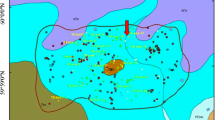
PCB levels remained stable in the sediment layers under field conditions. Surface sediments (0–26 cm) had elevated levels of nitrate+nitrite and sulfate, while subsurface sediments (27–35 cm) were characterized by high concentrations of ammonium, methane, and dissolved organic carbon. Oxic/anoxic conditions were also reflected in the bacterial community, with aerobic bacteria in the Bulkholderiales order enriched in the surface sediments, and anaerobic bacteria in the Clostridiales, Syntrophobacteriales, and Desulfobacterales orders more prevalent in the subsurface sediments. Although the chemical status was favorable and the potential degraders were present, PCB transformation did not take place under the prevailing environmental conditions. Under laboratory conditions, PCB transformation was positively influenced by lowered redox conditions with or without amendments with FeSO4, electron donors, or alternating anaerobic–aerobic conditions. The main PCB transformation process in all anaerobic treatments at 25°C was reductive dechlorination of 245-2’4’5’ hexachlorobiphenyl to 24-2’4’5’ pentachlorobiphenyl and 24-2’4’ tetrachlorobiphenyl, which occurred at rates between 60–85% in 177 days. Exposing sediments to air for an additional period of 100 days in the sequential anaerobic–aerobic treatment did not result in additional mineralization of PCBs. Temperature, electron donors, and redox conditions significantly affected the abundance of bacteria in the Bacteroidales, Hydrogenophilales, Pseudomonales, Myxococcales, Syntrophobacterales, Acidobacterales, Caldilineales, and Clostridiales orders. The PCB dechlorinator Dehalococcoides and other dechlorinating bacteria were detected in anaerobic sediments, including Desulfitobacterium, Desulfuromonas, Anaeromyxobacter, Geobacter, and Desulfomonile. The data suggested that shifts in the bacterial communities due to abiotic conditions affected PCB transformation, either directly (degraders) or indirectly (bacterial syntrophic associations with degraders).
Conclusions
The data indicated that elevated redox status and low temperature conditions accounted for stable PCB concentrations under field conditions in Ohio River sediments. In laboratory experiments, PCB removal increased under anaerobic conditions at 25°C, but was not affected by other varying conditions. Depending on the environmental conditions, several bacterial groups involved in the stepwise degradation of natural organic matter to dechlorinating substrates acetate and hydrogen were prevalent in sediments, including Clostridiales, Caldilineales, Syntrophobacterales, Bacteroidales, and Acidobacteriales. Measurements of the prevailing bacterial community composition and chemical–physical properties important for PCB degradation could be valuable prior to developing PCB bioremediation strategies in sediments.






Similar content being viewed by others
References
Acinas SG, Sarma-Rupavtarm R, Klepac-Ceraj V, Polz MF (2005) PCR-induced sequence artifacts and bias: insights from comparison of two 16S rRNA clone libraries constructed from the same sample. Appl Environ Microbiol 71:8966–8969
Alder AC, Haggblom MM, Oppenheimer SR, Young LY (1993) Reductive dechlorination of polychlorinated biphenyls in anaerobic sediments. Environ Sci Technol 27:530–538
ATSDR (2001) Toxicological profile for polychlorinated biphenyls (PCBs). Agency for Toxic Substances & Disease Registry. http://www.atsdr.cdc.gov/tfacts17.html Accessed 09 December 2009
Backer JG, Berardesco G, Rittmann BE, Stahl DA (2005) The role of syntrophic associations in sustaining anaerobic mineralization of chlorinated organic compounds. Environ Health Perspect 113:310–316
Bedard DL (2003) Polychlorinated biphenyls in aquatic sediments: environmental fate and outlook for biological treatment. In: Häggblom MM, Bossert I (eds) Dehalogenation: microbial processes and environmental applications. Kluwer Press, pp 443–465
Bedard DL, Quensen JF III (1995) Microbial reductive dechlorination of polychlorinated biphenyls. In: Young LY, Cerniglia CE (eds) Microbial transformations and degradation of toxic organic chemicals. Wiley, New York, pp 127–216
Bedard DL, Unterman R, Bopp L, Brennan MJ, Haberl ML, Johnson C (1986) Rapid assay for screening and characterizing microorganisms for the ability to degrade polychlorinated biphenyls. Appl Environ Microbiol 51:761–768
Bedard DL, Ritalahti KM, Löffler FE (2007) Dehalococcoides in a sediment-free, mixed culture metabolically dechlorinate the commercial polychlorinated biphenyl mixture Aroclor 1260. Appl Environ Microbiol 73:2513–2521
Borja J, Taleon DM, Auresenia J, Gallardo S (2005) Polychlorinated biphenyls and their biodegradation. Process Biochem 40:1999–2013
Briée C, Moreira D, López-Garcia P (2007) Archaeal and bacterial community composition of sediment and plankton from a suboxic freshwater pond. Res Microbiol 158:213–227
Brown JF, Bedard DL, Brennan MJ, Carnahan JC, Feng H, Wagner RE (1987) Polychlorinated biphenyl dechlorination in aquatic sediments. Science 236:709–712
Chang Y, Pfeffer JT, Chian ESK (1979) Comparative study of different iron compounds in inhibition of Sphaerotilus growth. Appl Environ Microbiol 38:385–389
Charney AL, Marbach EP (1962) Modified reagents for urea and ammonia. Clin Chem 8:130–132
Chen S, Song L, Dong X (2006) Sporacetigenium mesophilum gen. nov. sp. nov. isolated from an anaerobic digester treating municipal solid waste and sewage. Int J Syst Evol Microbiol 56:721–725
Cole JR, Chai B, Farris RJ, Wang Q, Kulam-Syed-Mohideen AS, McGarrell DM (2007) The ribosomal database project (RDP-II): introducing my RDP space and quality controlled public data. Nucleic Acids Res 35:D169–D172
Cutter LA, Watts JEM, Sowers KR, May HD (2001) Identification of a microorganism that links its growth to the reductive dechlorination of 2, 3, 5, 6-chlorobiphenyl. Environ Microbiol 3:699–709
Crutchfield JD, Burton HR (1998) Improved methods for the quantification of nitrate in plant materials. Anal Lett 22:555–571
D’Angelo EM, Reddy KR (1994) Diagenesis of organic matter in a wetland I. Distribution of dissolved nutrients in the soil and water column. J Environ Qual 23:928–936
Dann AL, Cooper RS, Bowman JP (2009) Investigation and optimization of passively operated compost-based system for remediation of acidic, highly iron- and sulfate-rich industrial waste water. Water Res 43:2302–2316
Drake HL, Horn MA, Wust PA (2009) Intermediary ecosystem metabolism as a main driver of methanogenesis in acidic wetland soil. Environ Microbiol Reports 1:307–318
Drzyzgal O, Gottschal JC (2002) Tetrachloroethene dehalorespiration and growth of Desulfitobacterium frappieri TCE1 in strict dependence on the activity of Desulfovibrio fructosivorans. Appl Environ Microbiol 68:642–649
Evans BS, Dudley CA, Klasson KT (1996) Sequential anaerobic-aerobic biodegradation of PCBs in soil slurry microcosms. Appl Biochem Biotechnol 57(58):885–894
Fazi S, Amalfitano S, Pernthaler J, Puddu A (2006) Bacterial communities associated with benthic organic matter in headwater stream microhabitats. Environ Microbiol 7:1633–1640
Felske A, Wolterink A, van Lis R, de Vos WM, Akkermans ADL (1999) Searching for predominant soil bacteria: 16S rDNA cloning versus strain cultivation. FEMS Microbiol Ecol 30:137–145
Field JA, Sierra-Alvarez R (2008) Microbial transformation and degradation of polychlorinated biphenyls. Environ Pollut 155:1–12
Fierer N, Jackson JA, Vilgalys R, Jackson RB (2005) Assessment of soil microbial community structure by use of taxon-specific quantitative PCR assays. Appl Environ Microbiol 71:4117–4120
Furukawa K (2000) Biochemical and genetic bases of microbial degradation of polychlorinated biphenyls (PCBs). J Gen Appl Microbiol 46:283–296
Gerbersdorf SU, Bittner R, Lubarsky H, Manz W, Paterson DM (2009) Microbial assemblages as ecosystem engineers of sediment stability. J Soils Sediments 9:640–652
Gomes NCM, Fagbola O, Costa R, Rumjanek NG, Buchner A, Mendona-Hagler L, Smalla K (2003) Dynamics of fungal communities in bulk and maize rhizosphere soil in the tropics. Appl Environ Microbiol 69:3758–3766
Gundersen DT, Krahling MD, Donosky JJ, Cable RG, Mims SD (1998) Polychlorinated biphenyls and chlordane in the gonads of paddlefish, Polyodon spathula, from the Ohio River. Bull Environ Contam Toxicol 61:650–657
Hackl E, Zechmeister-Boltenstern S, Bodrossy L, Sessitsch A (2004) Comparison of diversities and compositions of bacterial populations inhabiting natural forest soils. Appl Environ Microbiol 70:5057–5065
Hawker DW, Connell DW (1988) Octanol water partition-coefficients of polychlorinated biphenyl congeners. Environ Sci Technol 22:382–387
Heijs SK, Haese RR, van der Wielen PW, Forney LJ, van Elsas JD (2007) Use of 16S rRNA gene based clone libraries to assess microbial communities potentially involved in anaerobic methane oxidation in a Mediterranean cold seep. Microb Ecol 53:384–398
Hendrickson ER, Payne JA, Young RM, Starr MG, Perry MP, Fahnestock S, Ellis DE, Ebersole RC (2002) Molecular analysis of Dehalococcoides 16S ribosomal DNA from chloroethene-contaminated sites throughout North America and Europe. Appl Environ Microbiol 68:485–495
Hiraishi A (2008) Biodiversity of dehalorespiring bacteria with special emphasis on polychlorinated biphenyl/dioxin dechlorinators. Microbes Environ 23:1–12
Holliger C, Wohlfarth F, Diekert G (1999) Reductive dechlorination in the energy metabolism of anaerobic bacteria. FEMS Microbiol Rev 22:383–398
Holoman TRP, Elbertson MA, Cutter LA, May H, Sowers KR (1998) Characterization of a defined 2, 3, 5, 6 tetrachlorobiphenyl-ortho dechlorinating microbial community by comparitive sequence analysis of genes coding for 16S rRNA. Appl Environ Microbiol 64:3359–3367
Kloep F, Manz W, Röske I (2006) Multivariate analysis of microbial communities in the River Elbe (Germany) on different phylogenetic and spatial levels of resolution. FEMS Microbiol Ecol 56:79–94
Kumar S, Tamura K, Nei M (2004) MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, United Kingdom, pp 115–175
Liu Y, Balkwill DL, Aldrich HC, Drake GR, Boone DR (1999) Characterization of the anaerobic propionate-degrading syntrophs Smithella propionica gen. nov. sp. nov. and Syntrophobacter wolinii. Int J Syst Bacteriol 49:545–556
Maltseva O, Tsoi TV, Quensen JF III, Fukuda M, Tiedje JM (1999) Degradation of anaerobic reductive dechlorination products of Aroclor 1242 by four aerobic bacteria. Biodegradation 10:363–371
Master ER, Lai VWM, Kuipers B, Cullen WR, Mohn WW (2002) Sequential anaerobic-aerobic treatment of soil contaminated with weathered Aroclor 1260. Environ Sci Technol 36:100–103
Meijer SN, Ockenden WA, Sweetman A, Breivik K, Grimalt JO, Jones KC (2003) Global distribution and budget of PCBs and HCB in background surface soils: implications for sources and environmental processes. Environ Sci Technol 37:667–672
Nies L, Vogel TM (1990) Effects of organic substrates on dechlorination of Aroclor 1242 in anaerobic sediments. Appl Environ Microbiol 56:2612–2617
ORSANCO (2002) Ohio River total maximum daily load for PCBs. Ohio River Valley Water Sanitation Commission, Cincinnati, Ohio. www.epa.gov/reg3wapd/tmdl/pa_tmdl/OhioRiver/OhioRiverTMDL.pdf Accessed 09 December 2009
Owens WF, Stuckey DC, Healy JB, Young LY, McCarty PL (1979) Bioassay for monitoring biochemical methane production and anaerobic toxicity. Water Res 13:485–492
Pallud C, van Cappellen P (2006) Kinetics of microbial sulfate reduction in estuarine sediments. Geochim Cosmochim Acta 70:1148–1162
Peplies J, Lachmund C, Glöckner FO, Manz W (2006) A DNA microarrray platform based on direct detection of rRNA for characterization of freshwater sediment-related prokaryotic communities. Appl Environ Microbiol 72:4829–4838
Pieper DH (2005) Aerobic degradation of polychlorinated biphenyls. Appl Microbiol Biotechnol 67:170–191
Qiu X, Wu L, Huang H, McDonel PE, Palumbo AV, Tiedje JM (2001) Evaluation of PCR-generated chimeras, mutations, and heteroduplexes with 16S rRNA gene-based cloning. Appl Environ Microbiol 67:880–887
Quensen JF, Tiedje JM, Boyd SA (1988) Reductive dechlorination of polychlorinated biphenyls by anaerobic microorganisms from sediments. Science 242:752–754
Rhee GY, Bush B, Bethoney CM, DeNucci A, Oh HM, Sokol RC (1993) Anaerobic dechlorination of Aroclor 1242 as affected by some environmental conditions. Environ Toxicol Chem 12:1033–1039
Robertson LW, Hanson LG (2001) PCBs: recent advances in environmental toxicology and health effects. The University Press of Kentucky, Lexington
Rui J, Peng J, Lu Y (2009) Succession of bacterial populations during plant residue decomposition in rice field soil. Appl Environ Microbiol 75:4879–4886
Schipper L, Reddy KR (1995) In situ determination of detrital breakdown in wetland soil-floodwater profile. Soil Sci Soc Am J 59:565–568
Smidt H, de Vos WM (2004) Anaerobic microbial dehalogenation. Annu Rev Microbiol 58:43–73
Suflita JM, Horowitz A, Shelton DR, Tiedje JM (1982) Dehalogenation: a novel pathway for the anaerobic biodegradation of haloaromatic compounds. Science 218:1115–1117
Tringe SG, Hugenholtz P (2008) A renaissance for the pioneering 16S rRNA gene. Curr Opin Microbiol 11:442–446
Ueki A, Abe K, Kaku N, Watanabe K, Ueki K (2008) Bacteroides propionicifaciens sp. nov. isolated from rice-straw residue in a methanogenic reactor treating waste from cattle farms. Int J Syst Evol Microbiol 58:346–352
USACE (2006) Ohio River Water Management Information. 2006 Water Management Division Monthly Reports. http://www.lrd-wc.usace.army.mil/wmrep.html Accessed 09 December 2009
USEPA (2004) National listing of fish advisories. Office of Water. Environmental Protection Agency EPA-828-F-04-016. www.epa.gov/waterscience/fish/advisories/factsheet.pdf Accessed 09 December 2009
USEPA (2006) The Great River News Letter. Environmental Protection Agency EPA/620/N-06/002. www.epa.gov/emap/greatriver/March06.pdf Accessed 09 December 2009
Ward NL, Challacombe JF, Janssen PH et al (2009) Three genomes from the phylum Acidobacteria provide insight into the lifestyles of these microorganisms in soils. Appl Environ Microbiol 75:2046–2056
Wensui L, D’Angelo EM, Coyne MS (2008) Organic carbon effects on aerobic polychlorinated biphenyl removal and bacterial composition in soils and sediments. Chemosphere 70:364–373
Wiegel J, Wu Q (2000) Microbial reductive dechlorination of polychlorinated biphenyls. FEMS Microbiol Ecol 32:1–15
Wu Q, Bedard DL, Wiegel J (1996) Influence of incubation temperature on the microbial reductive dechlorination of 2, 3, 4, 6-tetrachlorobiphenyl in two freshwater sediments. Appl Environ Microbiol 62:4174–4179
Wu Q, Bedard DL, Weigel J (1997) Temperature determines the pattern of anaerobic microbial dechlorination of Aroclor 1260 primed by 2, 3, 4, 6-tetrachlorobiphenyl in Woods Pond sediment. Appl Environ Microbiol 63:4818–4825
Wu Q, Meier GP, Sowers KR, May HD (2002) Reductive dechlorination of polychlorinated benzenes by Bacterium DF-1, a polychlorinated biphenyl-dechlorinating microorganism. Environ Sci Technol 36(15):3290–3294
Wust PK, Horn MA, Drake HL (2009) Trophic links between fermenters and methanogens in a moderately acidic fen soil. Environ Microbiol 11:1395–1409
Yamada T, Sekiguchi Y, Hanada S, Imachi H, Ohashi A, Harada H, Kamagata Y (2006) Anaerolinea thermolimosa sp. nov. Levilinea saccharolytica gen. nov. sp. nov. and Leptolinea tardivitalis gen. nov. sp. nov. novel filamentous anaerobes, and description of the new classes Anaerolineae classis nov. and Caldilineae classis nov. in the bacterial phylum Chloroflexi. Int J Syst Evol Microbiol 56:1331–1340
Zwiernik MJ, Quensen JF III, Boyd SA (1998) FeSO4 amendments stimulate extensive anaerobic PCB dechlorination. Environ Sci Technol 32:3360–3365
Acknowledgments
This research was supported by the National Science Foundation CAREER Award no. 0134114 and the College of Agriculture at the University of Kentucky. Appreciation is also extended to Georgia Zeigler, Lilia Turcios, and Martin Vandiviere for their assistance in the molecular biology work. We would also like to express our appreciation to Dr. Sabine Gerbersdorf for her insightful comments that improved the paper.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Sabine Ulrike Gerbersdorf
Rights and permissions
About this article
Cite this article
D’Angelo, E., Nunez, A. Effect of environmental conditions on polychlorinated biphenyl transformations and bacterial communities in a river sediment. J Soils Sediments 10, 1186–1199 (2010). https://doi.org/10.1007/s11368-010-0218-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11368-010-0218-2