Abstract
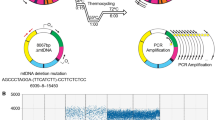
Mitochondria contain multiple copies of the circular mitochondrial genome (mtDNA) that encodes ribosomal RNAs and proteins locally translated for oxidative phosphorylation. Loss of mtDNA integrity, both altered copy number and increased mutations, is implicated in cellular dysfunction with aging. Published data on mtDNA copy number and aging is discordant which may be due to methodological limitations for quantifying mtDNA copy number. Existing quantitative PCR (qPCR) mtDNA copy number quantification methods provide only relative abundances and are problematic to normalize to different template input amounts and across tissues/sample types. As well, existing methods cannot quantify mtDNA copy number in subcellular isolates, such as isolated mitochondria and neuronal synaptic terminals, which lack nuclear genomic DNA for normalization. We have developed and validated a novel absolute mtDNA copy number quantitation method that uses chip-based digital polymerase chain reaction (dPCR) to count the number of copies of mtDNA and used this novel method to assess the literature discrepancy in which there is no clear consensus whether mtDNA numbers change with aging in skeletal muscle. Skeletal muscle in old mice was found to have increased absolute mtDNA numbers compared to young controls. Furthermore, young Sod1 −/− mice were assessed and show an age-mimicking increase in skeletal muscle mtDNA. These findings reproduce a number of previous studies that demonstrate age-related increases in mtDNA. This simple and cost effective dPCR approach should enable precise and accurate mtDNA copy number quantitation in mitochondrial studies, eliminating contradictory studies of mitochondrial DNA content with aging.





Similar content being viewed by others
References
Anson RM, Bohr VA (2000) Mitochondria, oxidative DNA damage, and aging. J Am Aging Assoc 23:199–218. doi:10.1007/s11357-000-0020-y
Bai RK, Perng CL, Hsu CH, Wong LJ (2004) Quantitative PCR analysis of mitochondrial DNA content in patients with mitochondrial disease. Ann N Y Acad Sci 1011:304–309
Baker M (2012) Digital PCR hits its stride. Nat Methods 9:541–544
Barja G (2013) Updating the mitochondrial free radical theory of aging: an integrated view, key aspects, and confounding concepts. Antioxid Redox Signal 19:1420–1445. doi:10.1089/ars.2012.5148
Barrientos A et al. (1997a) Qualitative and quantitative changes in skeletal muscle mtDNA and expression of mitochondrial-encoded genes in the human aging process. Biochem Mol Med 62:165–171
Barrientos A, Casademont J, Cardellach F, Estivill X, Urbano-Marquez A, Nunes V (1997b) Reduced steady-state levels of mitochondrial RNA and increased mitochondrial DNA amount in human brain with aging. Brain Res Mol Brain Res 52:284–289
Benz CC, Yau C (2008) Ageing, oxidative stress and cancer: paradigms in parallax nature reviews. Cancer 8:875–879. doi:10.1038/nrc2522
Chatterjee A, Mambo E, Sidransky D (2006) Mitochondrial DNA mutations in human cancer. Oncogene 25:4663–4674. doi:10.1038/sj.onc.1209604
Chaturvedi RK, Flint Beal M (2013) Mitochondrial diseases of the brain. Free Radic Biol Med 63:1–29. doi:10.1016/j.freeradbiomed.2013.03.018
Clay Montier LL, Deng JJ, Bai Y (2009) Number matters: control of mammalian mitochondrial DNA copy number. J Genet Genomics 36:125–131. doi:10.1016/S1673-8527(08)60099-5
D’Erchia AM et al. (2015) Tissue-specific mtDNA abundance from exome data and its correlation with mitochondrial transcription, mass and respiratory activity. Mitochondrion 20:13–21. doi:10.1016/j.mito.2014.10.005
Dickinson A et al. (2013) The regulation of mitochondrial DNA copy number in glioblastoma cells. Cell Death Differ 20:1644–1653. doi:10.1038/cdd.2013.115
Dimauro S, Davidzon G (2005) Mitochondrial DNA and disease. Ann Med 37:222–232. doi:10.1080/07853890510007368
Dimmock D, Tang LY, Schmitt ES, Wong LJ (2010) Quantitative evaluation of the mitochondrial DNA depletion syndrome. Clin Chem 56:1119–1127. doi:10.1373/clinchem.2009.141549
Ekstrand MI et al. (2004) Mitochondrial transcription factor A regulates mtDNA copy number in mammals. Hum Mol Genet 13:935–944. doi:10.1093/hmg/ddh109
Fernandez-Vizarra E, Enriquez JA, Perez-Martos A, Montoya J, Fernandez-Silva P (2011) Tissue-specific differences in mitochondrial activity and biogenesis. Mitochondrion 11:207–213. doi:10.1016/j.mito.2010.09.011
Ferreira R, Vitorino R, Alves RM, Appell HJ, Powers SK, Duarte JA, Amado F (2010) Subsarcolemmal and intermyofibrillar mitochondria proteome differences disclose functional specializations in skeletal muscle. Proteomics 10:3142–3154. doi:10.1002/pmic.201000173
Gadaleta MN, Rainaldi G, Lezza AM, Milella F, Fracasso F, Cantatore P (1992) Mitochondrial DNA copy number and mitochondrial DNA deletion in adult and senescent rats. Mutat Res 275:181–193
Hindson BJ et al. (2011) High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal Chem 83:8604–8610. doi:10.1021/ac202028g
Hindson CM et al. (2013) Absolute quantification by droplet digital PCR versus analog real-time PCR. Nat Methods 10:1003–1005. doi:10.1038/nmeth.2633
Huang JH, Hood DA (2009) Age-associated mitochondrial dysfunction in skeletal muscle: contributing factors and suggestions for long-term interventions. IUBMB Life 61:201–214. doi:10.1002/iub.164
Jang YC et al. (2010) Increased superoxide in vivo accelerates age-associated muscle atrophy through mitochondrial dysfunction and neuromuscular junction degeneration. FASEB J 24:1376–1390. doi:10.1096/fj.09-146308
Kowluru RA (2013) Mitochondria damage in the pathogenesis of diabetic retinopathy and in the metabolic memory associated with its continued progression. Curr Med Chem 20:3226–3233
Lanza IR et al. (2008) Endurance exercise as a countermeasure for aging. Diabetes 57:2933–2942. doi:10.2337/db08-0349
Lee HC, Yin PH, Lu CY, Chi CW, Wei YH (2000) Increase of mitochondria and mitochondrial DNA in response to oxidative stress in human cells. Biochem J 348(Pt 2):425–432
Malik AN, Czajka A (2013) Is mitochondrial DNA content a potential biomarker of mitochondrial dysfunction? Mitochondrion 13:481–492. doi:10.1016/j.mito.2012.10.011
Masser DR, Berg AS, Freeman WM (2013) Focused, high accuracy 5-methylcytosine quantitation with base resolution by benchtop next-generation sequencing. Epigenetics Chromatin 6:33. doi:10.1186/1756-8935-6-33
Masser DR et al. (2014) Hippocampal subregions exhibit both distinct and shared transcriptomic responses to aging and nonneurodegenerative cognitive decline. J Gerontol A Biol Sci Med Sci 69:1311–1324. doi:10.1093/gerona/glu091
Masuyama M, Iida R, Takatsuka H, Yasuda T, Matsuki T (2005) Quantitative change in mitochondrial DNA content in various mouse tissues during aging. Biochim Biophys Acta 1723:302–308. doi:10.1016/j.bbagen.2005.03.001
McInerny SC, Brown AL, Smith DW (2009) Region-specific changes in mitochondrial D-loop in aged rat CNS. Mech Ageing Dev 130:343–349. doi:10.1016/j.mad.2009.01.008
Miller FJ, Rosenfeldt FL, Zhang C, Linnane AW, Nagley P (2003) Precise determination of mitochondrial DNA copy number in human skeletal and cardiac muscle by a PCR-based assay: lack of change of copy number with age. Nucleic Acids Res 31:e61
Morrison T et al. (2006) Nanoliter high throughput quantitative PCR. Nucleic Acids Res 34:e123. doi:10.1093/nar/gkl639
Muller FL et al. (2006) Absence of CuZn superoxide dismutase leads to elevated oxidative stress and acceleration of age-dependent skeletal muscle atrophy. Free Radic Biol Med 40:1993–2004. doi:10.1016/j.freeradbiomed.2006.01.036
Nagahisa H, Okabe K, Iuchi Y, Fujii J, Miyata H (2016) Characteristics of skeletal muscle fibers of SOD1 knockout mice. Oxidative Med Cell Longev 2016:9345970. doi:10.1155/2016/9345970
Nicklas JA, Brooks EM, Hunter TC, Single R, Branda RF (2004) Development of a quantitative PCR (TaqMan) assay for relative mitochondrial DNA copy number and the common mitochondrial DNA deletion in the rat. Environ Mol Mutagen 44:313–320. doi:10.1002/em.20050
Pesce V et al. (2001) Age-related mitochondrial genotypic and phenotypic alterations in human skeletal muscle. Free Radic Biol Med 30:1223–1233
Phillips NR, Sprouse ML, Roby RK (2014) Simultaneous quantification of mitochondrial DNA copy number and deletion ratio: a multiplex real-time PCR assay. Sci Rep 4:3887. doi:10.1038/srep03887
Sanz A et al. (2007) Evaluation of sex differences on mitochondrial bioenergetics and apoptosis in mice. Exp Gerontol 42:173–182. doi:10.1016/j.exger.2006.10.003
Sharma K (2015) Mitochondrial hormesis and diabetic complications. Diabetes 64:663–672. doi:10.2337/db14-0874
Shmookler Reis RJ, Goldstein S (1983) Mitochondrial DNA in mortal and immortal human cells. Genome number, integrity, and methylation. J Biol Chem 258:9078–9085
Short KR, Bigelow ML, Kahl J, Singh R, Coenen-Schimke J, Raghavakaimal S, Nair KS (2005) Decline in skeletal muscle mitochondrial function with aging in humans. Proc Natl Acad Sci U S A 102:5618–5623. doi:10.1073/pnas.0501559102
Taylor RW, Turnbull DM (2005) Mitochondrial DNA mutations in human disease. Nat Rev Genet 6:389–402. doi:10.1038/nrg1606
van Leeuwen N, Beekman M, Deelen J, van den Akker EB, de Craen AJ, Slagboom PE, ‘t Hart LM (2014) Low mitochondrial DNA content associates with familial longevity: the Leiden longevity study. Age (Dordr) 36:9629. doi:10.1007/s11357-014-9629-0
VanGuilder HD, Brucklacher RM, Patel K, Ellis RW, Freeman WM, Barber AJ (2008) Diabetes downregulates presynaptic proteins and reduces basal synapsin I phosphorylation in rat retina. Eur J Neurosci 28:1–11. doi:10.1111/j.1460-9568.2008.06322.x
Veltri KL, Espiritu M, Singh G (1990) Distinct genomic copy number in mitochondria of different mammalian organs. J Cell Physiol 143:160–164. doi:10.1002/jcp.1041430122
Vogelstein B, Kinzler KW (1999) Digital PCR. Proc Natl Acad Sci U S A 96:9236–9241
Wanagat J, Ahmadieh N, Bielas JH, Ericson NG, Van Remmen H (2015) Skeletal muscle mitochondrial DNA deletions are not increased in CuZn-superoxide dismutase deficient mice. Exp Gerontol 61:15–19. doi:10.1016/j.exger.2014.11.012
Wang T, Sha H, Ji D, Zhang HL, Chen D, Cao Y, Zhu J (2014) Polar body genome transfer for preventing the transmission of inherited mitochondrial diseases. Cell 157:1591–1604. doi:10.1016/j.cell.2014.04.042
Warren L, Bryder D, Weissman IL, Quake SR (2006) Transcription factor profiling in individual hematopoietic progenitors by digital RT-PCR. Proc Natl Acad Sci U S A 103:17807–17812. doi:10.1073/pnas.0608512103
Welle S, Bhatt K, Shah B, Needler N, Delehanty JM, Thornton CA (2003) Reduced amount of mitochondrial DNA in aged human muscle. J Appl Physiol (1985) 94:1479–1484. doi:10.1152/japplphysiol.01061.2002
Acknowledgments
The authors wish to thank the Laboratory for Molecular Biology and Cytometry Research at OUHSC for providing Sanger sequencing assistance and Peter Eckhart for assistance with figure generation. This work was supported by the National Institutes of Health [National Eye Institute R01EY02176, R21EY024520 to WMF, T32EY023202 to DRM]; National Institute on Aging Oklahoma Nathan Shock Center (P30AG050911) to HVR, WMF; the Donald W. Reynolds Foundation [DRM & WMF]; The University of Oklahoma Health Sciences Center Graduate Student Association research grant to [DRM]; and in part by an award from Harold Hamm Diabetes Center at the University of Oklahoma [DRM].
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
All procedures were approved by the Institutional Animal Care and Use Committee at the Oklahoma Medical Research Foundation.
Competing interests
The authors declare that they have no competing interests.
About this article
Cite this article
Masser, D.R., Clark, N.W., Van Remmen, H. et al. Loss of the antioxidant enzyme CuZnSOD (Sod1) mimics an age-related increase in absolute mitochondrial DNA copy number in the skeletal muscle. AGE 38, 323–333 (2016). https://doi.org/10.1007/s11357-016-9930-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11357-016-9930-1