Abstract
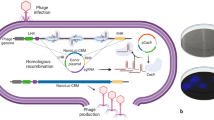
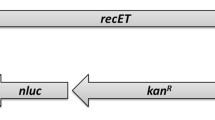
Water quality is a major safety consideration in environments that are impacted by human activity. The key challenge of the COMBITOX project is to develop a unique instrument that can accommodate several biodetector systems (see the accompanying COMBITOX papers) able to detect different pollutants such as bacteria, toxins, and heavy metals. The output signal chosen by our consortium is based on luminescence detection. Our group recently developed phage-based biosensors using gfp as a reporter gene to detect enteric bacteria in complex environments such as sea water, and the main challenge we faced was to adapt our biodetector to a luminescent signal that could fit the COMBITOX project requirements. Another key point was to use a substrate-independent reporter system in order to avoid substrate addition in the detection prototype. This paper describes the development of a phage-based biodetector using a luminescent and substrate-independent output to detect some enteric bacteria, such as Escherichia coli, in water samples. We have successfully engineered various prototypes using the HK620 and HK97 bacteriophages that use different packaging systems, and both proved functional for the integration of the full luxCDABE operon controlled by two different bacterial promoters. We show that the luxCDABE operon controlled by the PrplU bacterial promoter is the most efficient in terms of signal emission. The emission of luminescence is specific and allows the detection of 104 bacteria per milliliter in 1.5 h post-infection with neither a concentration nor enrichment step.






Similar content being viewed by others
References
Ackermann H-W (2009) Basic phage electron microscopy. Methods Mol. Biol. Clifton NJ 501:113–126
Ansaldi M, Bazin I, Cholat P, Rodrigue A, Pignol D (2015) Toward inline multiplex biodetection of metals, bacteria, and toxins in water networks: the COMBITOX project. Environ Sci Pollut Res Int. doi:10.1007/s11356-015-5582-4
Blattner FR, Plunkett G, Bloch CA, Perna NT, Burland V, Riley M, Collado-Vides J, Glasner JD, Rode CK, Mayhew GF et al (1997) The complete genome sequence of Escherichia coli K-12. Science 277:1453–1474
Byappanahalli MN, Nevers MB, Korajkic A, Staley ZR, Harwood VJ (2012) Enterococci in the environment. Microbiol Mol Biol Rev 76:685–706
Campbell A (1994) Comparative molecular biology of lambdoid phages. Annu Rev Microbiol 48:193–222
Clark AJ, Inwood W, Cloutier T, Dhillon TS (2001) Nucleotide sequence of coliphage HK620 and the evolution of lambdoid phages. J Mol Biol 311:657–679
D’Herelle F (2007) On an invisible microbe antagonistic toward dysenteric bacilli: brief note by Mr. F. D’Herelle, presented by Mr. Roux. 1917. Res Microbiol 158:553–554
Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci U S A 97:6640–6645
Derda R, Lockett MR, Tang SKY, Fuller RC, Maxwell EJ, Breiten B, Cuddemi CA, Ozdogan A, Whitesides GM (2013) Filter-based assay for Escherichia coli in aqueous samples using bacteriophage-based amplification. Anal Chem 85:7213–7220
Descamps, E., Franche, Nathalie, Meunier, D., Miclot, B., Larosa, P., Brutesco, C., Garcia, D., Bazin, I., Pignol, D., Cholat, P., et al. (in this issue). Semi-autonomous inline water analyzer: design of a common light detector for bacterial, phage and immunological biosensors.
Dhillon EK, Dhillon TS, Lai AN, Linn S (1980) Host range, immunity and antigenic properties of lambdoid coliphage HK97. J Gen Virol 50:217–220
Dhillon TS, Poon AP, Chan D, Clark AJ (1998) General transducing phages like Salmonella phage P22 isolated using a smooth strain of Escherichia coli as host. FEMS MicrobiolLett 161:129–133
Edberg SC, Rice EW, Karlin RJ, Allen MJ (2000) Escherichia coli: the best biological drinking water indicator for public health protection. Symp Ser Soc Appl Microbiol 29:106S–116S
Edgar R (2006) High-sensitivity bacterial detection using biotin-tagged phage and quantum-dot nanocomplexes. Proc Natl Acad Sci 103:4841–4845
Goodridge L, Abedon ST (2003) Bacteriophage biocontrol and bioprocessing: application of phage therapy to industry. SIM News 53:254–262
Haq IU, Chaudhry WN, Akhtar MN, Andleeb S, Qadri I (2012) Bacteriophages and their implications on future biotechnology: a review. Virol J 9:9
Hendrix R, Roberts JW, Stahl FW, Weisberg RA (1983) Lambda II. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Juhala RJ, Ford ME, Duda RL, Youlton A, Hatfull GF, Hendrix RW (2000) Genomic sequences of bacteriophages HK97 and HK022: pervasive genetic mosaicism in the lambdoid bacteriophages. J Mol Biol 299:27–51
Kim S, Kim M, Ryu S (2014) Development of an engineered bioluminescent reporter phage for the sensitive detection of viable Salmonella Typhimurium. Anal Chem 86:5858–5864
Kuhn JC (2007) Detection of Salmonella by bacteriophage Felix 01. Methods Mol. Biol. Clifton NJ 394:21–37
Li L, Mendis N, Trigui H, Oliver JD, Faucher SP (2014) The importance of the viable but non-culturable state in human bacterial pathogens. Front Microbiol 5
Loessner MJ, Rees CE, Stewart GS, Scherer S (1996) Construction of luciferase reporter bacteriophage A511::luxAB for rapid and sensitive detection of viable Listeria cells. Appl Environ Microbiol 62:1133–1140
Menouni R, Champ S, Espinosa L, Boudvillain M, Ansaldi M (2013) Transcription termination controls prophage maintenance in Escherichia coli genomes. Proc Natl Acad Sci U S A 110:14414–14419
Mosier-Boss PA, Lieberman SH, Andrews JM, Rohwer FL, Wegley LE, Breitbart M (2003) Use of fluorescently labeled phage in the detection and identification of bacterial species. Appl Spectrosc 57:1138–1144
Panis G, Duverger Y, Champ S, Ansaldi M (2010) Protein binding sites involved in the assembly of the KplE1 prophage intasome. Virology 404:41–50
Piuri M, Jacobs WR, Hatfull GF (2009) Fluoromycobacteriophages for rapid, specific, and sensitive antibiotic susceptibility testing of mycobacterium tuberculosis. PLoS ONE 4, e4870
Pond K, World Health Organization (2005) Water recreation and disease plausibility of associated infections: acute effects, sequelae, and mortality. Published on behalf of the World Health Organization by IWA Pub, London; Seattle
Prévéral S, Brutesco C, Descamps ECT, Escoffier C, Pignol D, Ginet N, Garcia D (2016) A bioluminescent arsenite biosensor designed for inline water analyzer. Environ Sci Pollut Res Int. doi:10.1007/s11356-015-6000-7
Preveral, S., Brutesco, C., Descamps, E., Escoffier, C., Ginet, N., Pignol, D., and Garcia, D (in this issue). A bioluminescent arsenite biosensor designed for inline water analyzer.
Schmelcher M, Loessner MJ (2014) Application of bacteriophages for detection of foodborne pathogens. Bacteriophage 4, e28137
Schofield DA, Molineux IJ, Westwater C (2009) Diagnostic bioluminescent phage for detection of Yersinia pestis. J Clin Microbiol 47:3887–3894
Springman R, Molineux IJ, Duong C, Bull RJ, Bull JJ (2012) Evolutionary stability of a refactored phage genome. ACS Synth Biol 1:425–430
Stewart JR, Gast RJ, Fujioka RS, Solo-Gabriele HM, Meschke JS, Amaral-Zettler LA, del Castillo E, Polz MF, Collier TK, Strom MS et al (2008) The coastal environment and human health: microbial indicators, pathogens, sentinels and reservoirs. Environ Health 7:S3
Toranzo AE, Magariños B, Romalde JL (2005) A review of the main bacterial fish diseases in mariculture systems. Aquaculture 246:37–61
Twort FW (1915) An investigation on the nature of ultra-microscopic viruses. Lancet 186:1241–1243
Vinay M, Franche N, Grégori G, Fantino J-R, Pouillot F, Ansaldi M (2015) Phage-based fluorescent biosensor prototypes to specifically detect enteric bacteria such as E. coli and Salmonella enterica Typhimurium. PLoS ONE 10, e0131466
Waddell TE, Poppe C (2000) Construction of mini-Tn10luxABcam/Ptac-ATS and its use for developing a bacteriophage that transduces bioluminescence to Escherichia coli O157:H7. FEMS Microbiol Lett 182:285–289
Zaslaver A, Bren A, Ronen M, Itzkovitz S, Kikoin I, Shavit S, Liebermeister W, Surette MG, Alon U (2006) A comprehensive library of fluorescent transcriptional reporters for Escherichia coli. Nat Methods 3:623–628
Acknowledgments
We thank all the members of the phage lab and the COMBITOX consortium for stimulating discussions. We are grateful to Artemis Costa (electronic microscopy facility) and Leon Espinosa for help with the microscopy and data analysis and to Marie-Agnès Petit for sending the HK97 lysogen.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Philippe Garrigues
Rights and permissions
About this article
Cite this article
Franche, N., Vinay, M. & Ansaldi, M. Substrate-independent luminescent phage-based biosensor to specifically detect enteric bacteria such as E. coli . Environ Sci Pollut Res 24, 42–51 (2017). https://doi.org/10.1007/s11356-016-6288-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-016-6288-y