Abstract
Introduction
There has been a growing interest towards creating defined mixed starter cultures for alcoholic fermentations. Previously, metabolite differences between single and mixed cultures have been explored at the endpoint of fermentations rather than during fermentations.
Objectives
To create metabolic footprints of metabolites that discriminate single and mixed yeast cultures at two key time-points during mixed culture alcoholic fermentations.
Methods
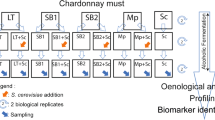
1H NMR- and GC–MS-based metabolomics was used to identify metabolites that discriminate single and mixed cultures of Lachancea thermotolerans (LT) and Saccharomyces cerevisiae (SC) during alcoholic fermentations.
Results
Twenty-two metabolites were found when comparing single LT and mixed cultures, including both non-volatiles (carbohydrate, amino acid and acids) and volatiles (higher alcohols, esters, ketones and aldehydes). Fifteen of these compounds were discriminatory only at the death phase initiation (T1) and fifteen were discriminatory only at the death phase termination (T2) of LT in mixed cultures. Eight metabolites were discriminatory at both T1 and T2. These results indicate that specific metabolic changes may be descriptive of different LT growth behaviors. Fifteen discriminatory metabolites were found when comparing single SC and mixed cultures. These metabolites were all volatiles, and twelve metabolites were discriminatory only at T2, indicating that LT-induced changes in volatiles occur during the death phase of LT in mixed cultures and not during their initial growth stage.
Conclusions
This work provides a detailed insight into yeast metabolites that differ between single and mixed cultures, and these data may be used for understanding and eventually predicting yeast metabolic changes in wine fermentations.






Similar content being viewed by others
Data availability
The data that support the findings of this study are available within the article and its supplementary information files.
References
Al-Hadid, Q., Roy, K., Munroe, W., Dzialo, M. C., Chanfreau, G. F., & Clarke, S. G. (2014). Histidine methylation of yeast ribosomal protein Rpl3p is required for proper 60S subunit assembly. Molecular and Cellular Biology, 34, 2903–2916.
Alonso-del-Real, J., Contreras-Ruiz, A., Castiglioni, G. L., Barrio, E., & Querol, A. (2017). The use of mixed populations of Saccharomyces cerevisiae and S. kudriavzevii to reduce ethanol content in wine: Limited aeration, inoculum proportions, and sequential inoculation. Frontiers in Microbiology, 8, 2087.
Bandara, A., Fraser, S., Chambers, P. J., & Stanley, G. A. (2009). Trehalose promotes the survival of Saccharomyces cerevisiae during lethal ethanol stress, but does not influence growth under sublethal ethanol stress. FEMS Yeast Research, 9, 1208–1216.
Cacho, J. I., Campillo, N., Viñas, P., & Hernández-Córdoba, M. (2013). Stir bar sorptive extraction with EG-silicone coating for bisphenols determination in personal care products by GC-MS. Journal of Pharmaceutical and Biomedical Analysis, 78–79, 255–260.
Ciani, M., Beco, L., & Comitini, F. (2006). Fermentation behaviour and metabolic interactions of multistarter wine yeast fermentations. International Journal of Food Microbiology, 108, 239–245.
Ciani, M., Comitini, F., Mannazzu, I., & Domizio, P. (2010). Controlled mixed culture fermentation: A new perspective on the use of non-Saccharomyces yeasts in winemaking. FEMS Yeast Research, 10, 123–133.
Comitini, F., Gobbi, M., Domizio, P., Romani, C., Lencioni, L., Mannazzu, I., et al. (2011). Selected non-Saccharomyces wine yeasts in controlled multistarter fermentations with Saccharomyces cerevisiae. Food Microbiology, 28, 873–882.
Dzialo, M. C., Park, R., Steensels, J., Lievens, B., & Verstrepen, K. J. (2017). Physiology, ecology and industrial applications of aroma formation in yeast. FEMS Microbiology Reviews, 41, S95–S128.
Howell, K. S., Cozzolino, D., Bartowsky, E. J., Fleet, G. H., & Henschke, P. A. (2006). Metabolic profiling as a tool for revealing Saccharomyces interactions during wine fermentation. FEMS Yeast Reviews, 6, 91–101.
Kemsawasd, V., Viana, T., Ardö, Y., & Arneborg, N. (2015). Influence of nitrogen sources on growth and fermentation performance of different wine yeast species during alcoholic fermentation. Applied Microbiology and Biotechnology, 99, 10191–10207.
Kim, H. K., Choi, Y. H., & Verpoorte, R. (2011). NMR-based plant metabolomics: Where do we stand, where do we go? Trends in Biotechnology, 29, 267–275.
Kristensen, M., Savorani, F., Ravn-Haren, G., Poulsen, M., Markowski, J., Larsen, F. H., et al. (2010). NMR and interval PLS as reliable methods for determination of cholesterol in rodent lipoprotein fractions. Metabolomics, 6, 129–136.
Lee, S., Do, S., Kim, S. Y., Kim, J., Jin, Y., & Lee, C. H. (2012). Mass spectrometry-based metabolite profiling and antioxidant activity of Aloe vera (Aloe barbadensis Miller) in different growth stages. Journal of Agricultural and Food Chemistry, 60, 11222–11228.
Liu, J., Arneborg, N., Toldam-Andersen, T. B., Zhang, S., Petersen, M. A., & Bredie, W. L. (2017). Impact of sequential co-culture fermentations on flavour characters of Solaris wines. European Food Research and Technology, 243, 437–445.
Liu, J., Toldam-Andersen, T. B., Petersen, M. A., Zhang, S., Arneborg, N., & Bredie, W. L. P. (2015). Instrumental and sensory characterisation of Solaris white wines in Denmark. Food Chemistry, 166, 133–142.
López-Rituerto, E., Avenoza, A., Busto, J. H., & Peregrina, J. M. (2013). NMR study of histidine metabolism during alcoholic and malolactic fermentations of wine and their influence on histamine production. Journal of Agricultural and Food Chemistry, 61, 9464–9469.
López-Rituerto, E., Cabredo, S., López, M., Avenoza, A., Busto, J. H., & Peregrina, J. M. (2009). A thorough study on the use of quantitative 1H NMR in Rioja red wine fermentation processes. Journal of Agricultural and Food Chemistry, 57, 2112–2118.
López-Rituerto, E., Savorani, F., Avenoza, A., Busto, J. H., Peregrina, J. M., & Engelsen, S. B. (2012). Investigations of La Rioja terroir for wine production using 1H NMR metabolomics. Journal of Agricultural and Food Chemistry, 60, 3452–3461.
Mapelli, V., Olsson, L., & Nielsen, J. (2008). Metabolic footprinting in microbiology: Methods and applications in functional genomics and biotechnology. Trends in Biotechnology, 26, 490–497.
Nicholson, J. K., & Lindon, J. C. (2008). Systems biology: Metabonomics. Nature, 455, 1054–1056.
Nilsson, M., Duarte, I. F., Almeida, C., Delgadillo, I., Goodfellow, B. J., Gil, A. M., et al. (2004). High-resolution NMR and diffusion-ordered spectroscopy of port wine. Journal of Agricultural and Food Chemistry, 52, 3736–3743.
Nørgaard, L., Saudland, A., Wagner, J., Nielsen, J. P., Munck, L., & Engelsen, S. B. (2000). Interval partial least squares regression (iPLS): A comparative chemometric study with an example from the near infrared spectroscopy. Applied Spectroscopy, 54, 413–419.
Ocón, A., Hampp, R., & Requena, N. (2007). Trehalose turnover during abiotic stress in arbuscular mycorrhizal fungi. New Phytologist, 174, 879–891.
Padilla, B., Gil, J. V., & Manzanares, P. (2016). Past and future of non-Saccharomyces yeasts: From spoilage microorganisms to biotechnological tools for improving wine aroma complexity. Frontiers in Microbiology, 7, 411.
Peng, C. T., Wen, Y., Tao, Y. S., & Lan, Y. Y. (2013). Modulating the formation of Meili wine aroma by prefermentative freezing process. Journal of Agricultural and Food Chemistry, 61, 1542–1553.
Puig-Castellví, F., Alfonso, I., Piña, B., & Tauler, R. (2015). A quantitative 1H NMR approach for evaluating the metabolic response of Saccharomyces cerevisiae to mild heat stress. Metabolomics, 11, 1612–1625.
Renault, P., Coulon, J., de Revel, G., Barbe, J. C., & Bely, M. (2015). Increase of fruity aroma during mixed T. delbrueckii/S. cerevisiae wine fermentation is linked to specific esters enhancement. International Journal of Food Microbiology, 207, 40–48.
Sadoudi, M., Tourdot-Maréchal, R., Rousseaux, S., Steyer, D., Gallardo-Chacón, J. J., Ballester, J., et al. (2012). Yeast-yeast interactions revealed by aromatic profile analysis of Sauvignon Blanc wine fermented by single or co-culture of non-Saccharomyces and Saccharomyces yeasts. Food Microbiology, 32, 243–253.
Savorani, F., Tomasi, G., & Engelsen, S. B. (2010). icoshift: A versatile tool for the rapid alignment of 1D NMR spectra. Journal of Magnetic Resonance, 202, 190–202.
Son, H. S., Hwang, G. S., Kim, K. M., Kim, E. Y., van den Berg, F., Park, W. M., et al. (2009). 1H NMR-based metabolomic approach for understanding the fermentation behaviors of wine yeast strains. Analytical Chemistry, 81, 1137–1145.
Sumner, L. W., Amberg, A., Barrett, D., Beale, M. H., Beger, R., Daykin, C. A., et al. (2007). Proposed minimum reporting standards for chemical analysis. Metabolomics, 3, 211–221.
Takahashi, H., Kai, K., Shinbo, Y., Tanaka, K., Ohta, D., Oshima, T., et al. (2008). Metabolomics approach for determining growth-specific metabolites based on Fourier transform ion cyclotron resonance mass spectrometry. Analytical and Bioanalytical Chemistry, 391, 2769–2782.
Tredoux, A., De Villiers, A., Májek, P., Lynen, F., Crouch, A., & Sandra, P. (2008). Stir bar sorptive extraction combined with GC-MS analysis and chemometric methods for the classification of South African wines according to the volatile composition. Journal of Agricultural and Food Chemistry, 56, 4286–4296.
Ugliano, M., & Henschke, P. A. (2009). Yeasts and wine flavour. In M. V. Moreno-Arribas & M. C. Polo (Eds.), Wine chemistry and biochemistry (pp. 313–392). New York: Springer.
Varela, C., Sengler, F., Solomon, M., & Curtin, C. (2016). Volatile flavour profile of reduced alcohol wines fermented with the non-conventional yeast species Metschnikowia pulcherrima and Saccharomyces uvarum. Food Chemistry, 209, 57–64.
Viana, T., Loureiro-Dias, M. C., & Prista, C. (2014). Efficient fermentation of an improved synthetic grape must by enological and laboratory strains of Saccharomyces cerevisiae. AMB Express, 4, 16.
Vold, S. (1987). Principal component analysis. Chemometrics and Intelligent Laboratory Systems, 2, 37–52.
Wang, X. C., Li, A. H., Dizy, M., Ullah, N., Sun, W. X., & Tao, Y. S. (2017). Evaluation of aroma enhancement for “Ecolly” dry white wines by mixed inoculation of selected Rhodotorula mucilaginosa and Saccharomyces cerevisiae. Food Chemistry, 228, 550–559.
Ward, J. L., Harris, C., Lewis, J., & Beale, M. H. (2003). Assessment of 1H NMR spectroscopy and multivariate analysis as a technique for metabolite fingerprinting of Arabidopsis thaliana. Phytochemistry, 62, 949–957.
Zara, G., Ciani, M., Domizio, P., Zara, S., Budroni, M., Carboni, A., et al. (2014). A culture-independent PCR-based method for the detection of Lachancea thermotolerans in wine. Annals of Microbiology, 64, 403–406.
Zhou, Q., Qian, Y., & Qian, M. C. (2015). Analysis of volatile phenols in alcoholic beverage by ethylene glycol-polydimethylsiloxane based stir bar sorptive extraction and gas chromatography-mass spectrometry. Journal of Chromatography A, 1390, 22–27.
Acknowledgements
The authors would like to thank Janne Margrethe Benjaminsen for excellent technical assistance.
Funding
This study was funded by Faculty of Science, University of Copenhagen and Chinese Scholarship Council (201406300048).
Author information
Authors and Affiliations
Contributions
NA, CP and TV conceived and designed research. CP conducted experiments. MAP and FHL contributed new reagents and analytical tools. CP and FHL analyzed data. CP and NA wrote the manuscript. All authors read, revised and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
Chuantao Peng, Tiago Viana, Mikael Agerlin Petersen, Flemming Hofmann Larsen and Nils Arneborg declares that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human or animal subjects performed by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Peng, C., Viana, T., Petersen, M.A. et al. Metabolic footprint analysis of metabolites that discriminate single and mixed yeast cultures at two key time-points during mixed culture alcoholic fermentations. Metabolomics 14, 93 (2018). https://doi.org/10.1007/s11306-018-1391-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-018-1391-3