Abstract
Metabolomics datasets are commonly acquired by either mass spectrometry (MS) or nuclear magnetic resonance spectroscopy (NMR), despite their fundamental complementarity. In fact, combining MS and NMR datasets greatly improves the coverage of the metabolome and enhances the accuracy of metabolite identification, providing a detailed and high-throughput analysis of metabolic changes due to disease, drug treatment, or a variety of other environmental stimuli. Ideally, a single metabolomics sample would be simultaneously used for both MS and NMR analyses, minimizing the potential for variability between the two datasets. This necessitates the optimization of sample preparation, data collection and data handling protocols to effectively integrate direct-infusion MS data with one-dimensional (1D) 1H NMR spectra. To achieve this goal, we report for the first time the optimization of (i) metabolomics sample preparation for dual analysis by NMR and MS, (ii) high throughput, positive-ion direct infusion electrospray ionization mass spectrometry (DI-ESI–MS) for the analysis of complex metabolite mixtures, and (iii) data handling protocols to simultaneously analyze DI-ESI–MS and 1D 1H NMR spectral data using multiblock bilinear factorizations, namely multiblock principal component analysis (MB-PCA) and multiblock partial least squares (MB-PLS). Finally, we demonstrate the combined use of backscaled loadings, accurate mass measurements and tandem MS experiments to identify metabolites significantly contributing to class separation in MB-PLS-DA scores. We show that integration of NMR and DI-ESI–MS datasets yields a substantial improvement in the analysis of metabolome alterations induced by neurotoxin treatment.
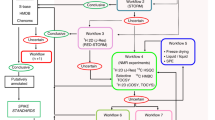
Graphical abstract





Similar content being viewed by others
References
Annesley, T. M. (2003). Ion suppression in mass spectrometry. Clinical Chemistry, 49, 1041–1044. doi:10.1373/49.7.1041.
Atherton, H. J., et al. (2006). A combined 1H-NMR spectroscopy- and mass spectrometry-based metabolomic study of the PPAR-α null mutant mouse defines profound systemic changes in metabolism linked to the metabolic syndrome. Physiological Genomics, 27, 178–186. doi:10.1152/physiolgenomics.00060.2006.
Barding, G. A., Beni, S., Fukao, T., Bailey-Serres, J., & Larive, C. K. (2013). Comparison of GC-MS and NMR for metabolite profiling of rice subjected to submergence stress. Journal of Proteome Research, 12, 898–909. doi:10.1021/pr300953k.
Beltran, A., et al. (2012). Assessment of compatibility between extraction methods for NMR- and LC/MS-based metabolomics. Analytical Chemistry, 84, 5838–5844. doi:10.1021/ac3005567.
Bove, J., Prou, D., Perier, C., & Przedborski, S. (2005). Toxin-induced models of Parkinson’s disease. NeuroRx, 2, 484–494.
Canelas, A. B., et al. (2009). Quantitative evaluation of intracellular metabolite extraction techniques for yeast metabolomics. Analytical Chemistry, 81, 7379–7389. doi:10.1021/ac900999t.
Chen, H., Pan, Z., Talaty, N., Raftery, D., & Cooks, R. G. (2006). Combining desorption electrospray ionization mass spectrometry and nuclear magnetic resonance for differential metabolomics without sample preparation. Rapid Communications in Mass Spectrometry, 20, 1577–1584. doi:10.1002/rcm.2474.
Cloarec, O., et al. (2005a). Statistical total correlation spectroscopy: An exploratory approach for latent biomarker identification from metabolic H-1 NMR data sets. Analytical Chemistry, 77, 1282–1289. doi:10.1021/Ac048630x.
Cloarec, O., et al. (2005b). Evaluation of the orthogonal projection on latent structure model limitations caused by chemical shift variability and improved visualization of biomarker changes in H-1 NMR spectroscopic metabonomic studies. Analytical Chemistry, 77, 517–526. doi:10.1021/Ac048803i.
Crockford, D. J., et al. (2006). Statistical heterospectroscopy, an approach to the integrated analysis of NMR and UPLC-MS data sets: Application in metabonomic toxicology studies. Analytical Chemistry, 78, 363–371. doi:10.1021/ac051444m.
Dai, H., Xiao, C., Liu, H., Hao, F., & Tang, H. (2010). Combined NMR and LC-DAD-MS analysis reveals comprehensive metabonomic variations for three phenotypic cultivars of Salvia miltiorrhiza bunge. Journal of Proteome Research, 9, 1565–1578. doi:10.1021/pr901045c.
De Meyer, T., et al. (2008). NMR-based characterization of metabolic alterations in hypertension using an adaptive, intelligent binning algorithm. Analytical Chemistry, 80, 3783–3790. doi:10.1021/Ac7025964.
Dettmer, K., Aronov, P. A., & Hammock, B. D. (2007). Mass spectrometry-based metabolomics. Mass Spectrometry Reviews, 26, 51–78. doi:10.1002/mas.20108.
Dieterle, F., Ross, A., Schlotterbeck, G., & Senn, H. (2006). Probabilistic quotient normalization as robust method to account for dilution of complex biological mixtures. Application in H-1 NMR metabonomics. Analytical Chemistry, 78, 4281–4290. doi:10.1021/Ac051632c.
Draper, J., Lloyd, A. J., Goodacre, R., & Beckmann, M. (2014). Flow infusion electrospray ionisation mass spectrometry for high throughput, non-targeted metabolite fingerprinting: A review. Metabolomics, 9, 4–29. doi:10.1007/s11306-012-0449-x.
Eriksson, L., Trygg, J., & Wold, S. (2008). CV-ANOVA for significance testing of PLS and OPLS (R) models. Journal of Chemometrics, 22, 594–600. doi:10.1002/Cem.1187.
Gu, H., Pan, Z., Xi, B., Asiago, V., Musselman, B., & Raftery, D. (2011). Principal component directed partial least squares analysis for combining nuclear magnetic resonance and mass spectrometry data in metabolomics: Application to the detection of breast cancer. Analytica Chimica Acta, 686, 57–63. doi:10.1016/j.aca.2010.11.040.
Halouska, S., & Powers, R. (2006). Negative impact of noise on the principal component analysis of NMR data. Journal of Magnetic Resonance, 178, 88–95. doi:10.1016/j.jmr.2005.08.016.
Jung, J.-Y., Jung, Y., Kim, J.-S., Ryu, D. H., & Hwang, G.-S. (2013). Assessment of peeling of Astragalus roots using 1H NMR- and UPLC-MS-based metabolite profiling. Journal of Agriculture and Food Chemistry, 61, 10398–10407. doi:10.1021/jf4026103.
Kamel, F., & Hoppin, J. A. (2004). Association of pesticide exposure with neurologic dysfunction and disease. Environmental Health Perspectives, 112, 950–958. doi:10.1289/ehp.7135.
Kanani, H., Chrysanthopoulos, P. K., & Klapa, M. I. (2008). Standardizing GC-MS metabolomics. Journal of Chromatography, B: Analytical Technologies in the Biomedical and Life Sciences, 871, 191–201. doi:10.1016/j.jchromb.2008.04.049.
Kell, D. B. (2004). Metabolomics and systems biology: Making sense of the soup. Current Opinion in Microbiology, 7, 296–307. doi:10.1016/j.mib.2004.04.012.
Kopka, J. (2006). Current challenges and developments in GC-MS based metabolite profiling technology. Journal of Biotechnology, 124, 312–322. doi:10.1016/j.jbiotec.2005.12.012.
Kuehnbaum, N. L., & Britz-McKibbin, P. (2013). New advances in separation science for metabolomics: Resolving chemical diversity in a post-genomic era. Chemical Reviews, 113, 2437–2468. doi:10.1021/cr300484s.
Lange, E., Tautenhahn, R., Neumann, S., & Gropl, C. (2008). Critical assessment of alignment procedures for LC-MS proteomics and metabolomics measurements. BMC Bioinformatics, 9, 375.
Lei, S., et al. (2014). Alterations in energy/redox metabolism induced by mitochondrial and environmental toxins: A specific role for glucose-6-phosphate-dehydrogenase and the pentose phosphate pathway in paraquat toxicity. ACS Chemical Biology. DOI:10.1021/cb400894a
Lenz, E. M., & Wilson, I. D. (2007). Analytical strategies in metabonomics. Journal of Proteome Research, 6, 443–458. doi:10.1021/pr0605217.
Lin, L., et al. (2010). Direct infusion mass spectrometry or liquid chromatography mass spectrometry for human metabonomics? A serum metabonomic study of kidney cancer. The Analyst, 135, 2970. doi:10.1039/c0an00265h.
Metz, T. O., et al. (2008). High-resolution separations and improved ion production and transmission in metabolomics. TrAC, Trends in Analytical Chemistry, 27, 205–214. doi:10.1016/j.trac.2007.11.003.
Mullen, A. R., et al. (2012). Reductive carboxylation supports growth in tumour cells with defective mitochondria. Nature, 481, 385–388. doi:10.1038/nature10642.
Nicholson, J. K., Lindon, J. C., & Holmes, E. (1999). “Metabonomics”: Understanding the metabolic responses of living systems to pathophysiological stimuli via multivariate statistical analysis of biological NMR spectroscopic data. Xenobiotica, 29, 1181–1189.
Pan, Z., & Raftery, D. (2007). Comparing and combining NMR spectroscopy and mass spectrometry in metabolomics. Analytical and Bioanalytical Chemistry, 387, 525–527. doi:10.1007/s00216-006-0687-8.
Skazov, R. S., Nekrasov, Y. S., Kuklin, S. A., & Simenel, A. A. (2006). Influence of experimental conditions on electrospray ionization mass spectrometry of ferrocenylalkylazoles. European Journal of Mass Spectrometry, 12, 137–142. doi:10.1255/ejms.795.
Smilde, A. K., Westerhuis, J. A., & de Jong, S. (2003). A framework for sequential multiblock component methods. Journal of Chemometrics, 17, 323–337. doi:10.1002/cem.811.
Smith, C. A., et al. (2005). METLIN: a metabolite mass spectral database. Therapeutic Drug Monitoring, 27, 747–751.
Taylor, P. J. (2005). Matrix effects: The Achilles heel of quantitative high-performance liquid chromatography-electrospray-tandem mass spectrometry. Clinical Biochemistry, 38, 328–334. doi:10.1016/j.clinbiochem.2004.11.007.
t’Kindt, R., et al. (2010). Metabolomics to unveil and understand phenotypic diversity between pathogen populations. PLOS Neglected Tropical Diseases, 4, e904. DOI:10.1371/journal.pntd.0000904.
Westerhuis, J. A., & Coenegracht, P. M. J. (1997). Multivariate modelling of the pharmaceutical two-step process of wet granulation and tableting with multiblock partial least squares. Journal of Chemometrics, 11, 379–392. doi:10.1002/(SICI)1099-128X(199709/10)11:5<379::AID-CEM482>3.0.CO;2-8.
Westerhuis, J. A., Kourti, T., & Macgregor, J. F. (1998). Analysis of multiblock and hierarchical PCA and PLS models. Journal of Chemometrics, 12, 301–321. doi:10.1002/(SICI)1099-128X(199809/10)12:5<301::AID-CEM515>3.0.CO;2-S.
Wishart, D. S., et al. (2007). HMDB: The human metabolome database. Nucleic Acids Research, 35, D521–D526.
Wishart, D. S., et al. (2009). HMDB: A knowledgebase for the human metabolome. Nucleic Acids Research, 37, D603–D610. doi:10.1093/nar/gkn810.
Wishart, D. S., et al. (2013). HMDB 3.0—the human metabolome database in 2013. Nucleic Acids Research, 41, D801–D807. doi:10.1093/nar/gks1065.
Wold, S. (1987). PLS modeling with latent variables in two or more dimensions. In Proceedings of PLS Model Building: Theory and Applications. Symposium Frankfurt am Main, September 23–25, 1987.
Worley, B., Halouska, S., & Powers, R. (2013). Utilities for quantifying separation in PCA/PLS-DA scores plots. Analytical Biochemistry, 433, 102–104. doi:10.1016/j.ab.2012.10.011.
Worley, B., & Powers, R. (2013). Multivariate analysis in metabolomics. Current Metabolomics, 1, 92–107. doi:10.2174/2213235x11301010092.
Worley, B., & Powers, R. (2014a). MVAPACK: A complete data handling package for NMR metabolomics. ACS Chemical Biology, 9, 1138–1144. doi:10.1021/cb4008937.
Worley, B., & Powers, R. (2014b). Simultaneous phase and scatter correction for NMR datasets. Chemometrics and Intelligent Laboratory Systems, 131, 1–6. doi:10.1016/j.chemolab.2013.11.005.
Xu, Y., Correa, E., & Goodacre, R. (2013). Integrating multiple analytical platforms and chemometrics for comprehensive metabolic profiling: Application to meat spoilage detection. Analytical and Bioanalytical Chemistry, 405, 5063–5074. doi:10.1007/s00216-013-6884-3.
Xu, F., Zou, L., & Ong, C. N. (2009). Multiorigination of chromatographic peaks in derivatized GC/MS metabolomics: A confounder that influences metabolic pathway interpretation. Journal of Proteome Research, 8, 5657–5665. doi:10.1021/pr900738b.
Zhang, B., Halouska, S., Schiaffo, C. E., Sadykov, M. R., Somerville, G. A., & Powers, R. (2011). NMR analysis of a stress response metabolic signaling network. Journal of Proteome Research, 10, 3743–3754. doi:10.1021/pr200360w.
Zhang, B., et al. (2013). Revisiting protocols for the NMR analysis of bacterial metabolomes. Journal of Integrated OMICS, 2, 120–137.
Acknowledgments
We would like to thank Dr. Jiantao Guo for providing us with the Escherichia coli Mach1 cell line. This manuscript was supported in part by funds from the National Institute of Health (R01 AI087668, R21 AI087561, R01 CA163649, P20 RR-17675, P30 GM103335), the University of Nebraska, the Nebraska Tobacco Settlement Biomedical Research Development Fund, and the Nebraska Research Council. The research was performed in facilities renovated with support from the National Institutes of Health (RR015468-01).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Darrell D. Marshall, Shulei Lei and Bradley Worley have equally contributed to this study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Marshall, D.D., Lei, S., Worley, B. et al. Combining DI-ESI–MS and NMR datasets for metabolic profiling. Metabolomics 11, 391–402 (2015). https://doi.org/10.1007/s11306-014-0704-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11306-014-0704-4