Abstract
Hyphal anastomosis testing and molecular methods have been the primary criteria employed to understand the evolutionary and taxonomic relationships of the soil-borne fungal plant pathogen Rhizoctonia solani species complex. In this study, a metabolomics-based approach for characterizing and identifying isolates of R. solani using gas chromatography/mass spectrometry (GC/MS) metabolite profiling and footprinting was developed. Multivariate and hierarchical cluster analyses of GC/MS data provided resolution of isolates belonging to anastomosis groups (AGs) 1–6, 9, and 10 of R. solani. Clustering of R. solani AG-3 isolates, based on host origin, was also observed and attributed to metabolite-biomarkers belonging to amino, carboxylic and fatty acids. The chemotaxonomic approach using metabolomics is a high-throughput methodology that complements existing molecular approaches for the taxonomic investigation of Rhizoctonia isolates and monitoring of fungal metabolism.




Similar content being viewed by others
References
Aliferis, K. A., & Jabaji, S. (2010a). 1H NMR and GC-MS metabolic fingerprinting of developmental stages of Rhizoctonia solani sclerotia. Metabolomics, 6, 96–108.
Aliferis, K. A., & Jabaji, S. (2010b). Metabolite composition and bioactivity of Rhizoctonia solani sclerotial exudates. Journal of Agriculture and Food Chemistry, 58, 7604–7615.
Allen, J., Davey, H. M., Broadhurst, D., et al. (2003). High-throughput classification of yeast mutants for functional genomics using metabolic footprinting. Nature Biotechnology, 21, 692–696.
Aoki, H., Sassa, T., & Tamura, T. (1963). Phytotoxic metabolites of Rhizoctonia solani. Nature, 200, 575.
Bartz, F., Danehower, D., Glassbrook, N., & Cubeta, M. (2010). The role of quinic acid in modulating the phenylacetic acid metabolic complex and the disease causing activity of Rhizoctonia solani AG-3. Inoculum, 61, 41.
Carling, D. E., Kuninaga, S., & Brainard, K. A. (2002). Hyphal anastomosis reactions, rDNA-internal transcribed spacer, and virulence levels among subsets of Rhizoctonia solani anastomosis group-2 (AG 2) and AG-BI. Phytopathology, 92, 43–50.
Ceresini, P. C., Shew, H. D., Vilgalys, R. J., et al. (2002). Genetic diversity of Rhizoctonia solani AG-3 from potato and tobacco in North Carolina. Mycologia, 94, 437–449.
Christensen, D., Foged, C., Rosenkrands, I., Nielsen, H. M., Andersen, P., & Agger, E. M. (2007). Trehalose preserves DDA/TDB liposomes and their adjuvant effect during freeze-drying. Biochimica et Biophysica Acta, 1768, 2120–2129.
Cubeta, M. A., & Vilgalys, R. (1997). Population biology of the Rhizoctonia solani complex. Phytopathology, 87, 480–484.
Ding, L., Qin, S., Li, F., Chi, X., & Laatsch, H. (2008). Isolation, antimicrobial activity, and metabolites of fungus Cladosporium sp. associated with red alga Porphyra yezoensis. Current Microbiology, 56, 229–235.
Efron, B., & Gong, G. (1983). A leisurely look at the bootstrap, the Jack-knife, and cross-validation. American Statistician, 37, 36–48.
Elbein, A. D. (1974). The metabolism of α,α-trehalose. In R.S. Tipson, & D. Horton (Eds.), Advances in carbohydrate chemistry and biochemistry (vol. 30, pp. 227–256) New York: Academic Press.
Eriksson, L., Johansson, E., Kettaneh-Wold, N., & Wold, S. (2001). Multi-and megavariate data analysis: Principles and applications. Umeå: Umetrics Academy.
Eriksson, L., Trygg, J., & Wold, S. (2009). PLS-trees®, a top-down clustering approach. Journal of Chemometrics, 23, 569–580.
Frisvad, J. C., Larsen, T. O., de Vries, O., et al. (2007). Secondary metabolite profiling, growth profiles and other tolls for species recognition and important Aspergillus mycotoxins. Studies in Mycology, 59, 31–37.
Frisvad, J. C., Andersen, B., & Thrane, U. (2008). The use of secondary metabolite profiling in chemotaxonomy of filamentous fungi. Mycological Research, 112, 231–240.
Gonzalez, D., Carling, D. E., Kuninaga, S., Vilgalys, R., & Cubeta, M. A. (2001). Ribosomal DNA systematics of Ceratobasidium and Thanatephorus with Rhizoctonia anamorphs. Mycologia, 93, 1138–1150.
Hettick, J. M., Green, B. J., Buskirk, A. D., et al. (2008). Discrimination of Aspergillus isolates at the species and strain level by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry fingerprinting. Analytical Biochemistry, 380, 276–281.
Himmelreich, U., Somorjai, R. L., Dolenko, B., et al. (2003). Rapid identification of Candida species by using nuclear magnetic resonance spectroscopy and a statistical classification strategy. Applied and Environmental Microbiology, 69, 4566–4574.
Hwang, B. K., Lim, S. W., Kim, B. S., Lee, J. Y., & Moon, S. S. (2001). Isolation and in vivo and in vitro antifungal activity of phenylacetic acid and sodium phenylacetate from Streptomyces humidus. Applied and Environmental Microbiology, 67, 3739–3745.
Iacobellis, N. S., & DeVay, J. E. (1987). Studies on pathogenesis of Rhizoctonia solani in beans: an evaluation of the possible role of phenylacetic acid and its hydroxy derivatives as phytotoxins. Physiology, Plant Pathology, 30, 421–432.
Iturriaga, G. (2008). The LEA proteins and trehalose loving couple: a step forward in anhydrobiotic engineering. Biochemical Journal, 410, e1–e2.
Jespersen, L., van der Kühle, A., & Petersen, K. M. (2000). Phenotypic and genetic diversity of Saccharomyces contaminants isolated from lager breweries and their phylogenetic relationship with brewing yeasts. Int. Journal of Food Microbiology, 60, 43–53.
Kadlec, Z., Šimek, P., Heydova, A., et al. (1994). Chemotaxonomic discrimination among the fungal genera Tolypocladium, Beauveria and Paecilomyces. Biochem. Systematics and Ecology, 22, 803–806.
Kell, D. B., Brown, M., Davey, H. M., Dunn, W. B., Spasic, I., & Oliver, S. G. (2005). Metabolic footprinting and systems biology: the medium is the message. Nature Reviews. Microbiology, 3, 557–565.
Kim, Y., Cho, J.-Y., Kuk, J.-H., et al. (2004). Identification and antimicrobial activity of phenylacetic acid produced by Bacillus licheniformis isolated from fermented soybean, Chungkook-Jang. Current Microbiology, 48, 312–317.
Kuninaga, S., Carling, D. E., Takeuchi, T., & Yokosawa, R. (2000). Comparison of rDNA-ITS sequences between potato and tabocco strains in Rhizoctonia solani AG-3. Journal of General Plant Pathology, 66, 2–11.
Laroche, J. P., Jabaji-Hare, S. H., & Charest, P. M. (1992). Differentiation of two anastomosis groups of Rhizoctonia solani by isozyme analysis. Phytopathology, 82, 1387–1393.
Larsen, T. O., Smedsgaard, J., Nielsen, K. F., Hansen, M. E., & Frisvad, J. (2005). Phenotypic taxonomy and metabolite profiling in microbial drug discovery. Natural Products Reports, 22, 672–695.
Mandava, N. B., Orellana, R. G., Warthen, J. D., Jr., et al. (1980). Phytotoxins in Rhizoctonia solani: isolation and biological activity of m-hydroxy- and m-methoxyphenylacetic acids. Journal of Agricultural and Food Chemistry, 28, 71–75.
Mas, S., Villas-Bôas, S. G., Hansen, M. E., Akesson, M., & Nielsen, J. (2007). A comparison of direct infusion MS and GC-MS for metabolic footprinting of yeast mutants. Biotechnology and Bioengineering, 96, 1014–1022.
Mendonça, A. L., Silva, C. E., Mesquita, F. L. T., et al. (2009). Antimicrobial activities of components of the glandular secretions of leaf cutting ants of the genus Atta. Antonie van Leeuwenhoek, 95, 295–303.
Mordue, J. E. M., Currah, R. S., & Bridge, P. D. (1989). An integrated approach to Rhizoctonia taxonomy: cultural, biochemical and numerical techniques. Mycological Research, 92, 78–90.
Nielsen, J., & Oliver, S. (2005). The next wave in metabolome analysis. Trends in Biotechnology, 23, 544–546.
Ogoshi, A. (1996). Introduction: The genus Rhizoctonia. In B. Sneh, S. Jabaji-Hare, S. Neate, & G. Dijst (Eds.), Rhizoctonia species: Taxonomy, molecular biology ecology, pathology and disease control (pp. 1–9). Dordrecht: Kluwer Academic Publishers.
Pope, G. A., MacKenzie, D. A., Defernez, M., et al. (2007). Metabolic footprinting as a tool for discriminating between brewing yeasts. Yeast, 24, 667–679.
Sharon, M., Kuninaga, S., Hyakumachi, M., & Sneh, B. (2006). The advancing identification and classification of Rhizoctonia spp. using molecular and biotechnological methods compared with the classical anastomosis grouping. Mycoscience, 47, 299–316.
Siefert, K. (2009). Progress towards DNA barcoding of fungi. Molecular Ecology Resources, 9, 83–89.
Smedsgaard, J., & Nielsen, J. (2004). Metabolite profiling of fungi and yeast: from phenotype to metabolome by MS and informatics. Journal of Experimental Botany, 56, 273–286.
Smedsgaard, J., Hansen, M. E., & Frisvad, J. C. (2004). Classification of terverticillate Penicillia by electrospray mass spectrometric profiling. Studies in Mycology, 49, 243–251.
Sneh, B. (1996). Anastomosis groups of multinucleate Rhizoctonia spp. In B. Sneh, S. Jabaji-Hare, S. Neate, & G. Dijst (Eds.), Rhizoctonia species: TAXONOMY, molecular biology, ecology, pathology and disease control (pp. 67–75). Dordrecht: Kluwer Academic Publishers.
Sonjak, S., Frisvad, J. C., & Gunde-Cimerman, N. (2009). Fingerprinting using extrolite profiles and physiological data shows sub-specific groupings of Penicillium crustosum strains. Mycological Research, 113, 836–841.
Sumner, L. W., Amberg, A., Barret, D., et al. (2007). Proposed minimum reporting standards for chemical analysis. Metabolomics, 3, 211–221.
Trygg, J., & Wold, S. (2002). Orthogonal projections to latent structures (OPLS). Journal of Chemometrics, 16, 119–128.
Ward, J. H. (1963). Hierarchical grouping to optimize an objective function. Journal of American Statistical Association, 58, 236–244.
Acknowledgments
Funding was provided by the Natural Sciences and Engineering Research Council of Canada (NSERC) discovery Grant to S. Jabaji.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
11306_2011_340_MOESM1_ESM.tif
Supplementary Fig. 1 Representative GC/MS total ion chromatograms of metabolite profiles of Rhizoctonia solani anastomosis groups (AGs) (TIFF 2985 kb)
11306_2011_340_MOESM2_ESM.tif
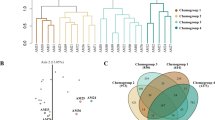
Supplementary Fig. 2 Orthogonal partial least squares-discriminant analysis (OPLS-DA) PC1/PC2 score plot (a) and corresponding OPLS-dendrogram (b) for R. solani AG-1 subgroups 1A and 1C. Metabolites contributing to the observed separation are displayed in the PLS coefficient plots (c) with values of scaled and centered PLS regression coefficients (CoeffCS) for the selected Y variables. The most influential variables to the observed separation are displayed with 95% jack-knifed confidence intervals. The ellipse in the score plots represents the Hotelling T2 with 95% confidence interval. Cluster distances of OPLS-dendrograms were calculated using the Ward’s method (TIFF 1175 kb)
11306_2011_340_MOESM3_ESM.tif
Supplementary Fig. 3 Orthogonal partial least squares-discriminant analysis (OPLS-DA) PC1/PC2 score plot (a) and corresponding OPLS-dendrogram (b) for R. solani AG-2 subgroups 2IIIB and 2IV. Metabolites contributing to the observed separation are displayed in the PLS coefficient plots (c) with values of scaled and centered PLS regression coefficients (CoeffCS) for the selected Y variables. The most influential variables to the observed separation are displayed with 95% jack-knifed confidence intervals. The ellipse in the score plots represents the Hotelling T2 with 95% confidence interval. Cluster distances of OPLS-dendrograms were calculated using the Ward’s method (TIFF 1239 kb)
11306_2011_340_MOESM4_ESM.tif
Supplementary Fig. 4 Representative GC/MS total ion chromatograms of metabolite profiles of R. solani AG-3 isolates (TIFF 3158 kb)
11306_2011_340_MOESM5_ESM.tif
Supplementary Fig. 5 OPLS-dendrograms for the prediction of R. solani AG-3 isolates based on the host. The isolates SJH-12 (a), ST-41 (b), DP-216 (c), QSU-028 (d), and RS-138 (e) were correctly classified by the model as potato isolates, whereas Bs-69 (f) was incorrectly classified as a tomato isolate. Isolates T1 (g) and T2 (h) isolates were correctly classified as tobacco isolates when used for validation. Cluster distances were calculated using the Ward’s method (TIFF 1527 kb)
Rights and permissions
About this article
Cite this article
Aliferis, K.A., Cubeta, M.A. & Jabaji, S. Chemotaxonomy of fungi in the Rhizoctonia solani species complex performing GC/MS metabolite profiling. Metabolomics 9 (Suppl 1), 159–169 (2013). https://doi.org/10.1007/s11306-011-0340-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11306-011-0340-1