Abstract
Azospirillum alphaproteobacteria, which live in the rhizosphere of many crops, are used widely as biofertilizers. Two-component signal transduction systems (TCSs) mediate the bacterial perception of signals and the corresponding adjustment of behavior facilitating the adaptation of bacteria to their habitats. In this study, we obtained the A. baldaniorum Sp245 mutant for the AZOBR_150176 gene, which encodes the TCS of the hybrid histidine kinase/response sensory regulator (HSHK-RR). Inactivation of this gene affected bacterial morphology and motility. In mutant Sp245-HSHKΔRR-Km, the cells were still able to synthesize a functioning polar flagellum (Fla), were shorter than those of strain Sp245, and were impaired in aerotaxis, elaboration of inducible lateral flagella (Laf), and motility in semiliquid media. The mutant showed decreased transcription of the genes encoding the proteins of the secretion apparatus, which ensures the assembly of Laf, Laf flagellin, and the repressor protein of translation of the Laf flagellin’s mRNA. The study examined the effects of polyethylene glycol 6000 (PEG 6000), an agent used to simulate osmotic stress and drought conditions. Under osmotic stress, the mutant was no longer able to use collective motility in semiliquid media but formed more biofilm biomass than did strain Sp245. Introduction into mutant cells of the AZOBR_150176 gene as part of an expression vector led to recovery of the lost traits, including those mediating bacterial motility under mechanical stress induced by increased medium density. The results suggest that the HSHK–RR under study modulates the response of A. baldaniorum Sp245 to mechanical and osmotic/water stress.






Similar content being viewed by others
Data Availability
All data generated or analyzed during this study are included in this published article (and its supplementary information files).
Abbreviations
- ANOVA:
-
Analysis of Variance
- CDS:
-
Coding Sequence
- CFU:
-
Colony-Forming Units
- Fla:
-
Polar Flagellum
- HSHK-RR:
-
Hybrid Sensor Histidine Kinase/Response Regulator
- HS-Taq PCR:
-
Hot Start-Taq Polymerase Chain Reaction
- Gm:
-
Gentamicin
- Km:
-
Kanamycin
- Laf:
-
Lateral Flagella
- LB:
-
Luria Bertani
- LSD:
-
Least Significant Difference
- MSM:
-
Malate Salt Medium
- PAS:
-
Per-ARNT-Sim
- PEG 6000:
-
Polyethylene Glycol 6000
- PB:
-
Phosphate Buffer
- PCR:
-
Polymerase Chain Reaction
- RT-PCR:
-
Reverse Transcription Polymerase Chain Reaction
- RTL:
-
Relative Transcription Level
- SDS-PAGE:
-
Sodium Dodecyl Sulfate–Polyacrylamide Gel
- Tc:
-
Tetracycline
- TCS:
-
Two-Component Signal Transduction System
- qRT-PCR:
-
Real-Time Quantitative PCR
References
Alexandre G, Greer SE, Zhulin IB (2000) Energy taxis is the dominant behavior in Azospirillum brasilense. J Bacteriol 182:6042–6048. https://doi.org/10.1128/jb.182.21.6042-6048.2000
Ansari FA, Jabeen M, Ahmad I (2021) Pseudomonas azotoformans FAP5, a novel biofilm-forming PGPR strain, alleviates drought stress in wheat plant. Int J Environ Sci Technol 18:3855–3870. https://doi.org/10.1007/s13762-020-03045-9
Appleby JL, Parkinson JS, Bourret RB (1996) Signal transduction via the multi-step phosphorelay: not necessarily a road less traveled. Cell 86:845–848. https://doi.org/10.1016/S0092-8674(00)80158-0
Baldani VLD, Baldani JI, Dobereiner J (1983) Effects of Azospirillum inoculation on root infection and nitrogen incorporation in wheat. Can J Microbiol 29:924–929. https://doi.org/10.1139/m83-148
Bashan Y (1986) Migration of the rhizosphere bacteria Azospirillum brasilense and Pseudomonas fluorescens towards wheat roots in the soil. J Gen Microbiol 132:3407–3414. https://doi.org/10.1099/00221287-132-12-3407
Baymiev AK, Yamidanov RS, Matniyazov RT, Blagova DK, Baymiev AK, Chemeris AV (2011) Preparation of fluorescent labeled nodule bacteria strains of wild legumes for their detection in vivo and in vitro. Mol Biol 45:904–910. https://doi.org/10.1134/S0026893311060033
Bellefontaine AF, Pierreux CE, Mertens P, Vandenhaute J, Letesson J-J, De Bolleet X (2002) Plasticity of a transcriptional regulation network among alpha-proteobacteria is supported by the identification of CtrA targets in Brucella abortus. Mol Microbiol 43(4):945–960. https://doi.org/10.1046/j.1365-2958.2002.02777.x
Belyakov AY, Burygin GL, Arbatsky NP, Shashkov AS, Selivanov NYu M, LYu KYA (2012) Shchyogolev SYu Identification of an O-linked repetitive glycan chain of the polar flagellum flagellin of Azospirillum brasilense Sp7. Carbohydr Res 1;361:127 – 32. https://doi.org/10.1016/j.carres.2012.08.019
Bible AN, Stephens BB, Ortega DR, Xie Z, Alexandre G (2008) Function of a chemotaxis-like signal transduction pathway in modulating motility, cell clumping, and cell length in the alphaproteobacterium Azospirillum brasilense. J Bacteriol 190:6365–6375. https://doi.org/10.1128/jb.00734-08
Biondi EG, Reisinger SJ, Skerker JM, Arif M, Perchuk BS, Ryan KR, Laub MT (2006) Regulation of the bacterial cell cycle by an integrated genetic circuit. Nature 444:899–904. https://doi.org/10.1038/nature05321
Borisov IV, Schelud’ko AV, Petrova LP, Katsy EI (2009) Changes in Azospirillum brasilense motility and the effect of wheat seedling exudates. Microb Res 164(5):578–587. https://doi.org/10.1016/j.micres.2007.07.003
Borland S, Oudart A, Prigent-Combaret C, Brochier-Armanet C, Wisniewski-Dyé F (2015) Genome-wide survey of two-component signal transduction systems in the plant growth-promoting bacterium Azospirillum. BMC Genomics 16:833. https://doi.org/10.1186/s12864-015-1962-x
Burygin GL, Shirokov AA, Shelud’ko AV, Katsy EI, Shchegolev SYu M LYu (2007) Detection of a surface sheath on Azospirillum brasilense polar flagellum. Microbiology 76(6):822–829. https://doi.org/10.1134/S0026261707060124
Chawla R, Gupta R, Lele TP, Lele PP (2020) A skeptic’s guide to bacterial mechanosensing. J Mol Biol 432(2):523–533. https://doi.org/10.1016/j.jmb.2019.09.004
Chutia J, Borah SP (2012) Water stress effects on leaf growth and chlorophyll content but not the grain yield in traditional rice (Oryza sativa L.) genotypes of Assam, India: II. Protein and proline status in seedlings under PEG induced water stress. Am J Plant Sci 3:971–980. https://doi.org/10.4236/ajps.2012.37115
Croes CL, Moens S, van Bastelaere E, Vanderleyden J, Michiels K (1993) The polar flagellum mediated Azospirillum brasilense adsorption to wheat roots. J Gen Microbiol 139:2261–2269. https://doi.org/10.1099/00221287-139-9-2261
Dubey AP, Pandey P, Singh VS, Mishra MN, Singh S, Mishra R, Tripathi AK (2020) An ECF41 family σ factor controls motility and biogenesis of lateral flagella in Azospirillum brasilense sp245. J Bacteriol 202(16):e00231–e00220. https://doi.org/10.1128/jb.00231-20
Errington J, Daniel RA, Scheffers DJ (2003) Cytokinesis in bacteria. Microbiol Mol Biol Rev 67:52–65. https://doi.org/10.1128/mmbr.67.1.52-65.2003
Ferreira NS, Sant’Anna FH, Reis VM, Ambrosini A, Volpiano CG, Rothballer M, Schwab S, Baura VA, Balsanelli E, Pedrosa FO et al (2020) Genome-based reclassification of Azospirillum brasilense Sp245 as the type strain of Azospirillum baldaniorum sp. nov. Int J Syst Evol Microbiol 70(12):6203–6212. https://doi.org/10.1099/ijsem.0.004517
Figurski DH, Helinski DR (1979) Replication of an origin-containing derivative of plasmid RK2 dependent on a plasmid function provided in trans. Proc Natl Acad Sci USA 76:1648–1652. https://doi.org/10.1073/pnas.76.4.1648
Filip’echeva YA, Shelud’ko AV, Prilipov AG, Burygin GL, Telesheva EM, Yevstigneyeva SS, Chernyshova MP, Petrova LP, Katsy EI (2018a) Plasmid AZOBR_p1-borne fabG gene for putative 3-oxoacyl-[acyl-carrier protein] reductase is essential for proper assembly and work of the dual flagellar system in the alphaproteobacterium Azospirillum brasilense Sp245. Can J Microbiol 64(2):107–118. https://doi.org/10.1139/cjm-2017-0561
Filip’echeva Y, Shelud’ko A, Prilipov A, Telesheva E, Mokeev D, Burov A, Petrova L, Katsy E (2018b) Chromosomal flhB1 gene of the alphaproteobacterium Azospirillum brasilense Sp245 is essential for correct assembly of both constitutive polar flagellum and inducible lateral flagella. Folia Microbiol 63:147–153. https://doi.org/10.1007/s12223-017-0543-6
Frolov A, Bilova T, Paudel G, Berger R, Balcke GU, Birkemeyer C, Wessjohann LA (2017) Early responses of mature Arabidopsis thaliana plants to reduced water potential in the agar-based polyethylene glycol infusion drought model. J Plant Physiol 208:70–83. https://doi.org/10.1016/j.jplph.2016.09.013
Fukami J, Cerezini P, Hungria M (2018) Azospirillum: benefits that go far beyond biological nitrogen fixation. AMB Expr 8:73–85. https://doi.org/10.1186/s13568-018-0608-1
Galperin MY (2005) A census of membrane-bound and intracellular signal transduction proteins in bacteria: bacterial IQ, extroverts and introverts. BMC Microbiol 5:35. https://doi.org/10.1186/1471-2180-5-35
Ganusova EE, Vo LT, Mukherjee T, Alexandre G (2021) Multiple CheY proteins control Surface-Associated Lifestyles of Azospirillum brasilense. Front Microbiol 12:664826. https://doi.org/10.3389/fmicb.2021.664826
Gavín R, Rabaan AA, Merino S, Tomás JM, Gryllos I, Shaw JG (2002) Lateral flagella of Aeromonas species are essential for epithelial cell adherence and biofilm formation. Mol Microbiol 43(2):383–397. https://doi.org/10.1046/j.1365-2958.2002.02750.x
Gullett JM, Bible A, Alexandre G (2017) Distinct domains of CheA confer unique functions in chemotaxis and cell length in Azospirillum brasilense Sp7. J Bacteriol 199(13):e00189–e00117. https://doi.org/10.1128/jb.00189-17
Haldar S, Sengupta S (2015) Plant-microbe cross-talk in the Rhizosphere: insight and biotechnological potential. The Open Microbiol J 9:1–7. https://doi.org/10.2174/1874285801509010001
Harshey RM, Partridge JD (2015) Shelter in a swarm. J Mol Biol 427:3683–3694. https://doi.org/10.1016/j.jmb.2015.07.025
Hendriksen NB (2022) Microbial biostimulants – the need for clarification in EU regulation /. Trends in Microbiol 30:311–313. https://doi.org/10.1016/j.tim.2022.01.008
Hoang TT, Karkhoff-Schweizer RR, Kutchma AJ, Schweizer HP (1998) A broad-host-range Flp-FRT recombination system for site-specific excision of chromosomally-located DNA sequences: application for isolation of unmarked Pseudomonas aeruginosa mutants. Gene 212(1):77–86. https://doi.org/10.1016/S0378-1119(98)00130-9
Ishii E, Eguchi Y (2021) Diversity in sensing and signaling of bacterial sensor histidine kinases. Biomolecules 11(10):1524. https://doi.org/10.3390/biom11101524
Jacob-Dubuisson F, Mechaly A, Betton JM, Antoine R (2018) Structural insights into the signalling mechanisms of two-component systems. Nat Rev Microbiol 16:585–593. https://doi.org/10.1038/s41579-018-0055-7
Keen NT, Tamaki S, Kobayashi D, Trollinger D (1988) Improved broad-host-range plasmids for DNA cloning in gram-negative bacteria. Gene 70:191–197. https://doi.org/10.1016/0378-1119(88)90117-5
Kim W, Killam T, Sood V, Surette MG (2003) Swarm-cell differentiation in Salmonella enterica serovar typhimurium results in elevated resistance to multiple antibiotics. J Bacteriol 185:3111–3117. https://doi.org/10.1128/jb.185.10.3111-3117.2003
Kumar S, Rai AK, Mishra MN, Shukla M, Singh PK, Tripathi AK (2012) RpoH2 sigma factor controls the photooxidative stress response in a nonphotosynthetic rhizobacterium, Azospirillum brasilense Sp7. Microbiol 158:2891–2902. https://doi.org/10.1099/mic.0.062380-0
Laub MT, Goulian M (2007) Specificity in two-component signal transduction pathways. Annu Rev Genet 41:121–145. https://doi.org/10.1146/annurev.genet.41.042007.170548
Lipa P, Janczarek M (2020) Phosphorylation systems in symbiotic nitrogen-fixing bacteria and their role in bacterial adaptation to various environmental stresses. Peer J 8:e8466. https://doi.org/10.7717/peerj.8466
Little K, Austerman J, Zheng J, Gibbs KA (2019) Cell shape and population migration are distinct steps of Proteus mirabilis swarming that are decoupled on high-percentage agar. J Bacteriol 201:e00726–e00718. https://doi.org/10.1128/jb.00726-18
Liu B, Yang F, Huang DS, Chou KC (2018) iPromoter-2L: a two-layer predictor for identifying promoters and their types by multi-window-based PseKNC. Bioinformatics 34:33–40. https://doi.org/10.1093/bioinformatics/btx579
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)). Method Methods 25(4):402–408. https://doi.org/10.1006/meth.2001.1262
McCarter L, Hilmen M, Silverman M (1988) Flagellar dynamometer controls swarmer cell differentiation of V. parahaemolyticus. Cell 54:345–351. https://doi.org/10.1016/0092-8674(88)90197-3
Moens S, Michiels K, Vanderleyden J (1995) Glycosylation of the flagellin of the polar flagellum of Azospirillum brasilense, a Gram-negative nitrogen-fixing bacterium. Microbiol 141(10):2651–2657. https://doi.org/10.1099/13500872-141-10-2651
Moens S, Schloter M, Vanderleyden J (1996) Expression of the structural gene, laf1, encoding the flagellin of the lateral flagella in Azospirillum brasilense Sp7. J Bacteriol 178:5017–5019. https://doi.org/10.1128/jb.178.16.5017-5019.1996
Mongiardini EJ, Quelas JI, Dardis C, Althabegoiti MJ, Lodeir AR (2017) Transcriptional control of the lateral-flagellar genes of Bradyrhizobium diazoefficiens. J Bacteriol 199:e00253–e00217. https://doi.org/10.1128/jb.00253-17
Oliveira ALM, Canuto E, Silva EE, Veronica MR, Baldani JI (2004) Survival of endophytic diazotrophic bacteria in soil under different moisture intensities. Braz J Microbiol 35:295–299. https://doi.org/10.1590/S1517-83822004000300005
O’Toole GA, Kolter R (1998) Initiation of biofilm formation in Pseudomonas fluorescens WCS365 proceeds via multiple, convergent signalling pathways: a genetic analysis. Mol Microbiol 28(3):449–461. https://doi.org/10.1046/j.1365-2958.1998.00797.x
Petrova LP, Yevstigneyeva SS, Borisov IV, Shelud’ko AV, Burygin GL, Katsy EI (2020) Plasmid gene AZOBR_p60126 impacts biosynthesis of lipopolysaccharide II and swarming motility in Azospirillum brasilense Sp245. J Basic Microbiol 60:613–623. https://doi.org/10.1002/jobm.201900635
Rotter C, Mühlbacher S, Salamon D, Schmitt R, Scharf B (2006) Rem, a new transcriptional activator of motility and chemotaxis in Sinorhizobium meliloti. J Bacteriol 188:6932–6942. https://doi.org/10.1128/jb.01902-05
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular Cloning: a Laboratory Manual, 2nd edn. Cold Spring Harbor Laboratory Press, New York
Scheludko AV, Katsy EI, Ostudin NA, Gringauz OK, Panasenko VI (1998) Novel classes of Azospirillum brasilense mutants with defects in the assembly and functioning of polar and lateral flagella. Mol Gen Mikrobiol Virusol (4): 33–37
Schelud’ko AV, Makrushin KV, Tugarova AV, Krestinenko VA, Panasenko VI, Antonyuk LP, Katsy EI (2009a) Changes in motility of the rhizobacterium Azospirillum brasilense in the presence of plant lectins. Microbiol Res 164(2):149–156. https://doi.org/10.1016/j.micres.2006.11.008
Schloter M, ,Moens S, Croes C, Reidel G, Esquenet M, Hartmann A, Michieis K (1994) Characterization of cell surface components of Azospirillum brasilense Sp7 as antigenic determinants for strain-specific monoclonal antibodies. Microbiol 140:823–828. https://doi.org/10.1089/hyb.1997.16.183
Shelud’ko AV, Filip’echeva YA, Telesheva EM, Yevstigneeva SS, Petrova LP, Katsy EI (2019) Polar flagellum of the alphaproteobacterium Azospirillum brasilense Sp245 plays a role in biofilm biomass accumulation and in biofilm maintenance under stationary and dynamic conditions. World J Microbiol Biotechnol 35(2):19. https://doi.org/10.1007/s11274-019-2594-0
Shelud’ko AV, Mokeev DI, Evstigneeva SS, Filip’echeva YuA, Burov AM, Petrova LP, Ponomareva EG, Katsy EI (2020) Cell ultrastructure in biofilms of Azospirillum brasilense. Microbiology (Moscow) 89: 50–63. https://doi.org/10.1134/S0026261720010142
Shelud’ko AV, Ponomareva EG, Varshalomidze OE, Vetchinkina EP, Katsy EI, Nikitina VE (2009b) Hemagglutinating activity and motility of the bacterium Azospirillum brasilense in the presence of various nitrogen sources. Microbiology 78:696–702. https://doi.org/10.1134/S0026261709060058
Shelud’ko AV, Shirokov AA, Sokolova MK, Sokolov OI, Petrova LP, Matora LYu, Katsy EI (2010) Wheat root colonization by Azospirillum brasilense strains with different motility. Microbiology 79:688–695. https://doi.org/10.1134/S0026261710050140
Shirokov A, Budanova A, Burygin G, Evseeva N, Matora L, Shchyogolev S (2020) Flagellin of polar flagellum from Azospirillum brasilense Sp245: isolation, structure, and biological activity. Int J Biol Macromol 147:1221–1227. https://doi.org/10.1016/j.ijbiomac.2019.10.092
Skerker JM, Laub MT (2004) Cell-cycle progression and the generation of asymmetry in Caulobacter crescentus. Nat Rev Microbiol 2(4):325–337. https://doi.org/10.1038/nrmicro864
Stephens BB, Loar SN, Alexandre G (2006) Role of CheB and CheR in the complex chemotactic and aerotactic pathway of Azospirillum brasilense. J Bacteriol 188:4759–4768. https://doi.org/10.1128/jb.00267-06
Tsang VCW, Peralta JM, Simons AR (1983) Enzyme-linked immunoelectrotransfer blot techniques (EITB) for studying the specificities of antigens and antibodies separated by gel electrophoresis Methods Enzymol 92: 377–391. https://doi.org/10.1016/0076-6879(83)92032-3
Ulrich LE, Koonin EV, Zhulin IB (2005) One-component systems dominate signal transduction in prokaryotes. Trends Microbiol 13:52–56. https://doi.org/10.1016/j.tim.2004.12.006
Vega-Baray B, Domenzain C, Rivera A, Alfaro-Lopez R, Gomez-Cesar E, Poggio S, Dreyfus G, Camarena L (2015) The flagellar set Fla2 in Rhodobacter sphaeroides is controlled by the CckA pathway and is repressed by organic acids and the expression of Fla1. J Bacteriol 197:833–847. https://doi.org/10.1128/jb.02429-14
Vieira J, Messing J (1982) The pUC plasmids, an M13mp7-derived system for insertion mutagenesis and sequencing with synthetic universal primers. Gene 19:259–268. https://doi.org/10.1016/0378-1119(82)90015-4
Viruega-Góngora VI, Acatitla-Jácome IS, Reyes-Carmona SR, Baca BE, Ramírez-Mata A (2020) Spatio-temporal formation of biofilms and extracellular matrix analysis in Azospirillum brasilense. FEMS Microbiol Lett 367:fnaa037. 10.1093
Willett JW, Herrou J, Briegel A, Rotskoff G, Crosson S (2015) Structural asymmetry in a conserved signaling system that regulates division, replication, and virulence of an intracellular pathogen. Proc Natl Acad Sci U S A 112(28):E3709–E3718. https://doi.org/10.1073/pnas.1503118112
Williams RH, Whitworth DE (2010) The genetic organisation of prokaryotic two-component system signalling pathways. BMC Genomics 11:720. https://doi.org/10.1186/1471-2164-11-720
Wisniewski-Dyé F, Borziak K, Khalsa-Moyers G, Alexandre G, Sukharnikov LO, Wuichet K, Hurst GB, McDonald WH, Robertson JS, Barbe V, Calteau A, Rouy Z, Mangenot S, Prigent-Combaret C, Normand P, Boyer M, Siguier P, Dessaux Y, Elmerich C, Condemine G, Krishnen G, Kennedy I, Paterson AH, González V, Mavingui P, Zhulin IB (2011) Azospirillum genomes reveal transition of bacteria from aquatic to terrestrial environments. PLoS Genet 7(12):e1002430. https://doi.org/10.1371/journal.pgen.1002430
Zhulin IB, Bespalov VA, Johnson MS, Taylor BL (1996) Oxygen taxis and proton motive force in Azospirillum brasilense. J Bacteriol 178:5199–5204. https://doi.org/10.1128/jb.178.17.5199-5204.1996
Acknowledgements
Our thanks go to Dmitry N. Tychinin for the translation of the original manuscript into English. We thank research center “Symbiosis” and immunochemistry laboratory IBPPM RAS for their support with immunofluorescence analysis and confocal microscopy.
Funding
This work was funded by the Russian Science Foundation (RSF) (project no. № 23-26-00271).
Author information
Authors and Affiliations
Contributions
All individuals listed as authors have made a substantive creative contribution to the work: AS supervised, wrote, reviewed, and edited the manuscript; IV, LP and LM contributed analyze, write, and edit the final manuscript; AS, IV, DM, ET, LP, AB, SY, AT, GB, AS and LYM performed research; AS and IV analyzed data; and all the authors read and approved the final submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflict of interest
The authors declare no real or perceived conflicts of interests.
Contributions
All individuals listed as authors have made a substantive creative contribution to the work: AS supervised, wrote, reviewed, and edited the manuscript; IV, LP and LM contributed analyze, write, and edit the final manuscript; AS, IV, DM, ET, LP, AB, SY, AT, GB, AS and LM performed research; AS and IV analyzed data; and all the authors read and approved the final submission.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig. 1
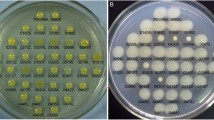
. A Average swimming rate of A. baldaniorum cells from 18–24-h-old cultures grown in a liquid MSM; B Light microscopy of the formation of a band of motile bacteria near the air–liquid interface. A suspension of cells from 18-h-old liquid cultures, which had been washed with 10 mM PB (pH 7.0), was placed in a 1-mm-diameter capillary tube, and the band formation time of motile bacteria in the oxygen gradient originating in the capillary liquid was recorded; C Trajectory of swimming cells from the liquid MSM. For blocks B and C, scale bars correspond to 10 μm; D Colonies formed by azospirilla within 48 h on MSM containing 0.2% Bacto agar and 1 mM succinic acid, galactose, fructose, aspartate, glutamate, or proline as the carbon source. Scale bars correspond to 10 mm
Supplementary Fig. 2
. A and B SDS–PAGE profiles of the extracellular proteins of strains Sp245 (2), Sp245.1062 (3), Sp245.1062(pRK415-150176) (4), and Sp245.1063 (1). (M): Standard protein marker (14.4–116-kDa) (Thermo Scientific, USA). Arrows indicate the positions of the flagellins of the polar (Fla) and lateral (Laf1) flagella. C Immunoblotting of total cellular proteins with antibodies interacting with the protein portion of the Sp245 Fla flagellin. Cells grown in liquid culture (A) and on a solid MSM (B and C) (18 and 36 h of growth, respectively) were used to isolate extracellular and total proteins
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shelud’ko, A., Volokhina, I., Mokeev, D. et al. Chromosomal gene of hybrid multisensor histidine kinase is involved in motility regulation in the rhizobacterium Azospirillum baldaniorum Sp245 under mechanical and water stress. World J Microbiol Biotechnol 39, 336 (2023). https://doi.org/10.1007/s11274-023-03785-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-023-03785-z