Abstract
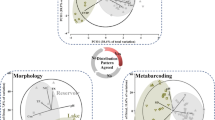
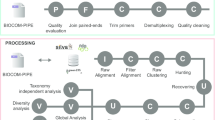
Metabarcoding using high throughput sequencing of amplicons of the 18S rRNA gene is one of the widely used methods for assessing the diversity of microeukaryotes in various ecosystems. We investigated the effectiveness of the V4 and V8-V9 regions of the 18S rRNA gene by comparing the results of metabarcoding microeukaryotic communities using the DADA2 (ASV), USEARCH-UNOISE3 (ZOTU), and USEARCH-UPARSE (OTU with 97% similarity) algorithms. Both regions showed similar levels of genetic variability and taxa identification accuracy. Richness for DADA2 datasets of both regions was lower than for UNOISE3 and UPARSE datasets, which is due to more accurate error correction in amplicons. Microeukaryotic communities (autotrophs and heterotrophs) structure identified using both regions showed a significant relationship with phytoplankton (autotrophs) communities structure based on microscopy in a seasonal freshwater sample series. The strongest relationship was found between the phytoplankton species and V8-V9 ASVs produced by DADA2.






Similar content being viewed by others
Data Availability
All authors certify that they have no affiliations with or involvement in any organization or entity with any financial interest or non-financial interest in the subject matter or materials discussed in this manuscript. The authors have no financial or proprietary interests in any material discussed in this article.
References
Adl SM, Bass D, Lane CE, Lukeš J, Schoch CL, Smirnov A et al (2019) Revisions to the classification, nomenclature, and diversity of eukaryotes. J Eukaryot Microbiol 66(1):4–119. https://doi.org/10.1111/jeu.12691
Annenkova NV, Giner CR, Logares R (2020) Tracing the origin of planktonic protists in an ancient lake. Microorganisms 8:543. https://doi.org/10.3390/microorganisms8040543
Badotti F, de Oliveira FS, Garcia CF, Vaz ABM, Fonseca PLC, Nahum LA, Oliveira G, Góes-Neto A (2017) Effectiveness of ITS and sub-regions as DNA barcode markers for the identification of Basidiomycota (Fungi). BMC Microbiol 17(1):1–12. https://doi.org/10.1186/s12866-017-0958-x
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B Methodol 57:289–300
Bik HM, Fournier D, Sung W, Bergeron RD, Thomas WK (2013) Intra-genomic variation in the ribosomal repeats of nematodes. PLoS ONE 8(10):e78230. https://doi.org/10.1371/journal.pone.0078230
Bondarenko NA, Ozersky T, Obolkina LA, Tikhonova IV, Sorokovikova EG, Sakirko MV et al (2019) Recent changes in the spring microplankton of Lake Baikal, Russia. Limnologica 75:19–29. https://doi.org/10.1016/j.limno.2019.01.002
Bradley IM, Pinto AJ, Guest JS (2016) Design and evaluation of Illumina MiSeq-compatible, 18S rRNA gene-specific primers for improved characterization of mixed phototrophic communities. Appl Environ Microb 82(19):5878–5891. https://doi.org/10.1128/AEM.01630-16
Bukin YS, Galachyants YP, Morozov IV, Bukin SV, Zakharenko AS, Zemskaya TI (2019) The effect of 16S rRNA region choice on bacterial community metabarcoding results. Sci Data 6:190007. https://doi.org/10.1038/sdata.2019.7
Burki F, Roger AJ, Brown MW, Simpson AG (2020) The new tree of eukaryotes. Trends Ecol Evol 35(1):43–55. https://doi.org/10.1016/j.tree.2019.08.008
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJ, Holmes SP (2016) DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods 13(7):581–583. https://doi.org/10.1038/nmeth.3869
Casey JM, Ransome E, Collins AG, Mahardini A, Kurniasih EM, Sembiring A et al (2021) DNA metabarcoding marker choice skews perception of marine eukaryotic biodiversity. Environ DNA 3(6):1229–1246. https://doi.org/10.1002/edn3.245
Chiu CH, Wang YT, Walther BA, Chao A (2014) An improved nonparametric lower bound of species richness via a modified good–turing frequency formula. Biometrics 70(3):671–682. https://doi.org/10.1111/biom.12200
Choi J, Park JS (2020) Comparative analyses of the V4 and V9 regions of 18S rDNA for the extant eukaryotic community using the Illumina platform. Sci Rep 10:6519. https://doi.org/10.1038/s41598-020-63561-z
Corell J, Rodríguez-Ezpeleta N (2014) Tuning of protocols and marker selection to evaluate the diversity of zooplankton using metabarcoding. Rev Invest 21:19–39
David GM, Moreira D, Reboul G, Annenkova NV, Galindo LJ, Bertolino P, López-Archilla AI, Jardillier L, López-García P (2020) Environmental drivers of plankton protist communities along latitudinal and vertical gradients in the oldest and deepest freshwater lake. Environ Microbiol 23:1436–1451. https://doi.org/10.1111/1462-2920.15346
De Santiago A, Pereira TJ, Mincks SL, Bik HM (2022) Dataset complexity impacts both MOTU delimitation and biodiversity estimates in eukaryotic 18S rRNA metabarcoding studies. Environ DNA 4(2):363–384. https://doi.org/10.1002/edn3.255
Deagle BE, Jarman SN, Coissac E, Pompanon F, Taberlet P (2014) DNA metabarcoding and the cytochrome c oxidase subunit I marker: not a perfect match. Biol lett 10(9):20140562. https://doi.org/10.1098/rsbl.2014.0562
Dixon P (2003) VEGAN, a package of R functions for community ecology. J Veg Sci 14(6):927–930. https://doi.org/10.1111/j.1654-1103.2003.tb02228.x
Dray S, Dufour AB (2007) The ade4 package: implementing the duality diagram for ecologists. J Stat Softw 22:1–20. https://doi.org/10.18637/jss.v022.i04
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10(10):996–998. https://doi.org/10.1038/nmeth.2604
Edgar RC (2016a) UNOISE2: improved error-correction for Illumina 16S and ITS amplicon sequencing. https://doi.org/10.1101/081257. BioRxiv 081257
Edgar RC (2016b) SINTAX: a simple non-bayesian taxonomy classifier for 16S and ITS sequences. https://doi.org/10.1101/074161. BioRxiv 074161
Faith DP, Minchin PR, Belbin L (1987) Compositional dissimilarity as a robust measure of ecological distance. Vegetatio 69(1):57–68. https://doi.org/10.1007/BF00038687
Ferrera I, Giner CR, Reñé A, Camp J, Massana R, Gasol JM, Garcés E (2016) Evaluation of alternative high-throughput sequencing methodologies for the monitoring of marine picoplanktonic biodiversity based on rRNA gene amplicons. Front Mar Sci 3:147. https://doi.org/10.3389/fmars.2016.00147
Fujisawa T, Barraclough TG (2013) Delimiting species using single-locus data and the generalized mixed Yule Coalescent approach: a revised method and evaluation on simulated data sets. Syst Biol 62(5):707–724. https://doi.org/10.1093/sysbio/syt033
Gao B, Chi L, Zhu Y, Shi X, Tu P, Li B et al (2021) An introduction to next generation sequencing bioinformatic analysis in gut microbiome studies. Biomolecules 11(4):530. https://doi.org/10.3390/biom11040530
Giebner H, Langen K, Bourlat SJ, Kukowka S, Mayer C, Astrin JJ et al (2020) Comparing diversity levels in environmental samples: DNA sequence capture and metabarcoding approaches using 18S and COI genes. Mol Ecol Resour 20(5):1333–1345. https://doi.org/10.1111/1755-0998.13201
Gong W, Marchetti A (2019) Estimation of 18S gene copy number in marine eukaryotic plankton using a next-generation sequencing approach. Front Mar Sci 6:219. https://doi.org/10.3389/fmars.2019.00219
Guillou L, Bachar D, Audic S, Bass D, Berney C, Bittner L et al (2012) The Protist Ribosomal reference database (PR2): a catalog of unicellular eukaryote small sub-unit rRNA sequences with curated taxonomy. Nucleic Acids Res 41(D1):D597–D604. https://doi.org/10.1093/nar/gks1160
Hadziavdic K, Lekang K, Lanzen A, Jonassen I, Thompson EM, Troedsson C (2014) Characterization of the 18S rRNA gene for designing universal eukaryote specific primers. PLoS ONE 9(2):e87624. https://doi.org/10.1371/journal.pone.0087624
Hill MO (1973) Diversity and evenness: a unifying notation and its consequences. Ecology 54(2):427–432. https://doi.org/10.2307/1934352
Hugerth LW, Muller EE, Hu YO, Lebrun LA, Roume H, Lundin D et al (2014) Systematic design of 18S rRNA gene primers for determining eukaryotic diversity in microbial consortia. PLoS ONE 9(4):e95567. https://doi.org/10.1371/journal.pone.0095567
Izmest’eva LR, Silow EA, Litchman E (2011) Long-term dynamics of Lake Baikal pelagic phytoplankton under climate change. Inland Water Biol 4:301–307. https://doi.org/10.1134/S1995082911030102
Joos L, Beirinckx S, Haegeman A, Debode J, Vandecasteele B, Baeyen S et al (2020) Daring to be differential: metabarcoding analysis of soil and plant-related microbial communities using amplicon sequence variants and operational taxonomical units. BMC Genomics 21(1):1–17. https://doi.org/10.1186/s12864-020-07126-4
Kozhova OM, Izmest’eva LR (1998) Lake Baikal: evolution and Biodiversity. Backhuys Publishers, Leiden, Netherlands
Kravtsova LS, Peretolchina TE, Triboy TI, Nebesnykh IA, Kupchinskiy AB, Tupikin AE, Kabilov MR (2021) The study of the diversity of hydrobionts from Listvennichny Bay of Lake Baikal by DNA metabarcoding. Russ J Genet 57(4):460–467. https://doi.org/10.1134/S1022795421040050
Leite BR, Vieira PEFR, Troncoso JS, Costa FO (2021) Comparing species detection success between molecular markers in DNA metabarcoding of coastal macroinvertebrates. Metabarcoding and Metagenomics 5:249–260. https://doi.org/10.3897/mbmg.5.70063
Liu YX, Qin Y, Chen T, Lu M, Qian X, Guo X, Bai Y (2021) A practical guide to amplicon and metagenomic analysis of microbiome data. Protein Cell 12:315–330. https://doi.org/10.1007/s13238-020-00724-8
Luo A, Ling C, Ho SYW, Zhu CD (2018) Comparison of methods for Molecular Species Delimitation across a range of speciation scenarios. Syst Biol 67(5):830–846. https://doi.org/10.1093/sysbio/syy011
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
McGarigal K, Cushman SA, Stafford S (2013) Multivariate statistics for wildlife and ecology research. Springer Science & Business Media
Mikhailov IS, Zakharova YR, Bukin YS, Galachyants YP, Petrova DP, Sakirko MV, Likhoshway YV (2019) Co-occurrence networks among bacteria and microbial eukaryotes of Lake Baikal during a spring phytoplankton bloom. Microb Ecol 77:96–109. https://doi.org/10.1007/s00248-018-1212-2
Mikhailov IS, Galachyants YP, Bukin YS, Petrova DP, Bashenkhaeva MV, Sakirko MV et al (2022) Seasonal succession and coherence among bacteria and microeukaryotes in Lake Baikal. Microb Ecol 84:404–422. https://doi.org/10.1007/s00248-021-01860-2
Nearing JT, Douglas GM, Comeau AM, Langille MG (2018) Denoising the denoisers: an independent evaluation of microbiome sequence error-correction approaches. PeerJ 6:e5364. https://doi.org/10.7717/peerj.5364
Nguyen L-T, Schmidt HA, von Haeseler A, Minh BQ (2014) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Bio Evol 32:268–274. https://doi.org/10.1093/molbev/msu300
O’hara RB (2005) Species richness estimators: how many species can dance on the head of a pin? J Anim Ecol 74:375–386
Oksanen J (2013) Vegan: ecological diversity. R project 368:1–11. https://cran.r-project.org/web/packages/vegan/vignettes/diversity-vegan.pdf access on 15.06.2022
Oksanen J, Blanchet FG, Friendly M, Kindt R, Legendre P, McGlinn D et al (2016) vegan: Community Ecology Package. Version 2.6–2. (https://CRAN.R-project.org/package=vegan access on 15.06.2022)
Oliveira MC, Repetti SI, Iha C, Jackson CJ, Díaz-Tapia P, Lubiana KMF et al (2018) High-throughput sequencing for algal systematics. Eur J Phycol 53(3):256–272. https://doi.org/10.1080/09670262.2018.1441446
Paradis E, Claude J, Strimmer K (2004) APE: analyses of phylogenetics and evolution in R language. Bioinformatics 20:289–290. https://doi.org/10.1093/bioinformatics/btg412
Pitsch G, Bruni EP, Forster D, Qu Z, Sonntag B, Stoeck T, Posch T (2019) Seasonality of planktonic freshwater ciliates: are analyses based on V9 regions of the 18S rRNA gene correlated with morphospecies counts? Front Microbiol 10:248. https://doi.org/10.3389/fmicb.2019.00248
Pomazkina GV, Belykh OI, Domysheva VM, Sakirko MV, Gnatovsky RY (2010) Structure and dynamics of phytoplankton of Southern Baikal (Russia). Inter J Algae 12(1):64–79. https://doi.org/10.1615/InterJAlgae.v12.i1.50
Popovskaya GI, Likhoshway YV, Genkal SI, Firsova AD (2006) The role of endemic diatom algae in the phytoplankton of Lake Baikal. Hydrobiologia 568:87–94. https://doi.org/10.1007/s10750-006-0328-4
Prodan A, Tremaroli V, Brolin H, Zwinderman AH, Nieuwdorp M, Levin E (2020) Comparing bioinformatic pipelines for microbial 16S rRNA amplicon sequencing. PLoS ONE 15(1):e0227434. https://doi.org/10.1371/journal.pone.0227434
Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig W, Peplies J, Glöckner FO (2007) SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res 35:7188–7196. https://doi.org/10.1093/nar/gkm864
Puillandre N, Lambert A, Brouillet S, Achaz G (2012) ABGD, Automatic Barcode Gap Discovery for primary species delimitation. Mol Ecol 21(8):1864–1877. https://doi.org/10.1111/j.1365-294X.2011.05239.x
Ríos-Castro R, Romero A, Aranguren R, Pallavicini A, Banchi E, Novoa B, Figueras A (2021) High-throughput sequencing of environmental DNA as a tool for monitoring eukaryotic communities and potential pathogens in a coastal upwelling ecosystem. Front Vet Sci 8. https://doi.org/10.3389/fvets.2021.765606
Rooney AP, Ward TJ (2005) Evolution of a large ribosomal RNA multigene family in filamentous fungi: birth and death of a concerted evolution paradigm. Proc Natl Acad Sci USA 102(14):5084–5089. https://doi.org/10.1073/pnas.0409689102
Rusch DB, Halpern AL, Sutton G, Heidelberg KB, Williamson S, Yooseph S et al (2007) The Sorcerer II Global Ocean Sampling Expedition: Northwest Atlantic through Eastern Tropical Pacific. PLoS Biol 5:e77. https://doi.org/10.1371/journal.pbio.0050077
Santi I, Kasapidis P, Karakassis I, Pitta P (2021) A comparison of DNA metabarcoding and microscopy methodologies for the study of aquatic microbial eukaryotes. Diversity 13(5):180. https://doi.org/10.3390/d13050180
Stoeck T, Bass D, Nebel M, Christen R, Meredith D (2010) Multiple marker parallel tag environmental DNA sequencing reveals a highly complex eukaryotic community in marine anoxic water. Mol Ecol 19:21–31. https://doi.org/10.1111/j.1365-294X.2009.04480.x
Thioulouse J, Dray S (2007) Interactive multivariate data analysis in R with the ade4 and ade4TkGUI packages. J Stat Softw 22:1–14. https://doi.org/10.18637/jss.v022.i05
Tragin M, Zingone A, Vaulot D (2018) Comparison of coastal phytoplankton composition estimated from the V4 and V9 regions of the 18S rRNA gene with a focus on photosynthetic groups and especially Chlorophyta. Environ Microbiol 20:506–520. https://doi.org/10.1111/1462-2920.13952
Větrovský T, Baldrian P (2013) The variability of the 16S rRNA gene in bacterial genomes and its consequences for bacterial community analyses. PLoS ONE 8(2):e57923. https://doi.org/10.1371/journal.pone.0057923
Walker G, Dorrell RG, Schlacht A, Dacks JB (2011) Eukaryotic systematics: a user’s guide for cell biologists and parasitologists. Parasitology 138(13):1638–1663. https://doi.org/10.1017/S0031182010001708
Xiao X, Sogge H, Lagesen K, Tooming-Klunderud A, Jakobsen KS, Rohrlack T (2014) Use of high throughput sequencing and light microscopy show contrasting results in a study of phytoplankton occurrence in a freshwater environment. PLoS ONE 9(8):e106510. https://doi.org/10.1371/journal.pone.0106510
Yi Z, Berney C, Hartikainen H, Mahamdallie S, Gardner M, Boenigk J, Cavalier-Smith T, Bass D (2017) High throughput sequencing of microbial eukaryotes in Lake Baikal reveals ecologically differentiated communities and novel evolutionary radiations. FEMS Microbiol Ecol 93:fix073. https://doi.org/10.1093/femsec/fix073
Zar JH (1972) Significance testing of the Spearman rank correlation coefficient. J Am Stat Assoc 67(339):578–580
Zhang J, Kapli P, Pavlidis P, Stamatakis A (2013) A general species delimitation method with applications to phylogenetic placements. Bioinformatics 29(22):2869–2876. https://doi.org/10.1093/bioinformatics/btt499
Acknowledgements
Anonymous reviewers are acknowledged for their honest and constructive comments which have greatly improved this work.
Funding
This work was supported by the Ministry of Science and Higher Education of Russian Federation project no. 0279–2021-0009 “Study of the role of selected genes and proteins of Baikal diatoms by methods of bioinformatics and physicochemical biology”.
Author information
Authors and Affiliations
Contributions
Yuri Bukin performed methodological planning, statistical analysis, prepared figures, wrote the main text of the manuscript. Ivan Mikhailov performed high-throughput sequencing data analysis, statistical analysis, prepared figures, wrote the main text of the manuscript, organization of expedition work and sampling. Darya Petrova carried out the DNA extraction, quality control of the original DNA preparations. Yuri Galachyants performed methodological planning, performed selection of primers for 18S rRNA. Yulia Zakharova organization of expedition work and sampling. Yelena Likhoshway provided guidance and methodological planning. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no competing interests to declare that are relevant to the content of this article.
Consent for publication
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bukin, Y.S., Mikhailov, I.S., Petrova, D.P. et al. The effect of metabarcoding 18S rRNA region choice on diversity of microeukaryotes including phytoplankton. World J Microbiol Biotechnol 39, 229 (2023). https://doi.org/10.1007/s11274-023-03678-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-023-03678-1