Abstract
Background
Targeted gene inactivation (TGI) is a widely used technique for the study of genes’ functions. There are many different methods for TGI, however, most of them are so complicated and time-consuming. New promising genetic engineering tools are developing for this purpose. In the present study, for the first time we disrupted a virulence gene from Salmonella enterica serovar Typhi (S. Typhi), located in the bacterial chromosome using CRISPR/Cas9 system and homology directed repair (HDR).
Methods
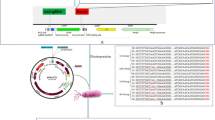
For this aim, pCas9 plasmid containing Cas9 enzyme and required proteins for homology directed recombination was transferred to S. Typhi by electroporation. On the other hand, a specific guide RNA (gRNA) was designed using CRISPOR online tool. Synthetic gRNA was cloned into pTargetF plasmid. Also, a DNA fragment (HDR fragment) was designed to incorporate into the bacterial chromosome following the cleavage of the bacterial genome by Cas9 enzyme. pTargetF containing gRNA and HDR fragment were co-transferred to S. Typhi containing pcas9 plasmid. The transformed bacteria were screened for recombination using PCR, restriction digestion and sequencing.
Results
The results of PCR, restriction digestion and sequencing showed the successful recombination of S. Typhi, in which the gidA gene is disrupted.
Conclusion
In the present study we aimed to develop a rapid and robust method for targeted gene inactivation in a bacterial species, S. Typhi. This procedure can be exploited for disruption of other Salmonella as well as other bacteria’s genes.






Similar content being viewed by others
References
Anzalone AV, Koblan LW, Liu DR (2020) Genome editing with CRISPR–Cas nucleases, base editors, transposases and prime editors. Nat Biotechnol 38(7):824–844
Barrangou R, van Pijkeren J-P (2016) Exploiting CRISPR–Cas immune systems for genome editing in bacteria. Curr Opin Biotechnol 37:61–68
Behrens B, Karber G (1983) Mathematics for naturalists and agriculturalists. PWN, Warszawa, pp 218–218
Chen K et al (2019) CRISPR/Cas genome editing and precision plant breeding in agriculture. Annu Rev Plant Biol 70(1):667–697
Cirillo DM et al (1998) Macrophage-dependent induction of the Salmonella pathogenicity island 2 type III secretion system and its role in intracellular survival. Mol Microbiol 30(1):175–188
Diallo M et al (2020) Adaptation and application of a two-plasmid inducible CRISPR-Cas9 system in Clostridium beijerinckii. Methods 172:51–60
Duffy MJ et al (2020) Targeting p53 for the treatment of cancer. Seminars in cancer biology, Elsevier, Amsterdam
Ebrahimi F et al (2018) Designing a Recombinant Vaccine Containing three bacterial proteins of EHEC, ETEC, and shigella dysentery antigens in E. coli and evaluation of its Humoral immunity in Mic. J Mazandaran Univ Med Sci 27(157):1–16
Feng Z et al (2013) Efficient genome editing in plants using a CRISPR/Cas system. Cell Res 23(10):1229–1232
Gao T et al (2016) GidA, a tRNA modification enzyme, contributes to the growth, and virulence of Streptococcus suis serotype 2. Front Cell Infect Microbiol 6:44
Gupta RM, Musunuru K (2014) Expanding the genetic editing tool kit: ZFNs, TALENs, and CRISPR-Cas9. J Clin Investig 124(10):4154–4161
Halloy F et al (2022) Innovative developments and emerging technologies in RNA therapeutics. RNA Biol 19(1):313–332
Han H (2018) RNA interference to knock down gene expression. Dis Gene Identif. https://doi.org/10.1007/978-1-4939-7471-9_16
Horvath P, Barrangou R (2010) CRISPR/Cas, the immune system of bacteria and archaea. Science 327(5962):167–170
Hou M et al (2020) Genetic editing of the virulence gene of Escherichia coli using the CRISPR system. PeerJ 8:e8881
Katalani C et al (2020) CRISPR-based diagnosis of infectious and noninfectious diseases. Biol procedures online 22(1):1–14
Kim H, Kim J-S (2014) A guide to genome engineering with programmable nucleases. Nat Rev Genet 15(5):321–334
Kinscherf TG, Willis DK (2002) Global regulation by gidA in Pseudomonas syringae. J Bacteriol 184(8):2281–2286
Lee AK, Detweiler CS, Falkow S (2000) OmpR regulates the two-component system SsrA-SsrB in Salmonella pathogenicity island 2. J Bacteriol 182(3):771–781
Levanova A, Poranen MM (2018) RNA interference as a prospective tool for the control of human viral infections. Front Microbiol 9:2151
Lostroh CP, Lee CA (2001) The HilA box and sequences outside it determine the magnitude of HilA-dependent activation of P prgH from Salmonella pathogenicity island 1. J Bacteriol 183(16):4876–4885
MacLachlan P, Sanderson K (1985) Transformation of Salmonella typhimurium with plasmid DNA: differences between rough and smooth strains. J Bacteriol 161(1):442–445
Meng X et al (2008) Targeted gene inactivation in zebrafish using engineered zinc-finger nucleases. Nat Biotechnol 26(6):695–701
Platt RJ et al (2014) CRISPR-Cas9 knockin mice for genome editing and cancer modeling. Cell 159(2):440–455
Rehl JM et al (2013) GidA expression in Salmonella is modulated under certain environmental conditions. Curr Microbiol 67(3):279–285
Setton J et al (2021) Synthetic lethality in cancer therapeutics: the next generationsynthetic lethality in cancer therapeutics: emerging concepts and biomarkers. Cancer Discov 11(7):1626–1635
Sha J et al (2004) Molecular characterization of a glucose-inhibited division gene, gidA, that regulates cytotoxic enterotoxin of Aeromonas hydrophila. Infect Immun 72(2):1084–1095
Shi R et al (2009) Structure-function analysis of Escherichia coli MnmG (GidA), a highly conserved tRNA-modifying enzyme. J Bacteriol 191(24):7614–7619
Shippy DC et al (2011) Biological and virulence characteristics of Salmonella enterica serovar Typhimurium following deletion of glucose-inhibited division (gidA) gene. Microb Pathog 50(6):303–3136
Singh P et al (2020) RNA interference nanotherapeutics for treatment of glioblastoma multiforme. Mol Pharm 17(11):4040–4066
Tolonen AC, Chilaka AC, Church GM (2009) Targeted gene inactivation in Clostridium phytofermentans shows that cellulose degradation requires the family 9 hydrolase Cphy3367. Mol Microbiol 74(6):300–1313
von Meyenburg K et al (1982) Promoters of the atp operon coding for the membrane-bound ATP synthase of Escherichia coli mapped by tn 10 insertion mutations. Mol Gen Genet MGG 188(2):240–248
Wang N et al (2022) Inactivation of a wheat protein kinase gene confers broad-spectrum resistance to rust fungi. Cell 185(16):2961–2974e19
Wierzba TF, Sanders JW (2019) The global burden of enteric fevers in the age of typhoid-conjugate vaccines. Lancet Infect Dis 19(4):340–341
Zhang T et al (2019) Removal of antibiotic resistance genes and control of horizontal transfer risk by UV, chlorination and UV/chlorination treatments of drinking water. Chem Eng J 358:589–597
Zhang N et al (2022) CRISPR/Cas9–Mediated genome editing for Pseudomonas fulva, a Novel Pseudomonas Species with Clinical, Animal, and Plant–Associated isolates. Int J Mol Sci 23(10):5443
Acknowledgements
This study is a part of Y.T. PhD thesis. Authors thanks the staff of pathobiology Department of Shahid Bahonar University of Kerman and Biology Research Center of Imam Hossein University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tarverdizadeh, Y., Khalili, M., Esmaeili, S. et al. Targeted gene inactivation in Salmonella Typhi by CRISPR/Cas9-assisted homologous recombination. World J Microbiol Biotechnol 39, 58 (2023). https://doi.org/10.1007/s11274-022-03504-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-022-03504-0