Abstract
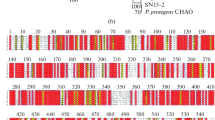
YmdB, which can regulate biofilm formation independently, has been reported to exist in Bacillus subtilis. The B. cereus 0–9 genome also encodes a YmdB-like protein, which has measureable phosphodiesterase activity, and 72.35% sequence identity to YmdB protein of B. subtilis 168. In this work, we studied the function of YmdB protein and its encoding gene, ymdB, in B. cereus 0–9. Our results indicated that YmdB protein is critical for the biofilm formation of B. cereus 0–9. In ΔymdB mutant, the transcriptional levels of sinR and hag were up-regulated, and those of genes closely related to biofilm formation, such as sipW, tasA and calY, were down-regulated. Deletion of ymdB gene stimulates the swarming motility of B. cereus 0–9, and enhances it to travel outward, but reduces its ability to form complex spatial structures on the solid surface of MSgg plates. Hence, it is considered that YmdB plays a key role in biofilm formation, and this effect is likely achieved through the function of repressor SinR in B. cereus 0–9. Furthermore, by comparing the amino acid sequences of YmdB by Basic Local Alignment Search Tool (BLAST) in Genebank, we found that YmdB homologues are present in a variety of bacteria (Including Gram-negative bacteria) except B. subtilis and B. cereus. All these bacteria come at different evolutionary distances and belong to different genera. Therefore, we believe that YmdB exists in many types of bacteria and plays an important role in the stress-resistance of bacteria to adapt to the environment. These results can help us to further understand the biocontrol characteristics of B. cereus 0–9.









Similar content being viewed by others
References
Arnaud M, Chastanet A, Débarbouillé M (2004) New vector for efficient allelic replacement in naturally nontransformable, low-GC-content, gram-positive Bacteria. Appl Environ Microb 70:6887–6891
Branda SS, González-Pastor JE, Dervyn E et al (2004) Genes involved in formation of structured multicellular communities by Bacillus subtilis. J Bacteriol 186:3970–3979
Branda SS, Gonzalez-Pastor JE, Ben-Yehuda S, Losick R, Kolter R (2001) Fruiting body formation by Bacillus subtilis. Proc Natl Acad Sci USA 98:11621–11626
Candela T, Fagerlund A, Buisson C et al (2019) CalY is a major virulence factor and a biofilm matrix protein. Mol Microbiol 111:1416–1429
Caro-Astorga J, Pérez-García A, De Vicente A, Romero D (2015) A genomic region involved in the formation of adhesin fibers in Bacillus cereus biofilms. Front Microbiol 5:1–11
Chu F, Kearns DB, Branda SS, Kolter R, Losick R (2006) Targets of the master regulator of biofilm formation in Bacillus subtilis. Mol Microbiol 59:1216–1228
Diethmaier C, Pietack N, Gunka K et al (2011) A novel factor controlling bistability in Bacillus subtilis: the YmdB protein affects flagellin expression and biofilm formation. J Bacteriol 193:5997–6007
Diethmaier C, Newman JA, Akos TK et al (2014) The YmdB phosphodiesterase is a global regulator of late adaptive responses in Bacillus subtilis. J bacteriol 196:265–275
Dubey G, Malli MG, Dubrovsky A, Amen T, Tsipshtein S, Rouvinski A (2016) Architecture and characteristics of bacterial nanotubes. Dev Cell 36:453–461
Fagerlund A, Dubois T, Økstad OA et al (2014) SinR control sentero toxin expression in Bacillus thuringiensis biofilms. PLoS ONE 9:e87532
Gao T, Li Y, Ding M, Chai Y, Wang Q (2017) The phosphotransferase system gene ptsI in Bacillus cereus regulates expression of sodA2 and contributes to colonization of wheat roots. Res Microbiol 168:524–535
Gao T, Ding M, Yang C, Fan H, Chai Y, Li Y (2019) The phosphotransferase system gene ptsH plays an important role in MnSOD production, biofilm formation, swarming motility, and root colonization in Bacillus cereus 905. Res Microbiol 170:86–96
Hobley L, Ostrowski A, Rao FV et al (2013) BslA is a self-assembling bacterial hydrophobin that coats the Bacillus subtilis biofilm. Proc Natl Acad Sci USA 110:13600–13605
Kang X, Wang L, Guo Y et al (2019) A comparative transcriptomic and proteomic analysis of hexaploid wheat’s responses to colonization by Bacillus velezensis and Gaeumannomyces graminis, both separately and combined. Mol Plant Microbe Interact 32:1336–1347
Kearns DB, Chu F, Branda SS, Kolter R, Losick R (2005) A master regulator for biofilm formation by Bacillus subtilis. Mol Microbiol 55:739–749
Kim T, Lee J, Kim K (2013) Escherichia coli YmdB regulates biofilm formation independently of its role as an RNase III modulator. BMC Microbiol 13:266
Kim M, Kim M, Kim K (2017) Ymdb-mediated down-regulation of sucA inhibits biofilm formation and induces apramycin susceptibility in Escherichia coli. Biochem Biophys Res Commun 483:252–257
Li M, Zhao J, Tang N, Sun H, Huang J (2018) Horizontal gene transfer from bacteria and plants to the arbuscular mycorrhizal fungus Rhizophagus irregularis. Front Plant Sci 9:701
Mäder U, Schmeisky AG, Flórez LA, Stülke J (2012) Subtiwiki—a comprehensive community resource for the model organism Bacillus subtilis. Nucleic Acids Res 40:D1278–1287
Mamou G, Malli MG, Rouvinski A, Rosenberg A, Ben-Yehuda S (2016) Early developmental program shapes colony morphology in bacteria. Cell Report 14:1850–1857
Majed R, Faille C, Kallassy M, Gohar M (2016) Bacillus cereus biofilms-same, only different. Front Microbiol 7:1054
Milton G, Draughn L, Bobay BG et al (2020) The solution structures and interaction of SinR and SinI: elucidating the mechanism of action of the master regulator switch for biofilm formation in Bacillus subtilis. J Mol Biol 432:343–357
Moons P, Chris M, Abram A (2009) Bacterial interactions in biofilms. Crit Rev Microbiol 35:157–168
Newman JA, Rodrigues C, Lewis RJ (2013) Molecular basis of the activity of SinR protein, the master regulator of biofilm formation in Bacillus subtilis. J Biol Chem 288:10766–10778
Ng WL, Bassler BL (2009) Bacterial quorum-sensing network architectures. Annu Rev Genet 43:197–222
Pflughoeft KJ, Sumby P, Koehler TM (2011) Bacillus anthracis sin locus and regulation of secreted proteases. J Bacteriol 193:631–639
Romero D (2013) Bacterial determinants of the social behavior of Bacillus subtilis. Res Microbiol 164:788–798
Romero D, Vlamakis H, Losick R, Kolter R (2014) Functional analysis of the accessory protein TapA in Bacillus subtilis amyloid fiber assembly. J Bacteriol 196:1505–1513
Shin DH, Proudfoot M, Lim HJ, Choi IK, Kim SH (2007) Structural and enzymatic characterization of dr1281: a calcineurin-like phosphoesterase from Deinococcus radiodurans. Proteins 70:1000–1009
Song Y, Miao Y, Song CP (2014) Behind the scenes: the roles of reactive oxygen species in guard cells. New Phytol 201:1121–1140
Stover AG, Driks A (1999a) Control of synthesis and secretion of the Bacillus subtilis protein YqxM. J Bacteriol 181:7065–7069
Stover AG, Driks A (1999b) Secretion, localization, and antibacterial activity of TasA, a Bacillus subtilis spore-associated protein. J Bacteriol 181:1664–1672
Vlamakis H, Chai Y, Beauregard P, Losick R, Kolter R (2013) Sticking together: building a biofilm the Bacillus subtilis way. Nat Rev Microbiol 11:157–168
Wan T, Liu ZM, Li LF et al (2018) A genome for gnetophytes and early evolution of seed plants. Nat Plants 4:82–89
Xu Y, Chen M, Zhang Y et al (2014) The phosphotransferase system gene ptsI in the endophytic bacterium Bacillus cereus is required for biofilm formation, colonization, and biocontrol against wheat sharp eyespot. FEMS Microbiol Lett 354:142–152
Yan M, Wang X, Deng J et al (2016) DNA methylation and cerebellar development, the regulation of Notch and Shh pathway. Ital J Zool 83:34–42
Zhang J, Wang H, Huang Q et al (2020) Four superoxide dismutases of Bacillus cereus 0–9 are non-redundant and perform different functions in diverse living conditions. World J Microbiol Biotechnol 36:12
Acknowledgements
This study was funded by National Nature Science Foundation of China (NSFC) (31572047) and NSFC (31701831).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, J., Wang, H., Xie, T. et al. The YmdB protein regulates biofilm formation dependent on the repressor SinR in Bacillus cereus 0–9. World J Microbiol Biotechnol 36, 165 (2020). https://doi.org/10.1007/s11274-020-02933-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-020-02933-z



