Abstract
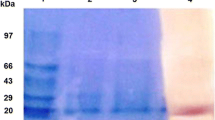
The poly(3-hydroxybutyrate-co-3-hydroxyvalerate) (PHBV)-degrading strain Acidovorax sp. HB01 was isolated from an activated sludge sample. A novel PHBV depolymerase with a molecular weight of 43.4 kDa was purified to homogeneity from the culture supernatant of the HB01 strain. The optimum pH and temperature of the PHBV depolymerase were 7.0 and 50 °C, respectively. The PHBV depolymerase can also degrade polyhydroxybutyrate, poly (3-hydroxybutyrate-co-4-hydroxybutyrate), and poly(caprolactone); however, the PHBV degradation activity of the depolymerase is higher than its activity against the other polymers. Effect of metal ions and various inhibitors on the PHBV depolymerase activity was examined. The addition of Na+, K+, and Ca2+ markedly increased the hydrolysis rate, whereas the enzyme activity was inhibited by Zn2+, Mg2+, Mn2+, and particularly by Cu2+ and Fe2+. Ethylenediaminetetraacetic acid was found to have a significant inhibitory effect. The main degradation product of depolymerase was identified as the 3-hydroxybutyric acid monomer and 3-hydroxyvaleric acid monomers via mass spectrometry.







Similar content being viewed by others
References
Breed EG, Murray D, Smith NR (1994) Berger’s manual of determinative bacteriology, 9th edn. The Williams and Wilkins co, Baltimore
Chanprateep S (2010) Current trends in biodegradable polyhydroxyalkanoates. J Biosci Bioeng 110:621–632
Chen GQ (2009) A microbial polyhydroxyalkanoates (PHA) based bio- and materials industry. Chem Soc Rev 38:2434–2446
Cousley RR (2009) A stent-guided mini-implant system. J Clinic Orthod 43:403–407
Feng LD, Wang Y, Inagawa Y, Kasuya K, Saito T, Doi Y, Inoue Y (2004) Enzymatic degradation behavior of comonomer compositionally fractionated bacterial poly(3-hydroxybutyrate-co-3-hydroxyvalerate)s by poly(3-hydroxyalkanoate) depolymerases isolated from Ralstonia pickettii T1 and Acidovorax sp. TP4. Polym Degrad Stabil 84:95–104
García MC, Hueso DKB (2010) Simultaneous kinetic determination of 3-hydroxybutyrate and 3-hydroxyvalerate in biopolymer degradation processes. Talanta 80:1436–1440
Gordon SH, Shogren RL, Imam SH, Govind NS, Greene RV (2000) A semiempirical model for predicting biodegradation profiles of individual polymers in starch-poly-(β-hydroxybutyrate-co-β-hydroxyvalerate) Bioplastic. J Appl Polym Sci 76:1767–1776
Han JS, Kim MN (2002) Purification and characterisation of extracellular poly(3-hydroxybutyrate) depolymerase from Penicillium simplicissimum LAR13. J Microbiol 40:20–25
Iyer S, Shah R, Sharma A, Jendrossek D, Desai A (2000) Purification of Aspergillus fumigatus (Pdf1) poly(3-hydroxybutyrate) depolymerase using a new, single-step substrate affinity chromatography method. J Polym Environ 8:197–203
Kim DY, Rhee YH (2003) Biodegradation of microbial and synthetic polyesters by fungi. Appl Microbiol Biotechnol 61:300–308
Kim DY, Yun JH, Kim HW (2002) Purification and characterisation of poly(3-hydroxybutyrate) depolymerase from a fungal isolate, Emericellopsis minima W2. J Microbiol 40:129–133
Kita K, Ishimaru K, Teraoka M, Yanase H, Kato N (1995) Properties of poly (3-hydroxybutyrate) depolymerases from a marine bacterium, Alcaligenes faecalis AE 122. Appl Environ Microb 61:1727–1730
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 277:680–685
Ohura T, Kasuya KI, Doi Y (1999) Cloning and characterization of the polyhydroxybutyrate depolymerase gene of Pseudomonas stutzeri and analysis of the function of substrate-binding domains. Appl Environ Microb 65:189–197
Quinteros R, Goodwin S, Lenz RW, Park WH (1999) Extracellular degradation of medium chain length poly(β-hydroxyalkanoates) by Comamonas sp. Int J Biol Macromol 25:135–143
Rhee YH, Kim YH, Shin KS, Rhee YH, Kim YH, Shin KS (2006) Characterization of an extracellular poly(3-hydroxyoctanoate) depolymerase from the marine isolate, Pseudomonas luteola M13-4. Enzyme Microb Tech 38:529–535
Shah AA, Hasan F, Hameed A, Ahmed S (2007) Isolation and characterization of poly (3-hydroxybutyrate-co-3-hydroxyvalerate) degrading bacteria and purification of PHBV depolymerase from newly isolated Bacillus sp. AF3. Int Biodeter Biodegr 60:109–115
Shirakura Y, Fukui T, Saito T, Okamoto Y, Narikawa T, Koide K, Tomita K, Takemasa T, Masamune S (1986) Degradation of poly(3-hydroxybutyrate) by poly(3-hydroxybutyrate) depolymerase from Alcaligenes faecalis T1. Biochim Biophys Acta 880:46–53
Takaki H, Kimoto A, Kodaira S, Nashimoto M, Takagi M (2006) Isolation of a gram-positive poly (3-hydroxybutyrate)(PHB)-degrading bacterium from compost, and cloning and characterization of a gene encoding PHB depolymerase of Bacillus megaterium N-18-25-9. FEMS Microbiol Lett 264:152–159
Tokiwa Y, Calabia BP (2004) Degradation of microbial polyesters. Biotechnol Lett 26:1181–1189
Xu XY, Li XT, Peng SW, Xiao JF, Liu C, Fang G, Chen KC, Chen GQ (2010) The behaviour of neural stem cells on polyhydroxyalkanoate nanofiber scaffolds. Biomaterials 31:3967–3975
Zhou HL, Wang ZY, Chen S, Liu DB, Xia H (2009) Purification and characterization of extracellular poly(β-hydroxybutyrate) depolymerase from Penicillium sp. DS9701-D2. Polym-Plast Technol 48:58–63
Acknowledgments
This work was supported by National Natural Science Foundation of China (Grant No. 31100099) and Science Foundation of Liaoning Shihua University (No. 2011XJJ-025).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, Z., Gao, J., Li, L. et al. Purification and characterization of an extracellular poly(3-hydroxybutyrate-co-3-hydroxyvalerate) depolymerase from Acidovorax sp. HB01. World J Microbiol Biotechnol 28, 2395–2402 (2012). https://doi.org/10.1007/s11274-012-1048-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-012-1048-8