Abstract
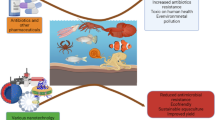
Anthropogenic activities are closely related to the emergence, spread, and evolution of antimicrobial resistance (AMR). In addition to the introduction of chemical pollutants into the environment, dam failure may contribute to the selection of AMR and metal tolerance, and horizontal gene transfer. In this context, environmental samples from unaffected and affected sites were analyzed to obtain bacterial strains and investigate AMR and metal tolerance, as well as antimicrobial resistance genes (ARGs) and metal tolerance genes (MTGs). A wide diversity of emerging bacterial pathogens has been detected, which are often associated with multidrug resistance and multimetal tolerance. Intrinsic and acquired ARGs and MTGs were identified in clinically important bacteria. Extended-spectrum β-lactamase-encoding genes were exclusive of Pseudomonas aeruginosa and Escherichia coli from sediments and soils of affected sites. Overall, P. aeruginosa, E. coli, and Klebsiella pneumoniae were the ones that presented ARGs and/or MTGs, as well as the coexistence of them, which increased in the affected sites. These findings support that mining pollution increased the amount of MTGs and that they co-occur with different ARGs in a great diversity of bacterial species.



Similar content being viewed by others
Data Availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Bonnet, M., Lagier, J. C., Raoult, D., & Khelaifia, S. (2019). Bacterial culture through selective and non-selective conditions: the evolution of culture media in clinical microbiology. New Microbes and New Infections, 34, 100622. https://doi.org/10.1016/j.nmni.2019.100622
Bourceret, A., Cébron, A., Tisserant, E., Poupin, P., Bauda, P., Beguiristain, T., & et al. (2015). The bacterial and fungal diversity of an aged PAH- and heavy metal-contaminated soil is affected by plant cover and edaphic parameters. Microbial Ecology, 71, 711–724. https://doi.org/10.1007/s00248-015-0682-8
Braz, V. S., Furlan, J. P. R., Passaglia, J., Falcão, J. P., & Stehling, E. G. (2018). Genotypic diversity and presence of β-lactamase encoding genes in Pseudomonas aeruginosa isolated from Brazilian soils. Applied Soil Ecology, 129, 94–97. https://doi.org/10.1016/j.apsoil.2018.05.005
Buch, A. C., Sautter, K. D., Marques, E. D., & Silva-Filho, E. V. (2020). Ecotoxicological assessment after the world’s largest tailing dam collapse (Fundão dam, Mariana, Brazil): effects on oribatid mites. Environmental Geochemistry and Health, 42, 3575–3595. https://doi.org/10.1007/s10653-020-00593-4
Chen, J., Li, J., Zhang, H., Shi, W., & Liu, Y. (2019). Bacterial Heavy-Metal and Antibiotic Resistance Genes in a Copper Tailing Dam Area in Northern China. Frontiers in Microbiology, 10, 1916. https://doi.org/10.3389/fmicb.2019.01916
Chen, X., Du, Z., Song, X., Wang, L., Wei, Z., Jia, L., & Zhao, R. (2023). Evaluating the occurrence frequency of horizontal gene transfer induced by different degrees of heavy metal stress. Journal of Cleaner Production, 328, 135371. https://doi.org/10.1016/j.jclepro.2022.135371
Cionek, V. M., Alves, G. H. Z., Tófoli, R. M., Rodrigues-Filho, J. L., & Dias, R. M. (2019). Brazil in the mud again: lessons I learned from Mariana dam collapse. Biodiversity and Conservation, 28, 1935–1938. https://doi.org/10.1007/s10531-019-01762-3
D’Costa, V. M., King, C. E., Kalan, L., Morar, M., Sung, W. W., Schwarz, C., et al. (2011). Antibiotic resistance is ancient. Nature, 477, 457–461. https://doi.org/10.1038/nature10388
de Castro, A. P., Fernandes Gda, R., & Franco, O. L. (2014). Insights into novel antimicrobial compounds and antibiotic resistance genes from soil metagenomes. Frontiers in Microbiology, 5, 489. https://doi.org/10.3389/fmicb.2014.00489
Deredjian, A., Colinon, C., Brothier, E., Favre-Bonté, S., Cournoyer, B., & Nazaret, S. (2011). Antibiotic and metal resistance among hospital and outdoor strains of Pseudomonas aeruginosa. Research in Microbiology, 162(7), 689–700. https://doi.org/10.1016/j.resmic.2011.06.007
Domínguez, D. C., Chacón, L. M., & Wallace, D. (2021). Anthropogenic Activities and the Problem of Antibiotic Resistance in Latin America: A Water Issue. Water, 13, 2693. https://doi.org/10.3390/w13192693
Farhad, M., Noor, M., Yasin, M. Z., Nizamani, M. H., Turan, V., & Iqbal, M. (2024). Interactive Suitability of Rice Stubble Biochar and Arbuscular Mycorrhizal Fungi for Improving Wastewater-Polluted Soil Health and Reducing Heavy Metals in Peas. Sustainability, 16, 634. https://doi.org/10.3390/su16020634
Fernandes, G. W., Goulart, F. F., Ranieri, B. D., Coelho, M. S., Dales, K., Boesche, N., et al. (2016). Deep into the mud: ecological and socio-economic impacts of the dam breach in Mariana, Brazil. Natureza & Conservação, 14, 35–45. https://doi.org/10.1016/j.ncon.2016.10.003
Furlan, J. P. R., dos Santos, L. D. R., Moretto, J. A. S., Ramos, M. S., Gallo, I. F. L., Alves, G. A. D., et al. (2020). Occurrence and abundance of clinically relevant antimicrobial resistance genes in environmental samples after the Brumadinho dam disaster, Brazil. Science of The Total Environment, 726, 138100. https://doi.org/10.1016/j.scitotenv.2020.138100
Furlan, J. P. R., Ramos, M. S., Rosa, R. D. S., Savazzi, E. A., & Stehling, E. G. (2022). Occurrence and genetic characteristics of multidrug-resistant Escherichia coli isolates co-harboring antimicrobial resistance genes and metal tolerance genes in aquatic ecosystems. International Journal of Hygiene and Environmental Health, 244, 114003. https://doi.org/10.1016/j.ijheh.2022.114003
Furlan, J. P. R., & Stehling, E. G. (2021). Multiple sequence types, virulence determinants and antimicrobial resistance genes in multidrug- and colistin-resistant Escherichia coli from agricultural and non-agricultural soils. Environmental Pollution, 288, 117804. https://doi.org/10.1016/j.envpol.2021.117804
Gaeta, N.C., de Carvalho, D.U., Fontana, H., Sano, E., Moura, Q., Fuga, B., & et al. (2022). Genomic features of a multidrug-resistant and mercury-tolerant environmental Escherichia coli recovered after a mining dam disaster in South America. Science of The Total Environment, 823, 153590. https://doi.org/10.1016/j.scitotenv.2022.153590
Glushakova, А. М., Kachalkin, А. V., Prokof’eva, T. V., & Lysak, L. V. (2022). Enterobacteriaceae in soils and atmospheric dust aerosol accumulations of Moscow city. Current Research in Microbial Sciences, 3, 100124. https://doi.org/10.1016/10.1016/j.crmicr.2022.100124
Hatje, V., Pedreira, R. M. A., de Rezende, C. E., Schettini, C. A. F., de Souza, G. C., Marin, D. C., et al. (2017). The environmental impacts of one of the largest tailing dam failures worldwide. Scientific Reports, 7, 10706. https://doi.org/10.1038/s41598-017-11143-x
Hernando-Amado, S., Coque, T. M., Baquero, F., & Martínez, J. L. (2019). Defining and combating antibiotic resistance from One Health and Global Health perspectives. Nature Microbiology, 4, 1432–1442. https://doi.org/10.1038/s41564-019-0503-9
Imran, M., Das, K. R., & Naik, M. M. (2019). Co-selection of multi-antibiotic resistance in bacterial pathogens in metal and microplastic contaminated environments: an emerging health threat. Chemosphere, 215, 846–857. https://doi.org/10.1016/j.chemosphere.2018.10.114
Keim, C.N., Serna, J.D.P., Acosta-Avalos, D., Neumann, R., Silva, A.S., Jurelevicius, D.A., & et al.. (2021). Dissimilatory Iron-Reducing Microorganisms Are Present and Active in the Sediments of the Doce River and Tributaries Impacted by Iron Mine Tailings from the Collapsed Fundão Dam (Mariana, MG, Brazil). Minerals, 11, 244. https://doi.org/10.3390/min11030244
Kim, D. W., & Cha, C. J. (2021). Antibiotic resistome from the One-Health perspective: understanding and controlling antimicrobial resistance transmission. Experimental & Molecular Medicine, 53, 301–309. https://doi.org/10.1038/s12276-021-00569-z
Lam, M. M. C., Wick, R. R., Watts, S. C., Cerdeira, L. T., Wyres, K. L., & Holt, K. E. (2021). A genomic surveillance framework and genotyping tool for Klebsiella pneumoniae and its related species complex. Nature Communications, 12, 4188. https://doi.org/10.1038/s41467-021-24448-3
Liao, X., Fang, L., Li, L., Sun, J., Li, X., Chen, M., & et al. (2015). Characterization of chromosomal qnrB and ampC alleles in Citrobacter freundii isolates from different origins. Infection, Genetics and Evolution, 35, 214-220. https://doi.org/10.1016/j.meegid.2015.07.011
Maciel-Guerra, A., Baker, M., Hu, Y., Wang, W., Zhang, X., Rong, J., et al. (2022). Dissecting microbial communities and resistomes for interconnected humans, soil, and livestock. The ISME Journal, 17, 21–35. https://doi.org/10.1038/s41396-022-01315-7
Madrazo, M., Esparcia, A., López-Cruz, I., Alberola, J., Piles, L., Viana, A., et al. (2021). Clinical impact of multidrug-resistant bacteria in older hospitalized patients with community-acquired urinary tract infection. BMC Infectious Diseases, 21, 1232. https://doi.org/10.1186/s12879-021-06939-2
Magiorakos, A. P., Srinivasan, A., Carey, R. B., Carmeli, Y., Falagas, M. E., & Giske, C. G. (2012). Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clinical Microbiology and Infection, 18, 268–281. https://doi.org/10.1111/j.1469-0691.2011.03570.x
Mourão, J., Novais, C., Machado, J., Peixe, L., & Antunes, P. (2015). Metal tolerance in emerging clinically relevant multidrug-resistant Salmonella enterica serotype 4,[5],12:i:- clones circulating in Europe. International Journal of Antimicrobial Agents, 45, 610–616. https://doi.org/10.1016/j.ijantimicag.2015.01.013
Nielsen, T. K., Browne, P. D., & Hansen, L. H. (2022). Antibiotic resistance genes are differentially mobilized according to resistance mechanism. Gigascience, 11, giac072. https://doi.org/10.1093/gigascience/giac072
Perelomov, L., Sizova, O., Rahman, M. M., Perelomova, I., Minkina, T., & Sokolov, S. (2022). Metal-Tolerant Bacteria of Wastewater Treatment Plant in a Large City. Sustainability, 14, 11335. https://doi.org/10.3390/su141811335
Pérez-Rodríguez, F., & Mercanoglu Taban, B. (2019). A State-of-Art Review on Multi-Drug Resistant Pathogens in Foods of Animal Origin: Risk Factors and Mitigation Strategies. Frontiers in Microbiology, 10, 2091. https://doi.org/10.3389/fmicb.2019.02091
Porsani, J. L., de Jesus, F. A. N., & Stangari, M. C. (2019). GPR Survey on an Iron Mining Area after the Collapse of the Tailings Dam I at the Córrego do Feijão Mine in Brumadinho-MG, Brazil. Remote Sensing, 11, 860. https://doi.org/10.3390/rs11070860
Quillaguamán, J., Guzmán, D., Campero, M., Hoepfner, C., Relos, L., & Mendieta, D. (2021). The microbiome of a polluted urban lake harbors pathogens with diverse antimicrobial resistance and virulence genes. Environmental Pollution, 273, 116488. https://doi.org/10.1016/j.envpol.2021.116488
Ramos, M. S., Furlan, J. P. R., Dos Santos, L. D. R., Rosa, R. D. S., Savazzi, E. A., & Stehling, E. G. (2023). Patterns of antimicrobial resistance and metal tolerance in environmental Pseudomonas aeruginosa isolates and the genomic characterization of the rare O6/ST900 clone. Environmental Monitoring and Assessment, 195(6), 713. https://doi.org/10.1007/s10661-023-11344-0
Riesenfeld, C. S., Goodman, R. M., & Handelsman, J. (2004). Uncultured soil bacteria are a reservoir of new antibiotic resistance genes. Environmental Microbiology, 6, 981–989. https://doi.org/10.1111/j.1462-2920.2004.00664.x
Rodrigue, A., Effantin, G., & Mandrand-Berthelot, M. A. (2005). Identification of rcnA (yohM), a nickel and cobalt resistance gene in Escherichia coli. Journal of Bacteriology, 187, 2912–2916. https://doi.org/10.1128/JB.187.8.2912-2916.2005
Samreen, A. I., Malak, H. A., & Abulreesh, H. H. (2021). Environmental antimicrobial resistance and its drivers: a potential threat to public health. Journal of Global Antimicrobial Resistance, 27, 101–111. https://doi.org/10.1016/j.jgar.2021.08.001
Santamarina, J. C., Torres-Cruz, L. A., & Bachus, R. C. (2019). Why coal ash and tailings dam disasters occur. Science, 364, 526–528. https://doi.org/10.1126/science.aax1927
Sojka, M., & Jaskuła, J. (2022). Heavy Metals in River Sediments: Contamination, Toxicity, and Source Identification—A Case Study from Poland. International Journal of Environmental Research and Public Health, 19, 10502. https://doi.org/10.3390/ijerph191710502
Squadrone, S. (2020). Water environments: metal-tolerant and antibiotic-resistant bacteria. Environmental Monitoring and Assessment, 192, 238. https://doi.org/10.1007/s10661-020-8191-8
Turan, V. (2024). Chapter 9: Bioremediation of Lithium and Nickel from Soil and Water. Lithium and Nickel Contamination in Plants and the Environment. World Scientific Series on Advances in Environmental Pollution Management, 219–230. https://doi.org/10.1142/9789811283123_0009
Ture, M., Kilic, M. B., & Altinok, I. (2021). Relationship Between Heavy Metal Accumulation in Fish Muscle and Heavy Metal Resistance Genes in Bacteria Isolated from Fish. Biological Trace Element Research, 199, 1595–1603. https://doi.org/10.1007/s12011-020-02246-0
Wang, Q., Liu, L., Hou, Z., Wang, L., Ma, D., Yang, G., et al. (2020). Heavy metal copper accelerates the conjugative transfer of antibiotic resistance genes in freshwater microcosms. Science of The Total Environment, 717, 137055. https://doi.org/10.1016/j.scitotenv.2020.137055
Wang, Q., Xu, Y., Liu, L., Li, L. Y., Lin, H., Wu, X. Y., et al. (2021). The prevalence of ampicillin-resistant opportunistic pathogenic bacteria undergoing selective stress of heavy metal pollutants in the Xiangjiang River, China. Environmental Pollution, 268, 115362. https://doi.org/10.1016/j.envpol.2020.115362
Wang, Y., Hou, N., Rasooly, R., Gu, Y., & He, X. (2021). Prevalence and Genetic Analysis of Chromosomal mcr-3/7 in Aeromonas From U.S. Animal-Derived Samples. Frontiers in Microbiology, 12, 667406. https://doi.org/10.3389/fmicb.2021.667406
Wright, M. H., Adelskov, J., & Greene, A. C. (2017). Bacterial DNA Extraction Using Individual Enzymes and Phenol/Chloroform Separation. Journal of Microbiology & Biology Education, 18, 18.2.48. https://doi.org/10.1128/jmbe.v18i2.1348
Wu, C., Zhang, G., Xu, W., Jian, S., Peng, L., Jia, D., et al. (2021). New Estimation of Antibiotic Resistance Genes in Sediment Along the Haihe River and Bohai Bay in China: A Comparison Between Single and Successive DNA Extraction Methods. Frontiers in Microbiology, 12, 705724. https://doi.org/10.3389/fmicb.2021.705724
Zagui, G. S., Moreira, N. C., Santos, D. V., Darini, A. L. C., Domingo, J. L., Segura-Muñoz, S. I., & Andrade, L. N. (2021). High occurrence of heavy metal tolerance genes in bacteria isolated from wastewater: A new concern? Environmental Research, 196, 110352. https://doi.org/10.1016/j.envres.2020.110352
Zhang, Y., Gu, A. Z., Cen, T., Li, X., He, M., Li, D., et al. (2018). Sub-inhibitory concentrations of heavy metals facilitate the horizontal transfer of plasmid-mediated antibiotic resistance genes in water environment. Environmental Pollution, 237, 74–82. https://doi.org/10.1016/j.envpol.2018.01.032
Zhuang, M., Achmon, Y., Cao, Y., Liang, X., Chen, L., & Wang, H. (2021). Distribution of antibiotic resistance genes in the environment. Environmental Pollution, 285, 117402. https://doi.org/10.1016/j.envpol.2021.117402
Acknowledgments
The authors thank the FAPESP [grant no. 2018/01890-3], the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) [grant no. 88882.180855/2018-01, 88887.314388/2019-00, 88882.332034/2019-01, 88882.332036/2019-01, 88882.332039/2019-01, 88887.361231/2019-00, 88887.519091/2020-00, and Finance Code 001] and the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) [grant no. 150712/2022-7, 130086/2021-5, 141016/2021-3, 308914/2019-8, 304905/2022-4, and 143965/2015-8] for fellowships.
Funding
This study was supported by Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP) [grant no. 2018/19539-0, 2021/01655-7, and 2018/24069-3].
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical Approval
Not required.
Competing Interest
The authors declare no conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 81 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Furlan, J.P.R., Ramos, M.S., dos Santos, L.D.R. et al. Effects of a Mining Dam Disaster on Antimicrobial-Resistant and Metal-Tolerant Bacterial Strains Recovered from Environmental Samples. Water Air Soil Pollut 235, 333 (2024). https://doi.org/10.1007/s11270-024-07171-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11270-024-07171-9