Abstract
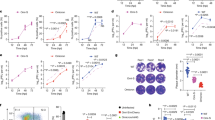
SARS-CoV-2 Omicron has the largest number of mutations among all the known SARS-CoV-2 variants. The presence of these mutations might explain why Omicron is more infectious and vaccines have lower efficacy to Omicron than other variants, despite lower virulence of Omicron. We recently established a long-term in vivo replication model by infecting Calu-3 xenograft tumors in immunodeficient mice with parental SARS-CoV-2 and found that various mutations occurred majorly in the spike protein during extended replication. To investigate whether there are differences in the spectrum and frequency of mutations between parental SARS-CoV-2 and Omicron, we here applied this model to Omicron. At 30 days after infection, we found that the virus was present at high titers in the tumor tissues and had developed several rare sporadic mutations, mainly in ORF1ab with additional minor spike protein mutations. Many of the mutant isolates had higher replicative activity in Calu-3 cells compared with the original SARS-CoV-2 Omicron virus, suggesting that the novel mutations contributed to increased viral replication. Serial propagation of SARS-CoV-2 Omicron in cultured Calu-3 cells resulted in several rare sporadic mutations in various viral proteins with no mutations in the spike protein. Therefore, the genome of SARS-CoV-2 Omicron seems largely stable compared with that of the parental SARS-CoV-2 during extended replication in Calu-3 cells and xenograft model. The sporadic mutations and modified growth properties observed in Omicron might explain the emergence of Omicron sublineages. However, we cannot exclude the possibility of some differences in natural infection.



Similar content being viewed by others
Data availability
All data needed to evaluate the conclusions in this manuscript have been included. The whole-genome sequencing data for the SARS-CoV-2 mutants analyzed in this study are available at the GISAID (https://gisaid.org) under the accession number: EPI_ISL_18454580. EPI_ISL_18454581. EPI_ISL_18454582. EPI_ISL_18454583. EPI_ISL_18454584. EPI_ISL_18454585. EPI_ISL_18454586. EPI_ISL_18454587. EPI_ISL_18454588. EPI_ISL_18454589. EPI_ISL_18454590. EPI_ISL_18454591. EPI_ISL_18454592. EPI_ISL_18454593. EPI_ISL_18454683. EPI_ISL_18454777. EPI_ISL_18454826. EPI_ISL_18454855. EPI_ISL_18454857. EPI_ISL_18454858. EPI_ISL_18454859. EPI_ISL_18454860. EPI_ISL_18454862. EPI_ISL_18454863. EPI_ISL_18454864. EPI_ISL_18454866. EPI_ISL_18454867.
References
1. Arora P, Zhang L, Rocha C, Sidarovich A, Kempf A, Schulz S et al (2022) Comparable neutralisation evasion of SARS-CoV-2 omicron subvariants BA.1, BA.2, and BA.3. Lancet Infect Dis 22: 766–767. https://doi:10.1016/S1473-3099(22)00224-9
2. Viana R, Moyo S, Amoako DG, Tegally H, Scheepers C, Althaus CL et al (2022) Rapid epidemic expansion of the SARS-CoV-2 Omicron variant in southern Africa. Nature 603: 679–686. https://doi:10.1038/s41586-022-04411-y
3. Nyberg T, Ferguson NM, Nash SG, Webster HH, Flaxman S, Andrews N et al (2022) Comparative analysis of the risks of hospitalisation and death associated with SARS-CoV-2 omicron (B.1.1.529) and delta (B.1.617.2) variants in England: a cohort study. Lancet 399: 1303–1312. https://doi:10.1016/S0140-6736(22)00462-7
4. Pulliam JRC, van Schalkwyk C, Govender N, von Gottberg A, Cohen C, Groome MJ et al (2022) Increased risk of SARS-CoV-2 reinfection associated with emergence of Omicron in South Africa. Science 376: eabn4947. https://doi:10.1126/science.abn4947
5. Miller J, Hachmann NP, Collier AY, Lasrado N, Mazurek CR, Patio RC et al (2023) Substantial Neutralization Escape by SARS-CoV-2 Omicron Variants BQ.1.1 and XBB.1. N Engl J Med 388: 662–664. https://doi:10.1056/NEJMc2214314
6. Tian D, Nie W, Sun Y, Ye Q (2022) The Epidemiological Features of the SARS-CoV-2 Omicron Subvariant BA.5 and Its Evasion of the Neutralizing Activity of Vaccination and Prior Infection. Vaccines (Basel) 10: 1699. https://doi:10.3390/vaccines10101699
7. Shao W, Zhang W, Fang X, Yu D, Wang X (2022) Challenges of SARS-CoV-2 Omicron Variant and appropriate countermeasures. J Microbiol Immunol Infect 55: 387–394. https://doi:10.1016/j.jmii.2022.03.007
8. Zhang Y, Zhang T, Fang Y, Liu J, Ye Q, Ding L. SARS-CoV-2 spike L452R mutation increases Omicron variant fusogenicity and infectivity as well as host glycolysis. Signal Transduct Target Ther. 2022 Mar 9;7(1):76. doi: 10.1038/s41392-022-00941-z
9. Chen J, Wang R, Gilby NB, Wei GW (2022) Omicron Variant (B.1.1.529): Infectivity, Vaccine Breakthrough, and Antibody Resistance. J Chem Inf Model 62: 412–422. https://doi:10.1021/acs.jcim.1c01451
10. Woolhouse ME, Gowtage-Sequeria S (2005) Host range and emerging and reemerging pathogens. Emerg Infect Dis 11:1842–1847. https://doi:10.3201/eid1112.050997
11. Lauring AS, Hodcroft EB (2021) Genetic Variants of SARS-CoV-2-What Do They Mean? JAMA 325: 529–531. https://doi:10.1001/jama.2020.27124
12. Kim D, Kim J, Kim M, Park H, Park S, Maharjan S et al (2023) Analysis of spike protein variants evolved in a novel in vivo long-term replication model for SARS-CoV-2. Front Cell Infect Microbiol 13:1280686. https://doi:10.3389/fcimb.2023.1280686
13. Kim D, Maharjan S, Kim J, Park S, Park JA, Park BK et al (2021) MUC1-C influences cell survival in lung adenocarcinoma Calu-3 cells after SARS-CoV-2 infection. BMB Rep 54: 425–430. https://doi:10.5483/BMBRep.2021.54.8.018
14. Park BK, Kim J, Park S, Kim D, Kim M, Baek K et al (2021) MERS-CoV and SARS-CoV-2 replication can be inhibited by targeting the interaction between the viral spike protein and the nucleocapsid protein. Theranostics 11: 3853–3867. https://doi:10.7150/thno.55647
15. Lei C, Yang J, Hu J, Sun X (2021) On the Calculation of TCID50 for Quantitation of Virus Infectivity. Virol Sin 36: 141–144. https://doi:10.1007/s12250-020-00230-5
16. Park S, Kim D, Kim J, Kwon HJ, Lee Y (2023) SARS-CoV-2 infection induces expression and secretion of lipocalin-2 and regulates iron in a human lung cancer xenograft model. BMB Rep 56: 669–674. https://doi.org/10.5483/BMBRep.2023-0177
17. Wu G, Kim D, Kim JN, Park S, Maharjan S, Koh H et al (2018) A Mucin1 C-terminal Subunit-directed Monoclonal Antibody Targets Overexpressed Mucin1 in Breast Cancer. Theranostics 8: 78–91. https://doi:10.7150/thno.21278
18. Park BK, Kim D, Park S, Maharjan S, Kim J, Choi JK et al (2021) Differential Signaling and Virus Production in Calu-3 Cells and Vero Cells upon SARS-CoV-2 Infection. Biomol Ther (Seoul) 29: 273–281. https://doi:10.4062/biomolther.2020.226
19. Baek K, Maharjan S, Akauliya M, Thapa B, Kim D, Kim J et al (2022) Comparison of vaccination efficacy using live or ultraviolet-inactivated influenza viruses introduced by different routes in a mouse model. PLoS One 17: e0275722. https://doi:10.1371/journal.pone.0275722
20. Kim D, Kim J, Park S, Kim M, Baek K, Kang M et al (2021) Production of SARS-CoV-2 N Protein-Specific Monoclonal Antibody and Its Application in an ELISA-Based Detection System and Targeting the Interaction Between the Spike C-Terminal Domain and N Protein. Front Microbiol 12: 726231. https://doi:10.3389/fmicb.2021.726231
21. Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30: 2114–2120. https://doi:10.1093/bioinformatics/btu170
22. Li H, Durbin R (2010) Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26: 589–595. https://doi:10.1093/bioinformatics/btp698
23. Tarasov A, Vilella AJ, Cuppen E, Nijman IJ, Prins P (2015) Sambamba: fast processing of NGS alignment formats. Bioinformatics 31: 2032–2034. https://doi:10.1093/bioinformatics/btv098
24. Li H (2011) A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27: 2987–2993. https://doi:10.1093/bioinformatics/btr509
25. Danecek P, Schiffels S, Durbin R (2016) Multiallelic calling model in bcftools (-m). http://samtools.github.io/bcftools/call-m.pdf. Accessed 17 Apr 2023.
26. Cingolani P, Platts A, Wang le L, Coon M, Nguyen T, Wang L (2012) A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 6: 80–92. https://doi:10.4161/fly.19695
27. Chumakov KM, Dragunsky EM, Norwood LP, Douthitt MP, Ran Y, Taffs RE (1994) Consistent selection of mutations in the 5’-untranslated region of oral poliovirus vaccine upon passaging in vitro. J Med Virol 42: 79–85. https://doi:10.1002/jmv.1890420115
28. Shi G, Li T, Lai KK, Johnson RF, Yewdell JW, Compton AA (2024) Omicron Spike confers enhanced infectivity and interferon resistance to SARS-CoV-2 in human nasal tissue. Nat Commun 15: 889. https://doi.org/10.1038/s41467-024-45075-8.
29. Carabelli AM, Peacock TP, Thorne LG, Harvey WT, Hughes J, COVID-19 Genomics UK Consortium et al (2023) SARS-CoV-2 variant biology: immune escape, transmission and fitness. Nat Rev Microbiol 21: 162–177. https://doi:10.1038/s41579-022-00841-7
30. Markov PV, Ghafari M, Beer M, Lythgoe K, Simmonds P, Stilianakis NI et al (2023) The evolution of SARS-CoV-2. Nat Rev Microbiol 21: 361–379. https://doi:10.1038/s41579-023-00878-2.
31. World Health Organization (WHO) (2023) Tracking SARS-CoV-2 variants. https://www.who.int/activities/tracking-SARS-CoV-2-variants. Accessed 2 Jan 2024
32. Espinosa-Gongora C, Berg C, Rehn M, Varg JE, Dillner L, Latorre-Margalef N et al (2023) Early detection of the emerging SARS-CoV-2 BA.2.86 lineage through integrated genomic surveillance of wastewater and COVID-19 cases in Sweden, weeks 31 to 38 2023. Euro Surveill 28: 2300595. https://doi:10.2807/1560-7917.ES.2023.28.46.2300595.
Acknowledgements
We extend our gratitude to the National Culture Collection for Pathogens in Osong, Korea, for providing the SARS-CoV-2 Omicron.
Funding
This research was supported by a grant from the National Research Foundation (grant number: NRF-2022M3A9I2082292) funded by the Ministry of Science and ICT in the Republic of Korea and by a grant of the Korea Health Technology R&D Project (grant number: HV23C0052) through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare in the Republic of Korea.
Author information
Authors and Affiliations
Contributions
MSP, YL and HJK conceived the study and contributed to conceptualization and fund acquisition. DK, JK, MSP, YL and HJK designed experiments. KB, DK, JK, BMK, MK and SK performed experiments in BSL-3 and ABSL-3 facility including cell culture, viral cultivation and animal experiments. KB, DK, JK, SP, HES and MHL contributed to sample processing for DNA sequencing. KB, DK, JK, HP and BMK performed viral RNA extraction, analyzed whole-genome sequences data and prepared figures. KB, DK, JK and SP performed qRT-PCR. KB and BMK carried out H&E staining and immunohistochemistry. KB, DK, JK, HP, BMK, SP, HES, MHL, MK and SK contributed to methodology and formal analysis. JK, SM, YL and HJK contributed to Writing-original draft. YL and HJK contributed to Writing – review & editing. All authors contributed to editing of the manuscript and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethical approval
All animal research was performed following the guidelines provided by the Guide for the Care and Use of Laboratory Animals from the National Veterinary Research & Quarantine Service of Korea. The Institutional Animal Care and Use Committee of Hallym University (Permit no. Hallym2022-52) approved the animal experiments in this study.
Additional information
Edited by Juergen Richt.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Baek, K., Kim, D., Kim, J. et al. Analysis of SARS-CoV-2 omicron mutations that emerged during long-term replication in a lung cancer xenograft mouse model. Virus Genes (2024). https://doi.org/10.1007/s11262-024-02067-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11262-024-02067-6