Abstract
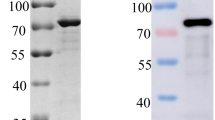
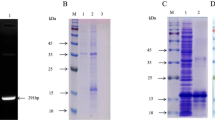
African swine fever virus (ASFV) is highly contagious and can cause lethal disease in pigs. ASFV p72 protein is a major capsid protein that presents as trimer in the virion. Epitopes on the surface of p72 trimer are considered as protective antigens. In this study, recombinant p72 protein and p72-baculovirus were constructed and obtained. Three monoclonal antibodies (mAbs) specific to ASFV p72 protein, designated as 1A3, 2B5 and 4A5, were generated. Among them, 4A5 showed strong reactivity with ASFV infected cells. Subsequently, the epitope recognized by 4A5 was mapped and identified using a series of overlapping peptides generated from p72 protein. IFA and western blot analyses showed that 4A5 recognized the linear epitope of p72 monomer located between amino acids 245–285 and recognized the conformational epitope located at the surface and top of the p72 trimer. These findings will enrich our knowledge regarding the epitope on p72 protein and provide valuable information for further characterization of the antigenicity and molecular functions of p72 protein.





Similar content being viewed by others
Data Availability
The data generated or analyzed during this study are included in this published article.
References
Dixon LK, Stahl K, Jori F, Vial L, Pfeiffer DU (2020) African swine fever epidemiology and control. Annu Rev Anim Biosci 8:221–246. https://doi.org/10.1146/annurev-animal-021419-083741
Ata EB, Li ZJ, Shi CW, Yang GL, Yang WT, Wang CF (2022) African swine fever virus: a raised global upsurge and a continuous threaten to pig husbandry. Microb Pathog 167:105561. https://doi.org/10.1016/j.micpath.2022.105561
Sanchez-Cordon PJ, Montoya M, Reis AL, Dixon LK (2018) African swine fever: a re-emerging viral disease threatening the global pig industry. Vet J 233:41–48. https://doi.org/10.1016/j.tvjl.2017.12.025
Zhou X, Li N, Luo Y, Liu Y, Miao F, Chen T, Zhang S, Cao P, Li X, Tian K, Qiu HJ, Hu R (2018) Emergence of African swine fever in China. Transbound Emerg Dis 65(6):1482–1484. https://doi.org/10.1111/tbed.12989
Alonso C, Borca M, Dixon L, Revilla Y, Rodriguez F, Escribano JM, Ictv Report C (2018) ICTV virus taxonomy profile: Asfarviridae. J Gen Virol 99(5):613–614. https://doi.org/10.1099/jgv.0.001049
Dixon LK, Chapman DA, Netherton CL, Upton C (2013) African swine fever virus replication and genomics. Virus Res 173(1):3–14. https://doi.org/10.1016/j.virusres.2012.10.020
Alejo A, Matamoros T, Guerra M, Andres G (2018) A proteomic atlas of the African swine fever virus particle. J Virol. https://doi.org/10.1128/JVI.01293-18
Salas ML, Andres G (2013) African swine fever virus morphogenesis. Virus Res 173(1):29–41. https://doi.org/10.1016/j.virusres.2012.09.016
Wang N, Zhao D, Wang J, Zhang Y, Wang M, Gao Y, Li F, Wang J, Bu Z, Rao Z, Wang X (2019) Architecture of African swine fever virus and implications for viral assembly. Science 366(6465):640–644. https://doi.org/10.1126/science.aaz1439
Revilla Y, Perez-Nunez D, Richt JA (2018) African swine fever virus biology and vaccine approaches. Adv Virus Res 100:41–74. https://doi.org/10.1016/bs.aivir.2017.10.002
Kollnberger SD, Gutierrez-Castaneda B, Foster-Cuevas M, Corteyn A, Parkhouse RME (2002) Identification of the principal serological immunodeterminants of African swine fever virus by screening a virus cDNA library with antibody. J Gen Virol 83(Pt 6):1331–1342. https://doi.org/10.1099/0022-1317-83-6-1331
Yu M, Morrissy CJ, Westbury HA (1996) Strong sequence conservation of African swine fever virus p72 protein provides the molecular basis for its antigenic stability. Arch Virol 141(9):1795–1802. https://doi.org/10.1007/BF01718302
Geng R, Sun Y, Li R, Yang J, Ma H, Qiao Z, Lu Q, Qiao S, Zhang G (2022) Development of a p72 trimer-based colloidal gold strip for detection of antibodies against African swine fever virus. Appl Microbiol Biotechnol 106(7):2703–2714. https://doi.org/10.1007/s00253-022-11851-z
Zhu W, Meng K, Zhang Y, Bu Z, Zhao D, Meng G (2021) Lateral flow assay for the detection of African swine fever virus antibodies using gold nanoparticle-labeled acid-treated p72. Front Chem 9:804981. https://doi.org/10.3389/fchem.2021.804981
Chen D, Wang D, Wang C, Wei F, Zhao H, Lin X, Wu S (2021) Application of an AlphaLISA method for rapid sensitive detection of African swine fever virus in porcine serum. Appl Microbiol Biotechnol 105(11):4751–4759. https://doi.org/10.1007/s00253-021-11339-2
Gomez-Puertas P, Rodriguez F, Oviedo JM, Ramiro-Ibanez F, Ruiz-Gonzalvo F, Alonso C, Escribano JM (1996) Neutralizing antibodies to different proteins of African swine fever virus inhibit both virus attachment and internalization. J Virol 70(8):5689–5694. https://doi.org/10.1128/jvi.70.8.5689-5694.1996
Liu S, Luo Y, Wang Y, Li S, Zhao Z, Bi Y, Sun J, Peng R, Song H, Zhu D, Sun Y, Li S, Zhang L, Wang W, Sun Y, Qi J, Yan J, Shi Y, Zhang X, Wang P, Qiu HJ, Gao GF (2019) Cryo-EM structure of the African swine fever virus. Cell Host Microbe 26(6):836–843. https://doi.org/10.1016/j.chom.2019.11.004
Liu Q, Ma B, Qian N, Zhang F, Tan X, Lei J, Xiang Y (2019) Structure of the African swine fever virus major capsid protein p72. Cell Res 29(11):953–955. https://doi.org/10.1038/s41422-019-0232-x
Achenbach JE, Gallardo C, Nieto-Pelegrin E, Rivera-Arroyo B, Degefa-Negi T, Arias M, Jenberie S, Mulisa DD, Gizaw D, Gelaye E, Chibssa TR, Belaye A, Loitsch A, Forsa M, Yami M, Diallo A, Soler A, Lamien CE, Sanchez-Vizcaino JM (2017) Identification of a new genotype of African swine fever virus in domestic pigs from Ethiopia. Transbound Emerg Dis 64(5):1393–1404. https://doi.org/10.1111/tbed.12511
Yin D, Geng R, Shao H, Ye J, Qian K, Chen H, Qin A (2022) Identification of novel linear epitopes in P72 protein of African swine fever virus recognized by monoclonal antibodies. Front Microbiol 13:1055820. https://doi.org/10.3389/fmicb.2022.1055820
Borca MV, Irusta P, Carrillo C, Afonso CL, Burrage T, Rock DL (1994) African swine fever virus structural protein p72 contains a conformational neutralizing epitope. Virology 201(2):413–418. https://doi.org/10.1006/viro.1994.1311
Wardley RC, Wilkinson PJ (1978) The growth of virulent African swine fever virus in pig monocytes and macrophages. J Gen Virol 38(1):183–186. https://doi.org/10.1099/0022-1317-38-1-183
Liu Y, Li Y, Xie Z, Ao Q, Di D, Yu W, Lv L, Zhong Q, Song Y, Liao X, Song Q, Wang H, Chen H (2021) Development and in vivo evaluation of MGF100–1R deletion mutant in an African swine fever virus Chinese strain. Vet Microbiol 261:109208. https://doi.org/10.1016/j.vetmic.2021.109208
Liu J, Mei N, Wang Y, Shi X, Chen H (2021) Identification of a novel immunological epitope on Hexon of fowl adenovirus serotype 4. AMB Express 11(1):153. https://doi.org/10.1186/s13568-021-01309-2
Muangkram Y, Sukmak M, Wajjwalku W (2015) Phylogeographic analysis of African swine fever virus based on the p72 gene sequence. Genet Mol Res 14(2):4566–4574. https://doi.org/10.4238/2015.May.4.15
Cobbold C, Windsor M, Wileman T (2001) A virally encoded chaperone specialized for folding of the major capsid protein of African swine fever virus. J Virol 75(16):7221–7229. https://doi.org/10.1128/JVI.75.16.7221-7229.2001
Epifano C, Krijnse-Locker J, Salas ML, Rodriguez JM, Salas J (2006) The African swine fever virus nonstructural protein pB602L is required for formation of the icosahedral capsid of the virus particle. J Virol 80(24):12260–12270. https://doi.org/10.1128/JVI.01323-06
Heimerman ME, Murgia MV, Wu P, Lowe AD, Jia W, Rowland RR (2018) Linear epitopes in African swine fever virus p72 recognized by monoclonal antibodies prepared against baculovirus-expressed antigen. J Vet Diagn Invest 30(3):406–412. https://doi.org/10.1177/1040638717753966
Acknowledgements
This research work was funded by the National Key Research and Development Program of China (2021YFD1801300 and 2021YFD1801401), National Nature Science Foundation (32170161) and Central Public-interest Scientific Institution Basal Research Fund (Y2022PT11). The authors would like to thank these.
Funding
This research work was funded by the National Key Research and Development Program of China (2021YFD1801300 and 2021YFD1801401), National Nature Science Foundation (32170161) and Central Public-interest Scientific Institution Basal Research Fund (Y2022PT11).
Author information
Authors and Affiliations
Contributions
The project was conceived and designed by HC. XD, YL, ZX et al. performed experiments. HC, JL, ZC and XD analyzed the data. JL and HC wrote the paper. HC supervised all the experiments. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Additional information
Edited by Juergen Richt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Duan, X., Liu, Y., Chen, Z. et al. Identification of monoclonal antibody targeting epitope on p72 trimeric spike of African swine fever virus. Virus Genes 59, 582–590 (2023). https://doi.org/10.1007/s11262-023-02003-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-023-02003-0