Abstract
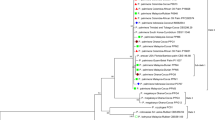
A distinct bipartite begomovirus was found associated with tomato plants showing yellowing, curling, and crumpling of the leaves, in a sub-temperate region in India. The complete DNA-A and DNA-B components were amplified through rolling circle amplification (RCA) using Φ-29 DNA polymerase and characterized. The DNA-A of the isolate was comprised of 2,756 nucleotides, encoding six open reading frames (ORFs) and DNA-B that of 2,725 nucleotides, encoding two ORFs. Genome organization of the isolate was typical of an old world bipartite begomovirus. Comparisons showed that DNA-A and its intergenic region (IR) have the highest sequence identity (86% and 84%, respectively) with the Tomato leaf curl New Delhi virus (ToLCNDV; DQ116885) and some other begomoviruses (>84%) reported from cucurbits and tomato. This data suggested that the isolate is a distinct begomovirus species for which a name Tomato leaf curl Palampur virus (ToLCPMV) is proposed. DNA-B showed the maximum sequence identity (73%) with Tomato leaf curl New Delhi virus-India-[Pakistan:Dargai:T5/6:2001] (AY150305). The common region (CR) of DNA-A and DNA-B showed 94% sequence similarity with each other. In the present study, phylogenetic relationship of this new species was also established with different begomoviruses reported from tomato and other begomoviruses showing highest homologies with complete DNA-A and DNA-B sequences. ToLCPMV is being reported from a sub-temperate region in India which was previously unaffected by begomoviruses and its whitefly vector.





Similar content being viewed by others
References
J. Stanley, D.M. Bisaro, R.W. Briddon, J.K. Brown, C.M. Fauquet, B.D. Harrison, et al., in Virus Taxonomy: VIII Report of the International Committee on Taxonomy of Viruses, ed. by C.M. Fauquet, M.A. Mayo, J. Maniloff, U. Desselberger, L.A. Ball, (Elsevier Academic Press, London, 2005), pp. 301–326
M. Padidam, R.N. Beachy, C.M. Fauquet, J. Gen. Virol. 76, 249–263 (1995). doi:10.1099/0022-1317-76-2-249
L. Hanley-Bowdoin, S.B. Settlage, B.M. Orozco, S. Nagar, D. Robert, Crit. Rev. Plant Sci. 18, 71–106 (1999). doi:10.1016/S0735-2689(99)00383-4
A.S. Moffat, Science 286, 1835 (1997). doi:10.1126/science.286.5446.1835
X.F. Cui, X.R. Tao, Y. Xie, C.M. Fauquet, X.P. Zhou, J. Virol. 78, 13966–13974 (2004). doi:10.1128/JVI.78.24.13966-13974.2004
R.W. Briddon, S. Mansoor, I.D. Bedford, M.S. Pinner, K. Saunders, J. Stanley, Virology 285, 234–243 (2001). doi:10.1006/viro.2001.0949
K. Saunders, A. Norman, S. Gucciardo, J. Stanley, Virology 324, 37–47 (2004). doi:10.1016/j.virol.2004.03.018
R.W. Briddon, J. Stanley, Virology 344, 198–210 (2006). doi:10.1016/j.virol.2005.09.042
S. Mansoor, R.W. Briddon, Y. Zafar, J. Stanley, Trends Plant Sci. 8, 128–134 (2003). doi:10.1016/S1360-1385(03)00007-4
X.P. Zhou, Y. Xie, Z.K. Zhang, Arch. Virol. 146, 1599–1606 (2001). doi:10.1007/s007050170081
Y. Xie, X.P. Zhou, Z.K. Zhang, Y.J. Qi, Chin. Sci. Bull. 47, 197–200 (2002). doi:10.1360/02tb9047
V.N. Fondong, J.S. Pita, M.E.C. Rey, A. de Kochko, R.N. Beachy, C.M. Fauquet, J. Gen. Virol. 81, 287–297 (2000)
M. Padidam, S. Sawyer, C.M. Fauquet, Virology 265, 218–225 (1999). doi:10.1006/viro.1999.0056
A.K. Inoue-Nagata, L.C. Albuquerque, W.B. Rocha, T. Nagata, J. Virol. Methods 116, 209–211 (2004). doi:10.1016/j.jviromet.2003.11.015
D. Haible, S. Kober, H. Jeske, J. Virol. Methods 135, 9–16 (2006). doi:10.1016/j.jviromet.2006.01.017
D. Knierim, E. Maiss, Arch. Virol. 152, 941–954 (2007). doi:10.1007/s00705-006-0914-910.1007/s00705-006-0914-9
S.L. Dellaporta, J. Wood, J.B. Hicks, Plant Mol. Biol. Rep. 1, 19–21 (1983). doi:10.1007/BF02712670
M.R. Rojas, R.L. Gilbertson, D.R. Russell, D.P. Maxwell, Plant Dis. 77, 340–347 (1993)
S.F. Altschul, W. Gish, W. Miller, E.W. Myers, D.J. Lipman, J. Mol. Biol. 215, 403–410 (1990)
S. Kumar, K. Tamura, M. Nei, Molecular Evolutionary Genetics Analysis Ver.1.02 and Tree Explorer Ver.2.12. (1993), http://www.megasoftware.net/
T. Vincze, J. Posfai, R.J. Roberts, Nucleic Acids Res. 31, 3688–3691 (2003). doi:10.1093/nar/gkg526
C.M. Fauquet, J. Stanley, Arch. Virol. 150, 2151–2179 (2005). doi:10.1007/s00705-005-0583-0
C.M. Fauquet, R.W. Briddon, J.K. Brown, E. Moriones, J. Stanley, M. Zerbini et al., Arch. Virol. 153, 783–821 (2008). doi:10.1007/s00705-008-0037-6
T. Ferreira Pde, T.O. Lemos, T. Nagata, A.K. Inoue-Nagata, J. Virol. Methods 147(2), 351–354 (2008). doi:10.1016/j.jviromet.2007.10.001
W. Chia-Ying, L. Yi-Chin, L. Na-Sheng, H. Yau-Heiu, T. Hsin-Tzu, L. Jye-Yann et al., J. Virol. Methods 147(2), 355–359 (2008). doi:10.1016/j.jviromet.2007.10.002
Acknowledgments
The authors are thankful to the Director, Institute of Himalayan Bioresource Technology (CSIR), Palampur, India for providing necessary research facilities and the Council of Scientific and Industrial Research (Govt. of India) for granting Senior Research Fellowship to Mr. Yogesh Kumar.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kumar, Y., Hallan, V. & Zaidi, A.A. Molecular characterization of a distinct bipartite begomovirus species infecting tomato in India. Virus Genes 37, 425–431 (2008). https://doi.org/10.1007/s11262-008-0286-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-008-0286-1