Abstract
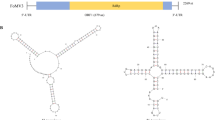
Okra mosaic virus (OkMV) is a tymovirus infecting members of the family Malvaceae. Early infections in okra (Abelmoschus esculentus) lead to yield losses of 12–19.5%. Besides intensive biological characterizations of OkMV only minor molecular data were available. Therefore, we determined the complete nucleotide sequence of a Nigerian isolate of OkMV. The complete genomic RNA (gRNA) comprises 6,223 nt and its genome organization showed three major ORFs coding for a putative movement protein (MP) of Mr 73.1 kDa, a large replication-associated protein (RP) of Mr 202.4 kDa and a coat protein (CP) of Mr 19.6 kDa. Prediction of secondary RNA structures showed three hairpin structures with internal loops in the 5′-untranslated region (UTR) and a 3′-terminal tRNA-like structure (TLS) which comprises the anticodon for valine, typical for a member of the genus Tymovirus. Phylogenetic comparisons based on the RP, MP and CP amino acid sequences showed the close relationship of OkMV not only to other completely sequenced tymoviruses like Kennedya yellow mosaic virus (KYMV), Turnip yellow mosaic virus (TYMV) and Erysimum latent virus (ErLV), but also to Calopogonium yellow vein virus (CalYVV), Clitoria yellow vein virus (CYVV) and Desmodium yellow mottle virus (DYMoV). This is the first report of a complete OkMV genome sequence from one of the various OkMV isolates originating from West Africa described so far. Additionally, the experimental host range of OkMV including several Nicotiana species was determined.




Similar content being viewed by others
References
G.P. Martelli, S. Sabanadzovic, N.A.G. Sabanadzovic, M.C. Edwards, T.W. Dreher, Arch. Virol. 147, 1837 (2002)
T.W. Dreher, M.C. Edwards, A.J. Gibbs, A.-L. Haenni, R.W. Hammond, I. Jupin, R. Koenig, S. Sabanadzovic, N. Abou Ghanem-Sabanadzovic, G.P. Martelli, in Virus Taxonomy: Eighth Report of the International Committee on Taxonomy of Viruses ed. by C.M. Fauquet, M.A. Mayo, J. Maniloff, U. Desselberger, L.A. Ball (Elsevier Academic Press, San Diego, 2005), pp. 1067–1076
G.P. Martelli, S. Sabanadzovic, N. Abouh-Ghanem Sabanadzovic, P. Saldarelli, Arch Virol 147, 1847 (2005)
K.L. Bransom, J.J. Weiland, T.W. Dreher, Virology 184, 351 (1991)
L. Givord, L. Hirth, Ann. Appl. Biol. 74, 359 (1973)
A.O. Lana, R.F. Bozarth, Phytopath. Z. 83, 77 (1975)
E.C.K. Igwegbe, Plant Dis. 67, 320 (1983)
E.C.K. Igwegbe, A.D. Hewings, V.D. Damsteegt, W.M. Dowler, Phytopathology 77, 987 (1987)
G. Thottappilly, Acta Virol. 40, 233 (1996)
R.F. Bozarth, A.O. Lana, R. Koenig, J. Reese, Phytopathology 67, 735 (1977)
L. Givord, Ann. Phytopathol. 9, 53 (1977)
L. Givord, L. Den Boer, Ann. Appl. Biol. 94, 235 (1980)
A.O. Lana, T.A. Taylor, Ann. Appl. Biol. 82, 361 (1976)
G.I. Atiri, M.F. Ivbijaro, A.D. Oladele, Trop. Agri. 68, 178 (1991)
J. Ndunguru, A.C. Rajabu, Scientia Hort. 99, 225 (2004)
T.C. Verwoerd, B.M. Dekker, A. Hoekema, Nucleic Acids Res. 25, 2362 (1980)
I. Bekesiova, J.P. Nap, L. Mlynarova, Plant Mol. Biol. Rep. 17, 269 (1999)
A.J. Dodds, J.T. Morris, L.R. Jordan, Ann. Rev. Phytopathol. 22, 151 (1984)
A.J. Dodds, in DsRNA in diagnosis, ed. by R.E.F. Matthews (CRC Press, Florida, 1993), pp. 274–289
A. Gibbs, A.M. Mackenzie, Meth. Mol. Biol. 81, 219 (1998)
M. Matz, D. Shagin, E. Bogdanova, O. Britanova, S. Lukyanov, L. Diatschenko, A. Chenchik, Nucleic Acids Res. 27, 1558 (1999)
J.D. Parker, P.S. Rabinovitch, G.C. Burmer, Nucleic Acids Res. 19, 3055 (1991)
T.M. Rose, Virol. J. 2, 20 (2005)
S.F. Altschul, T.L. Madden, A.A. Schäffer, Nucleic Acids Res. 25, 3389 (1997)
J.D. Thompson, T.J. Gibson, F. Plewniak, F. Jeanmougin, D.G. Higgins, Nucleic Acids Res. 25, 4876 (1997)
R.D.M. Page, Comput. Appl. Biosci. 12, 357 (1996)
J. Reeder, R. Giegerich, BMC Bioinf. 5, 104 (2004)
Y. Byun, K. Han, Nucleic Acids Res. 34 (web server issue), W416 (2006)
M.N. Rozanov, E.V. Koonin, A.E. Gorbalenya, J. Gen. Virol. 73, 2129 (1992)
M.N. Rozanov, G. Drugeon, A.L. Haenni, Arch. Virol. 140, 273 (1995)
E.V. Koonin, V.V. Dolja, Crit. Rev. Biochem. Mol. Biol. 28, 375 (1993)
M. Pagni, V. Ioannidis, L. Cerutti, M. Zahn-Zabal, C.V. Jongeneel, L. Falquet, Nucleic Acids Res. 32(web server issue), W332 (2004)
R. Koenig, C.W. Pleij, D.E. Lesemann, S. Loss, H.J. Vetten, Arch. Virol. 150, 2325 (2005)
S. Ding, J. Howe, P. Keese, A. Mackenzie, D. Meek, M. Osorio-Keese, M. Skotnicki, P. Srifah, M. Torronen, A. Gibbs, Nucleic Acids Res. 18, 1181 (1990)
J. Schirawski, A. Voyatzakis, B. Zaccomer, F. Bernardi, A.L. Haenni, J. Virol. 74, 11073 (2000)
K. Hellendoorn, P.J.A. Michiels, R. Buitenhuis, C.W.A. Pleij, Nucleic Acids Res. 24, 4910 (1996)
H.H. Bink, J. Schirawski, A.L. Haenni, C.W. Pleij, J. Virol. 77, 7452 (2003)
T.W. Dreher, J.B. Goodwin, Nucleic Acids Res. 26, 4356 (1998)
D. Matsuda, T.W. Dreher, Virology 321, 36 (2004)
M.H. de Smit, A.P. Gultyaev, M. Hilge, H.H. Bink, S. Barends, B. Kraal, C.W. Pleij, Nucleic Acids Res. 30, 4232–4240 (2002)
E.J. Mitchell, J.M. Bond, Arch. Virol. 150, 2347 (2005)
J.P. Briand, G. Jonard, H. Guilley, K. Richards, L. Hirth, Eur. J. Biochem. 72, 453 (1977)
M. Silberklang, A. Prochiantz, A.L. Haenni, U.L. Rajbhandary, Eur. J. Biochem. 72, 465 (1977)
T.W. Dreher, Mol. Plant Pathol. 5, 367 (2004)
C.T. Ranjith-Kumar, K. Gopinath, A.N.K. Jacob, V. Srividhya, P. Elango, H.S. Savithri, Arch. Virol. 143, 148 (1998)
P.L. Guy, J.L. Dale, M.A. Adena, A.J. Gibbs, Plant Pathol. 33, 337 (1984)
A.O. Lana, R.M. Gilmer, H.D. Cheda, D.O. Fatokun, Plant Dis. Rep. 58, 616 (1974)
J.J. Bernal, I. Jiménez, M. Moreno, M. Hord, C. Rivera, R. Koenig, E. Rodríguez-Cerezo, Phytopathology 90, 1098 (2000)
M.A.V. Alexandre, L.M.L. Duarte, E.B. Rivas, C.M. Chagas, M.M. Barradas, R. Koenig, Plant Dis. 84, 739 (2000)
C.M. Hayden, A.M. Mackenzie, A.J. Gibbs, Arch. Virol. 143, 191 (1998)
Acknowledgements
We thank Dirk Bellstedt, Kees Pleij and Theo Dreher for scientific discussion and Agnes Pietruszka and Marianne Koerbler for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
The nucleotide sequence data reported in this article have been submitted to the Genbank nucleotide sequence database and have been assigned the accession number EF554577.
Rights and permissions
About this article
Cite this article
Stephan, D., Siddiqua, M., Ta Hoang, A. et al. Complete nucleotide sequence and experimental host range of Okra mosaic virus . Virus Genes 36, 231–240 (2008). https://doi.org/10.1007/s11262-007-0181-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-007-0181-1