Abstract
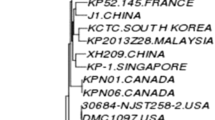
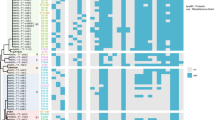
Urinary tract infections (UTIs) are pervasive in human and veterinary medicine, notably affecting companion animals. These infections frequently lead to the prescription of antibiotics, contributing to the rise of antimicrobial-resistant bacteria. This escalating concern is underscored by the emergence of a previously undocumented case: a high-risk clone, broad-spectrum cephalosporin-resistant K. pneumoniae ST147 strain, denoted USP-275675, isolated from a cat with UTI. Characterized by a multidrug-resistant (MDR) profile, whole genome sequencing exposed several antimicrobial-resistance genes, notably blaCTX-M-15, blaTEM-1B, blaSHV-11, and blaOXA-1. ST147, recognized as a high-risk clone, has historically disseminated globally and is frequently associated with carbapenemases and extended-spectrum β-lactamases. Notably, the core-genome phylogeny of K. pneumoniae ST147 strains isolated from urine samples revealed a unique aspect of the USP-276575 strain. Unlike its counterparts, it did not cluster with other isolates. However, a broader examination incorporating strains from both human and animal sources unveiled a connection between USP-276575 and a Portuguese strain from chicken meat. Both were part of a larger cluster of ST147 strains spanning various geographic locations and sample types, sharing commonalities such as IncFIB or IncR plasmids. This elucidates the MDR signature inherent in widespread K. pneumoniae ST147 strains carrying these plasmids, highlighting their pivotal role in disseminating antimicrobial resistance (AMR). Finally, discovering the high-risk clone K. pneumoniae ST147 in a domestic feline with a UTI in Brazil highlights the urgent need for thorough AMR surveillance through a One Health approach.


Similar content being viewed by others
Data Availability
Sequence data that support the findings of this study have been deposited in the Genbank with the primary accession code JAYMFY000000000.
References
Antimicrobial Resistance Collaborators (2022) Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. Lancet 399:629–655. https://doi.org/10.1016/S0140-6736(21)02724-0
Barsanti JA (2006) Genitourinary infections. In: Greene CE (ed) Infectious diseases of the dog and cat. Saunders/Elsevier, St Louis
Byron JK (2018) Urinary tract infection. Vet Clin Small Anim 49:211–221. https://doi.org/10.1016/j.cvsm.2018.11.005
Carvalho I, Safia Chenouf N, Cunha R, Martins C, Pimenta P, Pereira AR, Martínez-Álvarez S, Ramos S, Silva V, Igrejas G, Torres C, Poeta P (2021) Antimicrobial resistance genes and diversity of clones among ESBL- and acquired AmpC-producing Escherichia coli isolated from fecal samples of healthy and sick cats in Portugal. Antibiotics (Basel) 10(3):262. https://doi.org/10.3390/antibiotics10030262
Clegg S, Murphy CN (2012) Epidemiology and Virulence of Klebsiella pneumoniae. Microbiol Spect 4. https://doi.org/10.1128/microbiolspec.uti-0005-2012
CLSI (2021) Performed standards for antimicrobial susceptibility testing, 31st edn. CLSI supplement M100. Clinical and Laboratory Standards Institute. https://doi.org/10.1016/j.onehlt.2021.100236
Davies YM, Cunha MPV, Dropa M, Lincopan N, Gomes VTM, Moreno LZ, Sato MIZ, Moreno AM, Knöbl T (2022) Pandemic Clones of CTX-M-15 Producing Klebsiella pneumoniae ST15, ST147, and ST307 in Companion Parrots. Microorganisms 10:1412. https://doi.org/10.3390/microorganisms10071412
Doster E, Lakin SM, Dean CJ et al (2020) MEGARes 2.0: a database for classification of antimicrobial drug, biocide and metal resistance determinants in metagenomic sequence data. Nucleic Acids Res 48:561–569. https://doi.org/10.1093/nar/gkz1010
Ewers C, Stamm I, Pfeifer Y, Wieler LH, Kopp PA, Schønning K, Prenger-Berninghoff E, Scheufen S, Stolle I, Günther S, Bethe A (2014) Clonal spread of highly successful ST15-CTX-M-15 Klebsiella pneumoniae in companion animals and horses. J Antimicrob Chemother 69(10):2676–2680. https://doi.org/10.1093/jac/dku217
Jericó MM, Lorenzini F, Kanayama K (2014) Manual de obesidade felina. Manual de Obesidade canina e felina, ABEV, São Paulo
Jin M, Osman M, Green BA et al (2023) Evidence for the transmission of antimicrobial resistant bacteria between humans and companion animals: A scoping review. One Health 17:100593. https://doi.org/10.1016/j.onehlt.2023.100593
Marques C, Menezes J, Belas A et al (2019) Klebsiella pneumoniae causing urinary tract infections in companion animals and humans: population structure, antimicrobial resistance and virulence genes. J Antimicrob Chemother 74:594–602. https://doi.org/10.1093/jac/dky499
Nóbrega DB, Brocchi M (2014) An overview of extended-spectrum beta-lactamases in veterinary medicine and their public health consequences. J Infect Dev Ctries 8(8):954–960. https://doi.org/10.3855/jidc.4704
Ovejero CM, Escudero JA, Thomas-Lopez D et al (2017) Highly tigecycline-resistant Klebsiella pneumoniae sequence type 11 (ST11) and ST147 isolates from companion animals. Antimicrob Agents Chemother 61:e02640–e02716. https://doi.org/10.1128/AAC.02640-16
Pal C, Bengtsson-Palme J, Rensing C et al (2014) BacMet: antibacterial biocide and metal resistance genes database. Nucleic Acids Res 42:737–743. https://doi.org/10.1093/nar/gkt1252
Peirano G, Chen L, Kreiswirth BN et al (2020) Emerging antimicrobial-resistant high-risk Klebsiella pneumoniae clones ST307 and ST147. Antimicrob Agents Chemother 64:e01148–e01220. https://doi.org/10.1128/aac.01148-20
Prestinaci F, Pezzotti P, Pantosti A (2015) Antimicrobial resistance: a global multifaceted phenomenon. Pathog Glob Health 109:309–318. https://doi.org/10.1179/2047773215Y.0000000030
Rocha LKL, Santos Neto RL, Guimarães ACC et al (2014) Plasmid-mediated qnrA1 in Klebsiella pneumoniae ST147 in Recife, Brazil. Int J Infect Dis 26:49–50. https://doi.org/10.1016/j.ijid.2014.04.026
Rodrigues C, Machado E, Ramos H, Peixe L, Novais  (2014) Expansion of ESBL-producing Klebsiella pneumoniae in hospitalized patients: a successful story of international clones (ST15, ST147, ST336) and epidemic plasmids (IncR, IncFIIK). Int J Med Microbiol 304(8):1100–1108. https://doi.org/10.1016/j.ijmm.2014.08.003
Rosen DA, Hilliard JK, Tiemann KM et al (2016) Klebsiella pneumoniae FimK promotes virulence in murine pneumonia. J Infect Dis 213:649–658. https://doi.org/10.1093/infdis/jiv440
Rossolini GM, D’Andrea MM, Mugnaioli C (2008) The spread of CTX-M-type extended-spectrum beta-lactamases. Clin Microbiol Infect 14:33–41. https://doi.org/10.1111/j.1469-0691.2007.01867.x
Salgado-Caxito M, Benavides JA, Adell AD, Paes AC, Moreniswitt AI (2021) Global prevalence and molecular characterization of extended-spectrum β-lactamase producing-Escherichia coli in dogs and cats - A scoping review and metaanalysis. One Health 12:100236. https://doi.org/10.1016/j.onehlt.2021.100236
Tarkkanen AM, Virkola R, Clegg S et al (1997) Binding of the type 3 fimbriae of Klebsiella pneumoniae to human endothelial and urinary bladder cells. Infect Immun 65:1546–1549
Umeda K, Hase A, Fukuda A, Matsuo M, Horimoto T, Ogasawara J (2019) Prevalence and mechanisms of fluoroquinolone-resistant Escherichia coli among sheltered companion animals. Access Microbiol 2(1):acmi000077. https://doi.org/10.1099/acmi.0.000077
Vázquez-Jaramillo L, Cardozo-Herrera LK, Valencia NMPC (2023) A systematic review of tetracycline resistance genes in animals and derived products in Latin America and the Caribbean. Braz J Vet Res Anim Sci 60:e213883. https://doi.org/10.11606/issn.1678-4456.bjvras.2023.213883
Acknowledgements
The authors thank Claudia Stricagnolo for her valuable technical support, the São Paulo Research Foundation (FAPESP), and the National Council for Scientific and Technological Development (CNPq) for funding support. V.T.S.K. and N.C.G. are research fellows of FAPESP (2021/09813-0 and 2020/15008-0, respectively). M.B.H. is a research fellow of CNPq (310462/2021).
Author information
Authors and Affiliations
Contributions
V.T.S.S.: performed research and analyzed data A.H. and J.P.A.J. performed research and wrote the paper JSFN: wrote the paper M.B.H and N.C.G. conceived of or designed study, performed funding acquisition and supervision and wrote the paper.
Corresponding author
Ethics declarations
Animal ethics
All procedures were carried out in agreement with the Ethic Committee on Animal Use of the School of Veterinary Medicine and Animal Science (University of São Paulo) (CEUA/FMVZ) (# 2475050821).
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sakauchi, V.T.S., Haisi, A., Araújo Júnior, J.P. et al. Genomic analysis of the first multidrug-resistant ESBL-producing high-risk clonal lineage Klebsiella pneumoniae ST147 isolated from a cat with urinary tract infection. Vet Res Commun (2024). https://doi.org/10.1007/s11259-024-10396-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11259-024-10396-y