Abstract
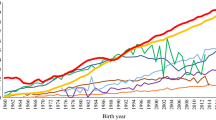
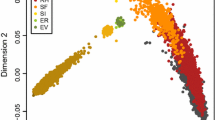
A vital requirement to design and implement a breeding program is to know the structure and genetic diversity of a population. The aim of this study was to characterize population structure and genetic diversity of the Colombian Simmental cattle. The pedigree file included 27,985 animals born from 1975 to 2017. The level of genetic diversity and breed structure was evaluated through probabilities of gene origin expressed via effective number of founders, ancestors and founders genomes. The inbreeding rates and the degree of genetic connectivity were estimated using a regression analysis and a genetic drift variance analysis, respectively. The lowest effective number of founders and ancestors were 50 and 38 by year, respectively. The average inbreeding by year of birth decreased from 5.06% in 1980 to 2.25% in 2017. The dairy line genetic contributions in the overall population increased significantly in the last 37 years, and the beef line contribution decreased. Regarding the genetic connectivity, Colombian regions (administrative divisions) with the largest cattle population had higher values. The results indicate that the availability of European and North American bulls contributes to genetic diversity by increasing the effective number of founders over time in the Colombian Simmental cattle population. However, the intensive use of relatively few founders causes an unbalanced genetic contribution and the loss of genetic diversity by gene pool erosion.




Similar content being viewed by others
References
Agung, P. P., Saputra, F., Septian, W. A., Syamsul, M, Zein, A. 2016. Study of genetic diversity among Simmental cross cattle in West Sumatra based on microsatellite markers, Asian-Australasian Journal of Animal Sciences, 29, 176-183.
American Simmental Association. History of the Simmental Breed 2018a. http://www.simmental.org/site/userimages/HistoryoftheSimmentalBreed.pdf. Accessed 15 Jul 2018.
American Simmental Association. SimGenetics Herdbook. 2018b. Retrieved from https://herdbook.org/simmapp/template/animalSearch,-AnimalSearch.vm. Accessed 15 Jul 2018.
Boichard, D. 2002. Pedig: a fortran package for pedigree analysis suited for large populations. 7th World Congress on Genetics Applied to Livestock Production.
Boichard, D., Maignel, L., Verrier, É. 1997. The value of using probabilities of gene origin to measure genetic variability in a population, Genetics Selection Evolution: GSE, 29, 5-23.
Borges, A., Mendes, C., Souza, P., Santos, L., Pagung, D., Carrillo, J., Martins, F. R. 2013. Population structure of Nellore cattle in northeastern Brazil, Revista Brasileira de Zootecnia, 42, 639-644.
Bouquet, A., Venot, E., Laloë, D., Forabosco, F., Fogh, A., Pabiou, T., Phocas, F. 2011. Genetic structure of the European Charolais and Limousin cattle meta-populations using pedigree analyses, Journal of Animal Science, 89, 1719-1730.
Crow, J., Kimura, M. 1970. An Introduction to Population Genetics Theory. University of Wisconsin, Madison, USA.
Gutierrez, J. P., Altarriba, J., Diaz, C., Quintanilla, R., Cañón, J., Piedrafita, J. 2003. Pedigree analysis of eight Spanish beef cattle breeds, Genetics Selection Evolution: GSE, 35, 1-21.
Hammami, H., Croquet, C., Stoll, J., Rekik, B., Gengler, N. 2007. Genetic diversity and joint-pedigree analysis of two importing Holstein populations, Journal of Dairy Science, 90, 3530-3541.
Hanocq, E., Boichard, D. 1999. Connectedness in the French Holstein cattle population, Genetics Selection Evolution: GSE, 31, 163-176.
Hernández-Castellano, L. E., Nalyy, J. E., Lindahl, J., Wanapat, M., Alhidary, I. A., Fangueiro, D., Grace, D., Ratto, M., Bambou, J. C., de Almeida, A, M. 2019. Dairy science and health in the tropics: challenges and opportunities for the next decades, Tropical Animal Health and Production, 51, 1009-1017.
Howard, J. T., Haile-mariam, M., Pryce, J. E., Maltecca, C. 2015. Investigation of regions impacting inbreeding depression and their association with the additive genetic effect for United States and Australia Jersey dairy cattle, BMC Genomics, 16: 813.
Kasinathan, P., Wei, H., Xiang, T., Molina, J. A., Metzger, J., Broek, D., Allan, M. F. 2015. Acceleration of genetic gain in cattle by reduction of generation interval. Scientific Reports, 5, 8674.
Kennedy, B., Trus, D. 1993. Considerations on genetic connectedness between management units under an animal model, Journal of Animal Science, 71, 2341-2352.
Lacy, R. C. 1989. Analysis of founder representation in pedigrees : founder equivalents and founder genome equivalents, Zoo Biology, 123, 111-123.
Leroy, G., Mary-Huard, T., Verrier, É., Danvy, S., Charvolin, E., Danchin-Burge, C. 2013. Methods to estimate effective population size using pedigree data: Examples in dog, sheep, cattle and horse, Genetics Selection Evolution: GSE, 45, 1-10.
Magalhaes, M., Mendes, C., Costa, J., Araujo, J., Napoles, C., Souza, P. 2016. Population genetic structure in the Holstein breed in Brazil, Tropical Animal Health and Production, 48, 331-336.
Maiwashe, A., Nephawe, K. A., Westhuizen, R. R., Van Der Mostert, B. E., Theron, H. E. 2006. Rate of inbreeding and effective population size in four major South African dairy cattle breeds, South African Journal of Animal Science, 36, 50-57.
Malhado, C. H. M., Carneiro, P. L. S., Malhado, A. C. M., Martins Filho, R., Bozzi, R., Ladle, R. J. 2010. Genetic improvement and population structure of the Nelore breed in Northern Brazil, Pesquisa Agropecuária Brasileira, 45, 1109-1116.
Marras, G., Gaspa, G., Sorbolini, S., Dimauro, C., Ajmone-Marsan, P., Valentini, A., MacCiotta, N. P. P. 2015. Analysis of runs of homozygosity and their relationship with inbreeding in five cattle breeds farmed in Italy, Animal Genetics, 46, 110-121.
Maximini, L., Fuerst-Waltl, B., Gredler, B., Baumung, R. 2011. Inbreeding depression on semen quality in Austrian dual-purpose Simmental bulls, Reproduction in Domestic Animals, 46, 102-104.
Mc Parland, S., Kearney, J. F., Rath, M., Berry, D. P. 2007. Inbreeding trends and pedigree analysis of Irish dairy and beef cattle populations. Journal of Animal Science, 85, 322-331.
Pegolo, N., Laloë, D., Oliveira, H., Lobo, R., Fouilloux, M. 2011. Trends of the genetic connectedness measures among Nelore beef cattle herds, Journal of Animal Breeding and Genetics, 129, 20-29.
R Core Time. 2014. R: A Language and Environment for Statistical Computing [Internet]. Vienna, Austria. Available: http://www.r-project.org/.
Rinderzucht Austria. 2018. Zuchtwertdatenbank. http://cgi.zar.at/cgi-bin/zw_default.pl. Accessed 15 Jul 2018.
Santana Jr, M. L., Pereira, R. J., Bignardi, A. B., Ayres, D. R., Menezes, G. R. O., Silva, L. O. C., Albuquerque, L. G. 2016. Structure and genetic diversity of Brazilian Zebu cattle breeds assessed by pedigree analysis, Livestock Science, 187, 6-15.
Sharma, R., Kishore, A., Mukesh, M., Ahlawat, S., Maitra, A., Pandey, A.K., Tantia, M. S. 2015. Genetic diversity and relationship of Indian cattle inferred from microsatellite and mitochondrial DNA markers, BMC Genetics, 16, 1-12.
Simmental Swiss Genetics. 2018. Swissgenetic. http://www.swissgenetics.com/Simmental. Accessed 15 Jul 2018.
Sölkner, J., Filipcic, L., Hampshire, N. 1998. Genetic variability of populations and similarity of subpopulations in Austrian cattle breeds determined by analysis of pedigrees, Animal Science, 67, 249-256.
Stachowicz, K., Sargolzaei, M., Miglior, F., Schenkel, F. S. 2011. Rates of inbreeding and genetic diversity in Canadian Holstein and Jersey cattle, Journal of Dairy Science, 94, 5160-5175.
VanRaden, P. M. 1992. Accounting for inbreeding and crossbreeding in genetic evaluation of large populations, Journal of Dairy Science, 75, 3136–3144.
Vozzi, P. A., Marcondes, C. R., Magnabosco, C. D. U., Antonio, L., Bezerra, F., Lôbo, R. B. 2006. Structure and genetic variability in Nellore (Bos indicus) cattle by pedigree analysis, Genetics and Molecular Biology, 485, 482-485.
Whitacre, L. K., Spangler, M. L. 2012. The Simmental breed: population structure and generation interval, Nebraska Beef Cattle Reports, Animal Science Department, 1, 1-3.
Acknowledgements
The authors wish to thank the Asociación Colombiana de Criadores de Ganado Simmental and the Departamento de Genética de la Corporación Colombiana Investigación Agropecuaria (AGROSAVIA) in Tibaitatá. The authors would also like to thank the research group GaMMA of the Universidad de Antioquia.
Funding
Universidad de Ciencias Aplicadas y Ambientales U.D.C.A and Colciencias (scholarship No 727 of 2015) financed the first author’s DSc studies.
Author information
Authors and Affiliations
Contributions
The work was designed by AA, RM and MCM. The statistical analyses involved AA, RM and MCM. The manuscript was written by AA and proofread by RM and MCM. All the authors read and approved the final version.
Corresponding author
Ethics declarations
This article does not contain any studies with animals performed by any of the authors.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Amaya, A., Martínez, R. & Cerón-Muñoz, M. Population structure and genetic diversity in Colombian Simmental cattle. Trop Anim Health Prod 52, 1133–1139 (2020). https://doi.org/10.1007/s11250-019-02111-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11250-019-02111-w