Abstract
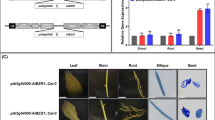
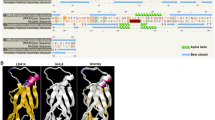
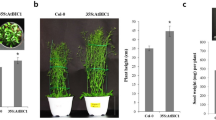
Seedling stand establishment is a critical factor affecting crop yield in low-precipitation agricultural regions. This is especially true for small seeded crops, such as Camelina (Camelina sativa) and canola (Brassica napus), that need to be planted shallow. Deeper planting would be desirable so that seeds can access soil moisture and bigger seeds could improve emergence and stand establishment by providing the energy necessary for seedling elongation. AHL (AT-Hook Containing, Nuclear Localized) genes play an important role in seedling growth and development. AHL proteins contain two structural units, the DNA-binding AT-hook motif and the Plant and Prokaryote Conserved (PPC) domain, required for protein–protein interactions. Our previous studies demonstrate that AtAHL29/SOB3 (Suppressor of phytochrome B-4 #3) regulates seedling development in Arabidopsis (Arabidopsis thaliana). Activation-tagged overexpression of AtSOB3 (Atsob3-D) represses the long-hypocotyl phenotype of an Arabidopsis phytochrome B mutant. In contrast, overexpression of the Atsob3-6 variant (Atsob3-6-OX), with a non-functional AT-hook, confers a long-hypocotyl phenotype. In this study, we demonstrate the role of Atsob3-D and Atsob3-6-OX in modulating seed size and hypocotyl length in the brassicas Arabidopsis and Camelina. In Arabidopsis, Atsob3-D reduces seed weight whereas Atsob3-6-OX increases seed weight and size when compared to the wild type. Similarly, Atsob3-6-OX transgenic Camelina seedlings are taller than the wild type, and produce larger and heavier seeds. These larger Atsob3-6-OX Camelina seeds also confer better emergence in deep-soil planting when compared to the wild type. Taken together, Atsob3-6-OX increases seed size, seed weight, seedling hypocotyl length and stand establishment in the oilseed crop Camelina.





Similar content being viewed by others
References
Ahmed SU, Zuberi MI (1973) Effects of Seed Size on Yield and some of its Components in Rapeseed, Brassica Campestris L. Var. Toria1. Crop Sci 13:119. https://doi.org/10.2135/cropsci1973.0011183X001300010039x
Aravind L (1998) AT-hook motifs identified in a wide variety of DNA-binding proteins. Nucleic Acids Res 26:4413–4421. https://doi.org/10.1093/nar/26.19.4413
Bertholdsson N-O, Kolodinska Brantestam A (2009) A century of Nordic barley breeding—Effects on early vigour root and shoot growth, straw length, harvest index and grain weight. Eur J Agronomy 30:266–274. https://doi.org/10.1016/j.eja.2008.12.003
Bodnaryk RP, Lamb RJ (1991) Influence of seed size in canola, Brassica napus L. and mustard, Sinapis alba L, on seedling resistance against flea beetles, Phyllotreta cruciferae (Goeze). Can J Plant Sci 71:397–404. https://doi.org/10.4141/cjps91-055
Elliott RH, Franke C, Rakow GFW (2008) Effects of seed size and seed weight on seedling establishment, vigour and tolerance of Argentine canola (Brassica napus) to flea beetles, Phyllotreta spp. Can J Plant Sci 88:207–217. https://doi.org/10.4141/CJPS07059
Favero DS, Jacques CN, Iwase A, Le KN, Zhao J, Sugimoto K, Neff MM (2016) SUPPRESSOR OF PHYTOCHROME B4-#3 Represses Genes Associated with Auxin Signaling to Modulate Hypocotyl Growth. Plant Physiol. https://doi.org/10.1104/pp.16.00405
Favero DS, Le KN, Neff MM (2017) Brassinosteroid signaling converges with SUPPRESSOR OF PHYTOCHROME B4-#3 to influence the expression of SMALL AUXIN UP RNA genes and hypocotyl growth. Plant J 89:1133–1145. https://doi.org/10.1111/tpj.13451
Fujimoto S, Matsunaga S, Yonemura M, Uchiyama S, Azuma T, Fukui K (2004) Identification of a novel plant MAR DNA binding protein localized on chromosomal surfaces. Plant Mol Biol 56:225–239. https://doi.org/10.1007/s11103-004-3249-5
Gendreau E, Traas J, Desnos T, Grandjean O, Caboche M, Höfte H (1997) Cellular basis of hypocotyl growth in Arabidopsis thaliana. Plant Physiol 114:295–305
Gnan S, Priest A, Kover PX (2014) The Genetic Basis of Natural Variation in Seed Size and Seed Number and Their Trade-Off Using Arabidopsis thaliana MAGIC Lines. Genetics 198:1751–1758. https://doi.org/10.1534/genetics.114.170746
Harker KN, O’Donovan JT, Smith EG, Johnson EN, Peng G, Willenborg CJ, Gulden RH, Moh R, Grenkow KS, Gill LA (2015) Seed size and seeding rate effects on canola emergence, development, yield and seed weight. Can J Plant Sci 95:1–8. https://doi.org/10.4141/cjps-2014-222
Harper JL, Lovell PH, Moore KG (1970) The Shapes and Sizes of Seeds. Annu Rev Ecol Syst 1:327–356. https://doi.org/10.1146/annurev.es.01.110170.001551
Heather DW, Sieczka JB (1991) Effect of seed size and cultivar on emergence and stand establishment of broccoli in crusted soil. J Am Soc Horticu Sci 116:946–949. https://doi.org/10.21273/JASHS.116.6.946
Jiang W-B, Huang H-Y, Hu Y-W, Zhu S-W, Wang Z-Y, Lin W-H (2013) Brassinosteroid Regulates Seed Size and Shape in Arabidopsis. Plant Physiol 162:1965–1977. https://doi.org/10.1104/pp.113.217703
Kagale S, Koh C, Nixon J, Bollina V, Clarke WE, Juteja R, Spillane C, Robinson SJ, Links MG, Clarke C, Higgins EE, Hubert T, Sharpe AG, Parkin IAP (2014) The emerging biofuel crop Camelina sativa retains a highly undifferentiated hexaploid genome structure. Nature Commun. https://doi.org/10.1038/ncomms4706
Kaydan D, Yagmur M (2008) Germination, seedling growth and relative water content of shoot in different seed sizes of triticale under osmotic stress of water and NaCl. Afr J Biotechnol 7:2862–2868
Khan N, Kazmi RH, Willems LAJ, van Heusden AW, Ligterink W, Hilhorst HWM (2012) Exploring the Natural Variation for Seedling Traits and Their Link with Seed Dimensions in Tomato. PLoS ONE 7:e43991. https://doi.org/10.1371/journal.pone.0043991
Lagercrantz U (1998) Comparative mapping between Arabidopsis thaliana and Brassica nigra indicates that Brassica genomes have evolved through extensive genome replication accompanied by chromosome fusions and frequent rearrangements. Genetics 150:1217–1228
Lagercrantz U, Lydiate DJ (1996) Comparative genome mapping in Brassica. Genetics 144:1903–1910
Li N, Li Y (2016) Signaling pathways of seed size control in plants. Curr Opin Plant Biol 33:23–32. https://doi.org/10.1016/j.pbi.2016.05.008
Lu C, Kang J (2008) Generation of transgenic plants of a potential oilseed crop Camelina sativa by Agrobacterium-mediated transformation. Plant Cell Rep 27:273–278. https://doi.org/10.1007/s00299-007-0454-0
Major DJ (1977) Influence of Seed Size on Yield and Yield Components of Rape1. Agron J 69:541. https://doi.org/10.2134/agronj1977.00021962006900040004x
Mendham NJ, Shipwa PA, Scott RK (1981) The effects of seed size, autumn nitrogen and plant population density on the response to delayed sowing in winter oil-seed rape (Brassica napus). J Agric Sci 96:417. https://doi.org/10.1017/S0021859600066181
Osborn TC, Kole C, Parkin IA, Sharpe AG, Kuiper M, Lydiate DJ, Trick M (1997) Comparison of flowering time genes in Brassica rapa, B. napus and Arabidopsis thaliana. Genetics 146:1123–1129
Pavlista AD, Isbell TA, Baltensperger DD, Hergert GW (2011) Planting date and development of spring-seeded irrigated canola, brown mustard and Camelina. Ind Crops Prod 33:451–456. https://doi.org/10.1016/j.indcrop.2010.10.029
Peng H, Zhao J, Neff MM (2015) ATAF2 integrates Arabidopsis brassinosteroid inactivation and seedling photomorphogenesis. Development 142:4129–4138. https://doi.org/10.1242/dev.124347
Raymond R, Nissen MS (1990) The A.T-DNA-binding domain of mammalian high mobility group I chromosomal proteins. A novel peptide motif for recognizing DNA structure. J Biol Chem 265:8573–658’2
Rensing SA, Lang D, Zimmer AD, Terry A, Salamov A, Shapiro H, Nishiyama T, Perroud PF, Lindquist EA, Kamisugi Y, Tanahashi T, Sakakibara K, Fujita T, Oishi K, Shin-l T, Kuroki Y, Toyoda A, Suzuki Y, Hashimoto S, Yamaguchi K, Sugano S, Kohora Y, Fujiyama A, Anterola A, Aoki S, Ashton N, Barbazuk WB, Barker E, Bennetzen JL, Blamkinship R, Cho SH, Dutcher SK, Estelle M, Fawcell JA, Gundlach H, Hanada K, Heyl A, Hicks KA, Hughes J, Lohr M, Mayer K, Melkozernov A, Murata T, Nelson DR, Pils B, Prigge M, Reiss B, Renner T, Rombauts S, Rushton PJ, Sanderfoot A, Schween G, Shiu SH, Stueber K, Theodoulou FL, Tu H, Van de Peer Y, Verrier PJ, Waters E, Wood A, Yang L, Cove D, Cuming AC, Husebe M, Lucas S, Mishler BD, Reski R, Grigoriev lV, Quatrano RS, Boore JL (2008) The Physcomitrella Genome Reveals Evolutionary Insights into the Conquest of Land by Plants. Science 319:64–69. https://doi.org/10.1126/science.1150646
Sadras VO (2007) Evolutionary aspects of the trade-off between seed size and number in crops. Field Crops Res. 100:125–138
Sadras and Egli (2008) Seed Size Variation in Grain Crops: allometric Relationships between Rate anDuration of Seed Growth. Crop Sci 48:1–8. https://doi.org/10.2135/cropsci2007.05.0292
Sadras VO, Slafer GA (2012) Environmental modulation of yield components in cereals: heritabilities reveal a hierarchy of phenotypic plasticities. Field Crops Research 127:215–224. https://doi.org/10.1016/j.fcr.2011.11.014
Schillinger WF, Young DL, Kennedy AC, Paulitz TC (2010) Diverse no-till irrigated crop rotations instead of burning and plowing continuous wheat. Field Crops Res 115(1):39–49. https://doi.org/10.1016/j.fcr.2009.10.001
Shonnard DR, Williams L, Kalnes TN (2010) Camelina-Derived Jet Fuel and Diesel: sustainable Advanced Biofuels. Environ Prog Sustain Energy 29(3):382–392. https://doi.org/10.1002/ep.10461
Street IH, Shah PK, Smith AM, Avery N, Neff MM (2008) The AT-hook-containing proteins SOB3/AHL29 and ESC/AHL27 are negative modulators of hypocotyl growth in Arabidopsis: sOB3/ESC: modulators of hypocotyl growth. Plant J 54:1–14. https://doi.org/10.1111/j.1365-313X.2007.03393.x
Sturrock CJ, Woodhall J, Brown M, Walker C, Mooney SJ, Ray RV (2015) Effects of damping-off caused by Rhizoctonia solani anastomosis group 2-1 on roots of wheat and oil seed rape quantified using X-ray Computed Tomography and real-time PCR. Front Plant Sci. https://doi.org/10.3389/fpls.2015.00461
Sun Y, Wang C, Wang N, Jiang X, Mao H, Zhu C, Wen F, Wang X, Lu Z, Yue G, Xu Z, Ye J (2017) Manipulation of Auxin Response Factor 19 affects seed size in the woody perennial Jatropha curcas. Sci Rep. https://doi.org/10.1038/srep40844
Tanabata et al (2012) SmartGrain: High-Throughput Phenotyping Software for Measuring Seed Shape through Image Analysis. PLANT PHYSIOLOGY 4, 1871-1880. https://doi.org/10.1104/pp.112.205120
Venable DL (1992) Size-number trade-offs and the variation of seed size with plant resource status. Am Nat 140:287–304
Zhao J, Favero DS, Peng H, Neff MM (2013) Arabidopsis thaliana AHL family modulates hypocotyl growth redundantly by interacting with each other via the PPC/DUF296 domain. Proc Natl Acad Sci 110:E4688–E4697. https://doi.org/10.1073/pnas.1219277110
Zhao J, Favero DS, Qiu J, Roalson EH, Neff MM (2014) Insights into the evolution and diversification of the AT-hook Motif Nuclear Localized gene family in land plants. BMC Plant Biol. https://doi.org/10.1186/s12870-014-0266-7
Zou J, Raman H, Guo S, Hu D, Wei Z, Luo Z, Long Y, Shi W, Fu Z, Du D, Meng J (2014) Constructing a densgenetic linkage map and mapping QTL for the traits of flower development in Brassica carinata. Theor Appl Genet 127:1593–1605. https://doi.org/10.1007/s00122-014-2321-z
Acknowledgements
This research project was originally supported by the Agriculture and Food Research Initiative competitive grant # 2013-67013-21666 of the USDA National Institute of Food and Agriculture (to M.M.N.). This research project was also supported by the Washington Oilseed Cropping Systems Competitive Grant Program (to M.M.N.), which is part of the Washington State Biofuels Initiative. This research project was also supported by the USDA National Institute of Food and Agriculture, Hatch umbrella project #1015621. The authors would like to thank Dr. Daniel Leroy Mullendore at the Franceschi Microscopy and Imaging Center, Washington State University for taking seedling images. We thank Dr. Ed Cahoon for donating the plant binary expression vector pBinGlyRed3 used in this study. We thank the following Neff lab members for their critical review and comments on this manuscript: Caitlin Jacques, Jessica Ortiz M.S., Hao Peng Ph.D., and Jonathan Schnore M.S..
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sharma Koirala, P., Neff, M.M. Improving seed size, seed weight and seedling emergence in Camelina sativa by overexpressing the Atsob3-6 gene variant. Transgenic Res 29, 409–418 (2020). https://doi.org/10.1007/s11248-020-00208-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11248-020-00208-9