Abstract
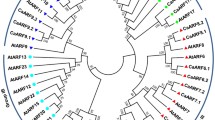
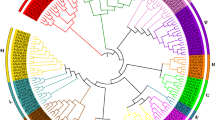
CUP-SHAPED COTYLEDON (CUC) transcription factors have a central regulatory function in plant growth and development. However, their involvement in kenaf (Hibiscus cannabinus L.) remains largely unexplored. In this study, we conducted a comprehensive analysis to identify six HcCUC genes in the kenaf genome. Through bioinformatic analysis, we found that the kenaf HcCUC genes share similar motifs and highly conserved gene structures. Phylogenetic analysis categorized the six HcCUC genes into two groups, that shared similarities with CUC2 or CUC3 genes from other species. Collinearity analysis revealed the formation of 6 syntenic gene pairs among the HcCUC genes, and 8 homologous gene pairs with three AtCUC genes from Arabidopsis. To investigate tissue-specific expression, we analyzed transcriptome data, that showed differential expression of HcCUC genes, particularly in leaves during the seedling stage, buds during the maturation stage, and anthers at the dual-core period. Functional characterization of HcCUC1 was achieved through its overexpression in Arabidopsis, resulting in elongated cotyledons, absent of petioles and increased number of rosette leaf and lateral branches. qRT-PCR analysis revealed that HcCUC1 potentially influences leaf and lateral branch development by up-regulating the expression of auxin-related genes (AtYUC2, AtYUC4, AtPIN1, AtPIN3, AtPIN4) and leaf shape-related genes (AtKNAT2, AtKNAT6). Notably, overexpression of HcCUC1 down-regulated the expression of flowering-related genes (AtFT, AtAP1, AtLFY, AtFUL), causing delayed flowering. Overall, our findings emphasize the pivotal role of HcCUC1 in regulating leaf and lateral branch growth, development, and flowering time, provide valuable insights into the function and genetic regulation of HcCUC genes.
Key message
Six HcCUC genes were identified in the kenaf genome, with HcCUC1 playing a role in regulating leaf and lateral branch growth and development, as well as flowering time.






Similar content being viewed by others
Data availability
The data generated and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Aida M, Ishida T, Fukaki H, Fujisawa H, Tasaka M (1997) Genes involved in organ separation in Arabidopsis: an analysis of the cup-shaped cotyledon mutant. Plant Cell 9(6):841–857. https://doi.org/10.1105/tpc.9.6.841
Aida M, Vernoux T, Furutani M, Traas J, Tasaka M (2002) Roles of PIN-FORMED1 and MONOPTEROS in pattern formation of the apical region of the Arabidopsis embryo. Development 129(17):3965–3974. https://doi.org/10.1242/dev.129.17.3965
Amasino R (2010) Seasonal and developmental timing of flowering. Plant J 61(6):1001–1013. https://doi.org/10.1111/j.1365-313X.2010.04148.x
Aslam M, She Z, Jakada BH, Fakher B, Greaves JG, Yan M, Chen Y, Zheng P, Cheng Y, Qin Y (2022) Interspecific complementation-restoration of phenotype in Arabidopsis cuc2cuc3 mutant by sugarcane CUC2 gene. BMC Plant Biol 22(1):47. https://doi.org/10.1186/s12870-022-03440-z
Bailey TL, Johnson J, Grant CE, Noble WS (2015) The MEME suite. Nucleic Acids Res 43(W1):W39–49. https://doi.org/10.1093/nar/gkv416
Baker CC, Sieber P, Wellmer F, Meyerowitz EM (2005) The early extra petals1 mutant uncovers a role for microRNA miR164c in regulating petal number in Arabidopsis. Curr Biol 15(4):303–315. https://doi.org/10.1016/j.cub.2005.02.017
Bilsborough GD, Runions A, Barkoulas M, Jenkins HW, Hasson A, Galinha C, Laufs P, Hay A, Prusinkiewicz P, Tsiantis M (2011) Model for the regulation of Arabidopsis thaliana leaf margin development. Proc Natl Acad Sci U S A 108(8):3424–3429. https://doi.org/10.1073/pnas.1015162108
Brewer PB, Dun EA, Gui R, Mason MG, Beveridge CA (2015) Strigolactone inhibition of branching Independent of Polar Auxin Transport. Plant Physiol 168(4):1820–1829. https://doi.org/10.1104/pp.15.00014
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, Xia R (2020) TBtools: an integrative Toolkit developed for interactive analyses of big Biological Data. Mol Plant 13(8):1194–1202. https://doi.org/10.1016/j.molp.2020.06.009
Chen P, Li Z, Luo D, Jia R, Lu H, Tang M, Hu Y, Yue J, Huang Z (2021) Comparative transcriptomic analysis reveals key genes and pathways in two different cadmium tolerance kenaf (Hibiscus cannabinus L.) cultivars. Chemosphere 263:128211. https://doi.org/10.1016/j.chemosphere.2020.128211
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16(6):735–743. https://doi.org/10.1046/j.1365-313x.1998.00343.x
Daimon Y, Takabe K, Tasaka M (2003) The CUP-SHAPED COTYLEDON genes promote adventitious shoot formation on calli. Plant Cell Physiol 44(2):113–121. https://doi.org/10.1093/pcp/pcg038
Danalatos NG, Archontoulis SV (2010) Growth and biomass productivity of kenaf (Hibiscus cannabinus, L.) under different agricultural inputs and management practices in central Greece. Ind Crops Prod 32(3):231–240. https://doi.org/10.1016/j.indcrop.2010.04.013
Fornara F, de Montaigu A, Coupland G (2010) SnapShot: control of flowering in Arabidopsis. Cell 141(3):550. https://doi.org/10.1016/j.cell.2010.04.024. 550 e1-2 doi
Gonzalez-Carranza ZH, Zhang X, Peters JL, Boltz V, Szecsi J, Bendahmane M, Roberts JA (2017) HAWAIIAN SKIRT controls size and floral organ number by modulating CUC1 and CUC2 expression. PLoS ONE 12(9):e0185106. https://doi.org/10.1371/journal.pone.0185106
Gordon SP, Heisler MG, Reddy GV, Ohno C, Das P, Meyerowitz EM (2007) Pattern formation during de novo assembly of the Arabidopsis shoot meristem. Development 134(19):3539–3548. https://doi.org/10.1242/dev.010298
Guo R, Xu X, Carole B, Li X, Gao M, Zheng Y, Wang X (2013) Genome-wide identification, evolutionary and expression analysis of the aspartic protease gene superfamily in grape. BMC Genomics 14:554. https://doi.org/10.1186/1471-2164-14-554
Hasson A, Plessis A, Blein T, Adroher B, Grigg S, Tsiantis M, Boudaoud A, Damerval C, Laufs P (2011) Evolution and diverse roles of the CUP-SHAPED COTYLEDON genes in Arabidopsis leaf development. Plant Cell 23(1):54–68. https://doi.org/10.1105/tpc.110.081448
Hibara K, Takada S, Tasaka M (2003) CUC1 gene activates the expression of SAM-related genes to induce adventitious shoot formation. Plant J 36(5):687–696. https://doi.org/10.1046/j.1365-313X.2003.01911.x
Hibara K, Karim MR, Takada S, Taoka K, Furutani M, Aida M, Tasaka M (2006) Arabidopsis CUP-SHAPED COTYLEDON3 regulates postembryonic shoot meristem and organ boundary formation. Plant Cell 18(11):2946–2957. https://doi.org/10.1105/tpc.106.045716
Kareem A, Durgaprasad K, Sugimoto K, Du Y, Pulianmackal AJ, Trivedi ZB, Abhayadev PV, Pinon V, Meyerowitz EM, Scheres B, Prasad K (2015) PLETHORA Genes Control Regeneration by a Two-Step Mechanism. Curr Biol 25(8):1017-30 doi:https://doi.org/10.1016/j.cub.2015.02.022
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol Biol Evol 35(6):1547–1549. https://doi.org/10.1093/molbev/msy096
Laufs P, Peaucelle A, Morin H, Traas J (2004) MicroRNA regulation of the CUC genes is required for boundary size control in Arabidopsis meristems. Development 131(17):4311–4322. https://doi.org/10.1242/dev.01320
Li Y, Xia T, Gao F, Li Y (2020) Control of plant branching by the CUC2/CUC3-DA1-UBP15 Regulatory Module. Plant Cell 32(6):1919–1932. https://doi.org/10.1105/tpc.20.00012
Liu CY, Xu HW, Jiang J, Wang S, Liu GF (2018) Analysis of the promoter features of BpCUC2 in Betula platyphylla x Betula pendula. Plant Cell Tiss Org 132(1):191–199. https://doi.org/10.1007/s11240-017-1324-2
Liu C, Xu H, Han R, Wang S, Liu G, Chen S, Chen J, Bian X, Jiang J (2019) Overexpression of BpCUC2 influences Leaf shape and Internode Development in Betula pendula. Int J Mol Sci 20(19). https://doi.org/10.3390/ijms20194722
Mallory AC, Dugas DV, Bartel DP, Bartel B (2004) MicroRNA regulation of NAC-domain targets is required for proper formation and separation of adjacent embryonic, vegetative, and floral organs. Curr Biol 14(12):1035–1046. https://doi.org/10.1016/j.cub.2004.06.022
Muller D, Leyser O (2011) Auxin, cytokinin and the control of shoot branching. Ann Bot 107(7):1203–1212. https://doi.org/10.1093/aob/mcr069
Nikovics K, Blein T, Peaucelle A, Ishida T, Morin H, Aida M, Laufs P (2006) The balance between the MIR164A and CUC2 genes controls leaf margin serration in Arabidopsis. Plant Cell 18(11):2929–2945. https://doi.org/10.1105/tpc.106.045617
Ramesh M (2016) Kenaf (Hibiscus cannabinus L.) fibre based bio-materials: a review on processing and properties. Prog Mater Sci 78–79:1–92. https://doi.org/10.1016/j.pmatsci.2015.11.001
Su W, Ren Y, Wang D, Huang L, Fu X, Ling H, Su Y, Huang N, Tang H, Xu L, Que Y (2020) New insights into the evolution and functional divergence of the CIPK gene family in Saccharum. BMC Genomics 21(1):868. https://doi.org/10.1186/s12864-020-07264-9
Taoka K, Yanagimoto Y, Daimon Y, Hibara K, Aida M, Tasaka M (2004) The NAC domain mediates functional specificity of CUP-SHAPED COTYLEDON proteins. Plant J 40(4):462–473. https://doi.org/10.1111/j.1365-313X.2004.02238.x
Wang Y, Tang H, Debarry JD, Tan X, Li J, Wang X, Lee TH, Jin H, Marler B, Guo H, Kissinger JC, Paterson AH (2012) MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res 40(7):e49. https://doi.org/10.1093/nar/gkr1293
Wang J, Bao J, Zhou B, Li M, Li X, Jin J (2021) The osa-miR164 target OsCUC1 functions redundantly with OsCUC3 in controlling rice meristem/organ boundary specification. New Phytol 229(3):1566–1581. https://doi.org/10.1111/nph.16939
Wei F, Tang D, Li Z, Kashif MH, Khan A, Lu H, Jia R, Chen P (2019) Molecular cloning and subcellular localization of six HDACs and their roles in response to salt and drought stress in kenaf (Hibiscus cannabinus L). Biol Res 52(1):20. https://doi.org/10.1186/s40659-019-0227-6
Wen S, Li J, Hao Z, Wei L, Ma J, Zong Y, Li H (2022) Overexpression of the LcCUC2-like gene in Arabidopsis thaliana alters the cotyledon morphology and increases rosette leaf number. PeerJ 10:e12615. https://doi.org/10.7717/peerj.12615
Xiong Y, Jiao Y (2019) The diverse roles of Auxin in regulating Leaf Development. Plants (Basel) 8(7). https://doi.org/10.3390/plants8070243
Yu CS, Lin CJ, Hwang JK (2004) Predicting subcellular localization of proteins for Gram-negative bacteria by support vector machines based on n-peptide compositions. Protein Sci 13(5):1402–1406. https://doi.org/10.1110/ps.03479604
Yue J, Tang M, Zhang H, Luo D, Cao S, Hu Y, Huang Z, Wu Q, Wu X, Pan J, Chen C, Wang C, Chen P (2022) The transcription factor HcERF4 confers salt and drought tolerance in kenaf (Hibiscus cannabinus L). Plant Cell Tissue and Organ Culture (PCTOC) 150(1):207–221. https://doi.org/10.1007/s11240-022-02260-1
Zhang LW, Xu Y, Zhang XT, Ma XK, Zhang LL, Liao ZY, Zhang Q, Wan XB, Cheng Y, Zhang JS, Li DX, Zhang LM, Xu JT, Tao AF, Lin LH, Fang PP, Chen S, Qi R, Xu XM, Qi JM, Ming R (2020) The genome of kenaf (Hibiscus cannabinus L.) provides insights into bast fibre and leaf shape biogenesis. Plant Biotechnol J 18(8):1796–1809. https://doi.org/10.1111/pbi.13341
Zheng G, Wei W, Li Y, Kan L, Wang F, Zhang X, Li F, Liu Z, Kang C (2019) Conserved and novel roles of miR164-CUC2 regulatory module in specifying leaf and floral organ morphology in strawberry. New Phytol 224(1):480–492. https://doi.org/10.1111/nph.15982
Acknowledgements
This research work was supported by the National Natural Science Foundation of China (Grant No. 31960368).
Author information
Authors and Affiliations
Contributions
QW: Data curation; Methodology, Formal analysis, Roles/Writing - original draft. CC; JY: Investigation, Formal analysis. SM: Writing-review & editing.SC; XL: Investigation, Data curation, Formal analysis. MW; HZ: Software, Methodology, Formal analysis. XW; CW; DL: Formal analysis, Validation. PC: Conceptualization, Methodology, Writing-review & editing, Funding acquisition, Project administration.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by Manoj Prasad.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, Q., Chen, C., Yue, J. et al. Genome-wide identification of CUC gene family and functional analysis of HcCUC1 in kenaf. Plant Cell Tiss Organ Cult 155, 91–102 (2023). https://doi.org/10.1007/s11240-023-02555-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-023-02555-x