Abstract
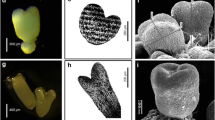
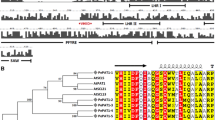
Carica papaya L. plantlets, normally exhibit low rooting capacity when cultured in vitro. It has been suggested in other species that auxin concentration at root tissues, is the result of a reflux system driven by auxin influx transporters (AIT; AUX1/LAX) and auxin efflux transporters (AET; PIN), that regulates the mechanism of initiation and development of lateral roots. Therefore, in the present paper, we studied the structure, phylogeny and the expression patterns of the whole family of AIT and AET in C. papaya, and their possible relation with the limited capacity to generate adventitious roots of in vitro cultured papaya plantlets. We found 4 AUX1/LAX genes (CpAUX1, CpLAX1, CpLAX2, CpLAX3) and 6 PIN genes (CpPIN1, CpPIN2, CpPIN3, CpPIN4, CpPIN5, CpPIN6) within the genome of C. papaya. The expression patterns and levels of those genes were studied in stem-base and root tissues from C. papaya cv. Maradol plants under four different treatments: (1) in vitro plantlets without IBA (that did not generate roots), (2) in vitro plantlets treated with 2 mg L−1 IBA (that did generate roots), (3) de-rooted seedlings treated with the same concentration of IBA (that also generated adventitious roots), and (4) intact seedlings used as controls. Histological studies made on the stem base and root tissues from all treatments showed that the IBA-induced roots were histologically equivalent, to those naturally formed in intact seedlings. In vitro plantlets non-treated with IBA had low expression of all auxin transporters genes in stem-base tissues and they were unable to produce roots. On the contrary, in vitro plantlets treated with IBA experienced a marked increase in the expression of most auxin transporters genes, in particular of CpLAX3 and CpPIN2, and they were capable to produce roots. Those roots generated in the IBA-treated in vitro plantlets, showed expression levels and patterns of auxin transporter genes, equivalent to those shown in both, the IBA-treated de-rooted seedlings, and in the naturally formed roots from intact seedlings.








Similar content being viewed by others
Abbreviations
- AET:
-
Auxin efflux transporters
- AIT:
-
Auxin influx transporters
- AUX1/LAX:
-
AIT genes
- IAA:
-
Indole-acetic acid
- IBA:
-
Indole-3-butyric acid
- L:
-
Leaf
- LD:
-
Long-distance
- NAA:
-
1-Naphthaleneacetic acid
- PAT:
-
Polar auxin transport
- PIN:
-
AET genes
- PPFD:
-
Photosynthetic photon flux density
- REL:
-
Relative expression levels
- RH:
-
Relative humidity
- R:
-
Root
- SB:
-
Stem-base
- SD:
-
Short-distance
- T:
-
Temperature
- 2,4-D:
-
2,4-Dichlorophenoxyacetic acid
References
Abas L, Benjamins R, Malenica N, Paciorek T, Wišniewska J, Moulinier-Anzola JC, Sieberer T, Friml J, Luschnig C (2006) Intracellular trafficking and proteolysis of the auxin-efflux facilitator PIN2 are involved in root gravitropism. Nat Cell Biol 8:249–256. doi:10.1038/ncb1369
Adamowski M, Friml J (2015) PIN-dependent auxin transport: action, regulation, and evolution. Plant Cell 27:20–32. doi:10.1105/tpc.114.134874
Bainbridge K, Guyomarc’h S, Bayer E, Swarup R, Bennett M, Mandel T, Kuhlemeier C (2008) Auxin influx carriers stabilize phyllotactic patterning. Genes Dev 22:810–823. doi:10.1101/gad.462608
Ballester A, San-José MC, Vidal N, Fernández-Lorenzo JL, Vieitez AM (1999) Anatomical and biochemical events during in vitro rooting of microcuttings from juvenile and mature phases of chestnut. Ann Bot 83:619–629. doi:10.1006/anbo.1999.0865
Benková E, Michniewicz M, Sauer M, Teichmann T, Seifertová D, Jürgens G, Friml J (2003) Local, efflux-dependent auxin gradients as a common module for plant organ formation. Cell 115:591–602. doi:10.1016/S0092-8674(03)00924-3
Berlin GP, Miksche JP (1976) Botanical microtechnique and cytochemistry, 3rd edn. Iowa State University Press, Ames, p 254
Blilou I, Xu J, Wildwater M, Willemsen V, Paponov I, Friml J, Heidstra R, Mitsuhiro A, Palme K, Scheres B (2005) The PIN auxin efflux facilitator network controls growth and patterning in Arabidopsis roots. Nature 433:39–44. doi:10.1038/nature03184
Chen Y, Hao X, Cao J (2014) Small auxin upregulated RNA (SAUR) gene family in maize: identification, evolution, and its phylogenetic comparison with Arabidopsis, rice, and sorghum. J Integr Plant Biol 56:133–150. doi:10.1111/jipb.12127
Dall Bosco C, Dovzhenko A, Palme K (2012) Intracellular auxin transport in pollen. Plant Signal Behav 7:1504–1505. doi:10.4161/psb.21953
Dal Cin V, Barbaro E, Danesin M, Murayama H, Velasco R, Ramina A (2009) Fruitlet abscission: a cDNA-AFLP approach to study genes differentially expressed during shedding of immature fruits reveals the involvement of a putative auxin hydrogen symporter in apple (Malus domestica L. Borkh). Gene 442:26–36. doi:10.1016/j.gene.2009.04.009
De Smet I, Tetsumura T, De Rybel B, Frey NF, Laplaze L, Casimiro I, Swarup R, Naudts M, Vanneste S, Audenaert D, Inzé D, Bennett MJ, Beeckman T (2007) Auxin-dependent regulation of lateral root positioning in the basal meristem of Arabidopsis. Development 134:681–690. doi:10.1242/dev.02753
Ding Z, Wang B, Moreno I, Dupláková N, Simon S, Carraro N, Reemmer J, Pěnčík A, Chen X, Tejos R, Skůpa P, Pollmann S, Mravec J, Petrášek J, Zažímalova E, Honys D, Rolčík J, Murphy A, Orellana A, Geisler M, Friml J (2012) ER-localized auxin transporter PIN8 regulates auxin homeostasis and male gametophyte development in Arabidopsis. Nat Commun 3:941–947. doi:10.1038/ncomms1941
Drew RA (1987) The effects of medium composition and cultural conditions on in vitro root initiation and growth of papaya (Carica papaya L.). J Hortic Sci Biotechnol 62:551–556. http://www.jhortscib.org/Vol62/62_4/20.htm. Accessed 6 Jun 2015
Drew RA, Miller RM (1993) Nutritional and cultural factors affecting rooting of papaya (Carica papaya L.) in vitro. Hortic Sci 64:767–773. doi:10.1080/14620316.1989.11516019
Drew RA, McComb JA, Considine JA (1993) Rhizogenesis and root growth of Carica papaya L. in vitro in relation to auxin sensitive phase and use of riboflavin. Plant Cell Tissue Organ Cult 33:1–7. doi:10.1007/BF01997591
Dubrovsky JG, Gambetta GA, Hernandez-Barrera A, Shishkova S, Gonzalez I (2006) Lateral root initiation in Arabidopsis: developmental window, spatial patterning, density and predictability. Ann Bot 97:903–915. doi:10.1093/aob/mcj604
Dubrovsky JG, Sauer M, Napsucialy-Mendivil S, Ivanchenko MG, Friml J, Shishkova S, Celenza J, Benková E (2008) Auxin acts as a local morphogenetic trigger to specify lateral root founder cells. Proc Natl Acad Sci 105:8790–8794. doi:10.1073/pnas.0712307105
Fang H, Zago MK, Abas L, Van Marion A, Galván-Ampudia CS, Offringa R (2010) Phosphorylation of conserved PIN motifs directs Arabidopsis PIN1 polarity and auxin transport. Plant Cell 22:1129–1142. doi:10.1105/tpc.109.072678
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791. doi:10.1016/j.bse.2010.10.005
Forestan C, Farinati S, Varotto S (2012) The maize PIN gene family of auxin transporters. Front Plant Sci 3:1–16. doi:10.3389/fpls.2012.00016
Friml J, Palme K (2002) Polar auxin transport—old questions and new concepts? Plant Mol Biol 49:273–284. doi:10.1023/A:1015248926412
Friml J, Yang X, Michniewicz M, Weijers D, Quint A, Tietz O, Benjamins R, Ouwerkerk P, Ljung K, Sandberg G, Hooykaas P, Palme K, Offringa R (2004) A PINOID-dependent binary switch in apical-basal PIN polar targeting directs auxin efflux. Science 306:862–865. doi:10.1126/science.1100618
Gälweiler L, Guan C, Müller A, Wisman E, Mendgen K, Yephremov A, Palme K (1998) Regulation of polar auxin transport by AtPIN1 in Arabidopsis vascular tissue. Science 282:2226–2230. doi:10.1126/science.282.5397.2226
Ganguly A, Park M, Kesawat MS, Cho HT (2014) Functional analysis of the hydrophilic loop in intracellular trafficking of Arabidopsis PIN-FORMED proteins. Plant Cell 26:1570–1585. doi:10.3410/f.718335431.793494308
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. In: Nucleic acids symposium series No. 41. Oxford University Press. pp. 95–98. http://brownlab.mbio.ncsu.edu/JWB/papers/1999Hall1.pdf
Himanen K, Boucheron E, Vanneste S, de Almeida Engler J, Inzé D, Beeckman T (2002) Auxin-mediated cell cycle activation during early lateral root initiation. Plant Cell 14:2339–2351. doi:10.1105/tpc.004960
Hoffmann M, Hentrich M, Pollmann S (2011) Auxin–oxylipin crosstalk: relationship of antagonists. J Integr Plant Biol 53:429–445. doi:10.1111/j.1744-7909.2011.01053.x
Idrovo EFM, Peraza-Echeverria S, Fuentes G, Santamaría JM (2012) In silico cloning and characterization of the TGA (TGACG MOTIF-BINDING FACTOR) transcription factors subfamily in Carica papaya. Plant Physiol Biochem 54:113–122. doi:10.1016/j.plaphy.2012.02.011
Iliev I, Kitin P, Funada R (2001) Morphological and anatomical study on in vitro root formation of silver birch (Betula pendula Roth.). Prop Orn Plants 1:10–19
Islam R, Rahman SM, Hossain M, Joarder OI (1993) In vitro clonal propagation of papaya (Carica papaya L.). Pak J Bot 25:189–192. http://www.pakbs.org/pjbot/PDFs/25(2)/PJB25(2)13.pdf
Ivanchenko MG, Zhu J, Wang B, Medvecká E, Du Y, Azzarello E, Mancuso S, Megraw M, Filichkin S, Dubrovsky JG, Friml J, Geisler M (2015) The cyclophilin A DIAGEOTROPICA gene affects auxin transport in both root and shoot to control lateral root formation. Development 142:712–721. doi:10.1242/dev.113225
Jiménez VM, Mora-Newcomer E, Gutiérrez-Soto MV (2014) Biology of the papaya plant. In: Ming R, Moore PH (eds) Genetics and genomics of papaya, plant genetics and genomics: crops and models 10. Springer Science + Business Media, New York, pp 17–33
Jones DT, Taylor WR, Thornton JM (1992) The rapid generation of mutation data matrices from protein sequences. Comput Appl Biosci 8:275–282. doi:10.1093/bioinformatics/8.3
Kerr ID, Bennett MJ (2007) New insight into the biochemical mechanisms regulating auxin transport in plants. Biochem J 401:613–622. doi:10.1042/BJ20061411
Kramer EM, Bennett MJ (2006) Auxin transport: a field in flux. Trends Plant Sci 11:382–386. doi:10.1016/j.tplants.2006.06.002
Křeček P, Skůpa P, Libus J, Naramoto S, Tejos R, Friml J, Zažímalová E (2009) The PIN-FORMED (PIN) protein family of auxin transporters. Genome Biol 10:1–11. doi:10.1186/gb-2009-10-12-249
Laskowski M, Grieneisen VA, Hofhuis H, Ten Hove CA, Hogeweg P, Marée AF, Scheres B (2008) Root system architecture from coupling cell shape to auxin transport. PLoS Biol 6:e307. doi:10.1371/journal.pbio.0060307
Lavenus J, Goh T, Roberts I, Guyomarc’h S, Lucas M, De Smet I, Fukaki H, Beeckman T, Bennett M, Laplaze L (2013) Lateral root development in Arabidopsis: fifty shades of auxin. Trends Plant Sci 18:450–458. doi:10.1016/j.tplants.2013.04.006
Magdalita PM, Persley DM, Godwin ID, Drew RA, Adkins SW (1997) Screening Carica papaya × C. cauliflora hybrids for resistance to papaya ringspot virus-type P. Plant Pathol 46:837–841. doi:10.1046/j.1365-3059.1997.d01-90.x
Malabadi RB, Vijayakumar S, Mulgund GS, Nataraja K (2011) Induction of somatic embryogenesis in papaya (Carica papaya L). Res Biotechnol 2:40–55
Marchant A, Bhalerao R, Casimiro I, Eklöf J, Casero PJ, Bennett M, Sandberg G (2002) AUX1 promotes lateral root formation by facilitating indole-3-acetic acid distribution between sink and source tissues in the Arabidopsis seedling. Plant Cell 14:589–597. doi:10.1105/tpc.010354
McManus JFA (1961) Periodate oxidation techniques. General cytochemical methods. Academic Press, New York, pp 171–201
Medina R, Faloci MM, Gonzalez AM, Mroginski LA (2007) In vitro cultured primary roots derived from stem segments of cassava (Manihot esculenta) can behave like storage organs. Ann Bot 99:409–423. doi:10.1093/aob/mcl272
Michniewicz M, Zago MK, Abas L, Weijers D, Schweighofer A, Meskiene I, Heisler MG, Ohno C, Zhang J, Huang F, Schwab R, Weigel D, Meyerowitz EM, Luschnig C, Offringa R, Friml J (2007) Antagonistic regulation of PIN phosphorylation by PP2A and PINOID directs auxin flux. Cell 130:1044–1056. doi:10.1016/j.cell.2007.07.033
Ming R, Hou S, Feng Y, Yu Q, Dionne-Laporte A, Saw JH, Senin P et al (2008) The draft genome of the transgenic tropical fruit tree papaya (Carica papaya Linnaeus). Nature 452:991–996. doi:10.1038/nature06856
Moreno-Risueno MA, Van Norman JM, Moreno A, Zhang J, Ahnert SE, Benfey PN (2010) Oscillating gene expression determines competence for periodic Arabidopsis root branching. Science 329:1306–1311. doi:10.1126/science.1191937
Mravec J, Skůpa P, Bailly A, Hoyerová K, Křeček P, Bielach A, Petrášek J, Zhang J, Gaykova V, Stierhof YD, Dobrev PI, Schwarzerová K, Rolčík J, Seifertová D, Luschnig C, Benková E, Zažímalová E, Geisler M, Friml J (2009) Subcellular homeostasis of phytohormone auxin is mediated by the ER-localized PIN5 transporter. Nature 459:1136–1140. doi:10.1038/nature08066
Muday GK, DeLong A (2001) Polar auxin transport: controlling where and how much. Trends Plant Sci 6:535–542. doi:10.1016/S1360-1385(01)02101-X
Müller A, Guan C, Gälweiler L, Tänzler P, Huijser P, Marchant A, Parry G, Bennett M, Wisman E, Palme K (1998) AtPIN2 defines a locus of Arabidopsis for root gravitropism control. EMBO J 17:6903–6911. doi:10.1093/emboj/17.23.6903
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497. doi:10.1111/j.1399-3054.1962.tb08052.x
Paponov IA, Teale WD, Trebar M, Blilou I, Palme K (2005) The PIN auxin efflux facilitators: evolutionary and functional perspectives. Trends Plant Sci 10:170–177. doi:10.1016/j.tplants.2005.02.009
Parry G, Delbarre A, Marchant A, Swarup R, Napier R, Perrot-Rechenmann C, Bennett MJ (2001a) Novel auxin transport inhibitors phenocopy the auxin influx carrier mutation aux1. Plant J 25:399–406. doi:10.1046/j.1365-313x.2001.00970.x
Parry G, Marchant A, May S, Swarup R, Swarup K, James N, Graham N, Allen T, Martucci T, Yemm A, Napier R, Manning K, King G, Bennett M (2001b) Quick on the uptake: characterization of a family of plant auxin influx carriers. J Plant Growth Regul 20:217–225. doi:10.1007/s003440010030
Peraza-Echeverría S, Santamaria JM, Gabriela Fuentes, Mariana Menéndez, Angel Vallejo M, Virginia Herrera (2012) The NPR1 family of transcription cofactors in papaya: insights into its structure, phylogeny and expression. Genes Genom 34:379–390. doi:10.1007/s13258-011-0218-7
Petrásëk J, Mravec J, Bouchard R, Blakeslee JJ, Abas M, Seifertová D, Wisniewska J, Tadele Z, Kubeš M, Čovanová M, Dhonukshe P, Skůpa P, Benková E, Perry L, Křeček P, Lee OR, Fink GR, Geisler M, Murphy AS, Luschnig C, Zažímalová E, Friml J (2006) PIN proteins perform a rate-limiting function in cellular auxin efflux. Science 312:914–918. doi:10.1126/science.1123542
Quint M, Gray WM (2006) Auxin signaling. Curr Opin Plant Biol 9:448–453. doi:10.1016/j.pbi.2006.07.006
Raven JA (1975) Transport of indoleacetic acid in plant cells in relation to pH and electrical potential gradients, and its significance for polar IAA transport. New Phytol 74:163–172. doi:10.1111/j.1469-8137.1975.tb02602.x
Reinhardt D, Pesce ER, Stieger P, Mandel T, Baltensperger K, Bennett M, Traas J, Friml J, Kuhlemeier C (2003) Regulation of phyllotaxis by polar auxin transport. Nature 426:255–260. doi:10.1038/nature02081
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sawchuk MG, Scarpella E (2013) Control of vein patterning by intracellular auxin transport. Plant Signal Behav 8:e27205. doi:10.4161/psb.27205
Schnabel EL, Frugoli J (2004) The PIN and LAX families of auxin transport genes in Medicago truncatula. Mol Genet Genom 272:420–432. doi:10.1007/s00438-004-1057-x
Shen C, Bai YH, Wang S, Zhang S, Wu YR, Chen M, Jiang DA, Qi YH (2010) Expression profile of PIN, AUX/LAX and PGP auxin transporters gene families in Sorghum bicolor under phytohormone and abiotic stress. FEBS J 277:2954–2969. doi:10.1111/j.1742-4658.2010.07706.x
Shen C, Yue R, Youhuang B, Feng R, Sun T, Wang Y, Tie S, Wang H (2015) Identification and analysis of Medicago truncatula auxin transporter gene families uncover their roles in responses to Sinorhizobium meliloti infection. Plant Cell Physiol 56:1930–1943. doi:10.1093/pcp/pcv113
Swarup R, Kramer EM, Perry P, Knox K, Ottoline LH, Haseloff J, Beemster G, Bhalerao R, Bennett MJ (2005) Root gravitropism requires lateral root cap and epidermal cells for transport and response to a mobile auxin signal. Nat Cell Biol 7:1057–1065. doi:10.1038/ncb1316
Swarup K, Benková E, Swarup R, Casimiro I, Péret B, Yang Y, Parry G, Nielsen E, De Smet I, Vanneste S, Levesque MP, Carrier D, James N, Calvo V, Ljung K, Kramer E, Roberts R, Graham N, Marillonnet S, Patel K, Jones J, Taylor C, Schachtman D, May S, Sandberg G, Benfey P, Friml J, Kerr I, Beeckman T, Laplaze L, Bennett MJ (2008) The auxin influx carrier LAX3 promotes lateral root emergence. Nat Cell Biol 10:946–954. doi:10.1038/ncb1754
Talavera C, Espadas F, Contreras F, Fuentes G, Santamaría JM (2007) Acclimatization, rooting and field establishment of micropropagated papaya plants. Acta Hortic 812:373–378. doi:10.17660/ActaHortic.2009.812.52
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739. doi:10.1093/molbev/msr121
Teale WD, Ditengou FA, Dovzhenko AD, Li X, Molendijk AM, Ruperti B, Paponov I, Palme K (2008) Auxin as a model for the integration of hormonal signal processing and transduction. Mol Plant 1:229–237. doi:10.1093/mp/ssn006
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882. doi:10.1093/nar/25.24.4876
Titapiwatanakun B, Murphy AS (2009) Post-transcriptional regulation of auxin transport proteins: cellular trafficking, protein phosphorylation, protein maturation, ubiquitination, and membrane composition. J Exp Bot 60:1093–1107. doi:10.1093/jxb/ern240
Ugartechea-Chirino Y, Swarup R, Swarup K, Péret B, Whitworth M, Bennett M, Bougourd S (2010) The AUX1 LAX family of auxin influx carriers is required for establishment of embryonic root cell organization in Arabidopsis thaliana. Ann Bot 105:277–289. doi:10.1093/aob/mcp287
Vandenbussche F, Petrášek J, Žádníková P, Hoyorevá K, Pešek B, Raz V, Swarup R, Bennett M, Zažímalová E, Benková E, Van Der Straeten D (2010) The auxin influx carriers AUX1 and LAX3 are involved in auxin-ethylene interactions during apical hook development in Arabidopsis thaliana seedlings. Development 137:597–606. doi:10.1242/dev.040790
Vanneste S, Friml J (2009) Auxin: a trigger for change in plant development. Cell 136:1005–1016. doi:10.1016/j.cell.2009.03.001
Visser EJW, Heijink CJ, Van Hout KJGM, Voesenek LACJ, Barendse GWM, Blom CWPM (1995) Regulatory role of auxin in adventitious root formation in two species of Rumex, differing in their sensitivity to waterlogging. Physiol Plant 93:116–122. doi:10.1034/j.1399-3054.1995.930117.x
Wabnik K, Kleine-Vehn J, Govaerts W, Friml J (2011) Prototype cell-to-cell auxin transport mechanism by intracellular auxin compartmentalization. Trends Plant Sci 16:468–475. doi:10.1016/jplants.2011.05.002
Wang JR, Hu H, Wang GH, Li J, Chen JY, Wu P (2009) Expression of PIN genes in rice (Oryza sativa L.): tissue specificity and regulation by hormones. Mol Plant 2:823–831. doi:10.1093/mp/ssp023
Whelan S, Goldman N (2001) A general empirical model of protein evolution derived from multiple protein families using a maximum-likelihood approach. Mol Biol Evol 18:691–699. doi:10.1093/oxfordjournals.molbev.a003851
Wu G, Lewis DR, Spalding EP (2007) Mutations in Arabidopsis multidrug resistance-like ABC transporters separate the roles of acropetal and basipetal auxin transport in lateral root development. Plant Cell 19:1826–1837. doi:10.1105/tpc.106.048777
Xu Y, Zhang S, Guo H, Wang S, Xu L, Li C et al (2014) OsABCB14 functions in auxin transport and iron homeostasis in rice (Oryza sativa L.). Plant J 79:106–117. doi:10.1111/tpj.12544
Yu TA, Yeh SD, Cheng YH, Yang JS (2000) Efficient rooting for establishment of papaya plantlets by micropropagation. Plant Cell Tissue Organ Cult 61:29–35. doi:10.1023/A:10064-759-0143-9
Yue R, Tie S, Sun T, Zhang L, Yang Y, Qi J, Yan S, Han X, Wang H, Shen C (2015) Genome-wide identification and expression profiling analysis of ZmPIN, ZmPILS, ZmLAX and ZmABCB auxin transporter gene families in maize (Zea mays L.) under various abiotic stresses. PLoS ONE 10:1–23. doi:10.1371/journal.pone.0118751
Zažímalová E, Křeček P, Skůpa P, Hoyerová K, Petrášek J (2007) Polar transport of the plant hormone auxin—the role of PIN-FORMED (PIN) proteins. Cell Mol Life Sci 64:1621–1637. doi:10.1007/s00018-007-6566-4
Zažímalová E, Murphy AS, Yang H, Hoyerová K, Hošek P (2010) Auxin transporters—why so many? Cold Spring Harb Perspect Biol 2:1–14. doi:10.1101/cshperspect.a001552
Acknowledgments
This work was funded by CONACYT, México (Project No. CB155356). H.E.M. acknowledges a scholarship (254647) granted by CONACYT.
Author contribution
E.M.H., First author, PhD student, performed the gene expression studies and the bioinformatic analysis; E.M.H., F.O.G., C.L.A., I.E.F. and R.Z.L. contributed to the writing of the paper; F.O.G. and I.E.F., supervision on the expression and bioinformatics analysis; C.L.A., Assisted in the gene expression studies; C.T.M. and E.G.F., Assisted in the tissue culture work, including the experimental set up for root induction; B.P.F., Responsible of the anatomical studies; S.J.M., Corresponding author. General conception of the project. Design of the experimental strategy, responsible for the writing of the paper.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest and the presented work is compliant with ethical standards of PCTOC. All the authors read and approved the manuscript in its final form.
Rights and permissions
About this article
Cite this article
Estrella-Maldonado, H., Fuentes Ortíz, G., Chan León, A.C. et al. The papaya CpAUX1/LAX and CpPIN genes: structure, phylogeny and expression analysis related to root formation on in vitro plantlets. Plant Cell Tiss Organ Cult 126, 187–204 (2016). https://doi.org/10.1007/s11240-016-0989-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-016-0989-2