Abstract
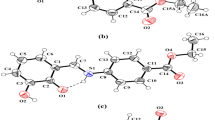
The in situ condensation reaction of 2-hydrazinobenzothiazole with salicylaldehyde, 3,4-dihydroxybenzaldehyde, 2,4-dihydroxybenzaldehyde, 2,5-dihydroxybenzaldehyde, 2,3-dihydroxybenzaldehyde, 2-hydroxy-1-naphthaldehyde, 2-methoxy-1-naphthaldehyde, 4-methoxy-1-naphthaldehyde and 6-methoxy-2-naphthaldehyde produced 9 hydrazone Schiff bases (L1–L9, respectively) which were identified and characterized by elemental analysis, IR and NMR spectroscopy. The crystal and molecular structures of four Schiff bases (L1, L7–L9) have been determined by the single-crystal X-ray diffraction method confirming the imino form of L1 and the amino tautomeric form of L7–L9 compounds. Molecular structure analysis also confirmed that reported compounds are E-isomers relative to exo C = N imino bond. The Nhydrazino–H group of amino tautomers forms Nhydrazino–H···Nthiazolyl intermolecular hydrogen bonds shaping molecules into R 22 (8) rings, while imino tautomer of L1 forms C(4) infinite helical chains via Nthiazolyl–H···Nhydrazino type of intermolecular hydrogen bond. The methoxy group (L7–L9) further shaped these primary supramolecular synthons into different supramolecular arrangements via C–H···O, C–H···N and C–H···S intermolecular hydrogen bonds. The role of aryl substituents in the shaping and stabilization of supramolecular architectures of L1, L7–L9 is supported by quantum chemical calculations. Strong antiproliferative effects on tumor cells and cytotoxic effects on fibroblasts are shown for all ligands L1–L9 with exception of L6 and L7 that had no effect on fibroblast cells.










Similar content being viewed by others
References
Mortimer CG, Wells G, Crochard JP, Stone EL, Bradshaw TD, Stevens MFG, Westwell AD (2006) J Med Chem 49:179–185
Al-Tel TH, Al-Qawasmeh RA, Zaarour R (2011) Eur J Med Chem 46:1874–1881
Ćaleta I, Kralj M, Marjanović M, Bertosa B, Tomić S, Pavlović G, Pavelić K, Karminski-Zamola G (2009) J Med Chem 52:1744–1756
Pattan SR, Suresh C, Pujar VD, Reddy VVK, Rasal VP, Koti BC (2005) Indian J Chem 44:2404–2408
Siddiqui N, Pandeya SN, Khau SA, Stables J, Rana A, Alam M, Arshad MF, Bhat MA (2007) Bioorg Med Chem Lett 17:255–259
Gurupadayya BM, Gopal M, Padmashali B, Vaidya VP (2005) Ind J Heterocycl Chem 15:169–172
Pavlović G, Racané L, Čičak H, Tralić-Kulenović V (2009) Dyes Pigments 83:354–362
Allen FH (2002) Acta Crystallogr B58:380–388
Groom CR, Bruno IJ, Lightfoot MP, Ward SC (2016) Acta Crystallogr B72:171–179
Lindgren EB, Yoneda JD, Leal KZ, Nogueira AF, Vasconcelos TRA, Wardell JL, Wardell AMV (2013) J Mol Struct 1036:19–27
Nogueira AF, Vasconcelos TRA, Wardell JL, Wardell SMSV (2011) Z Kristallogr 226:846–860
Vanucci-Bacqué C, Carayon C, Bernis C, Camare V, Nègre-Salvayre A, Bedos-Belval F, Baltas M (2014) Bioorg Med Chem 22:4269–4276
Patil SA, Weng CM, Huang PC, Hong FE (2009) Tetrahedron 65:2889–2897
Lindgren EB, de Brito MA, Vasconselos TRA, de Morales MO, Montenegro RC, Yoneda JD, Leal KZ (2014) Eur J Med Chem 86:12–16
Girish SR, Revankar VK, Mahale VB (1996) Transit Met Chem 21:401–405
X’Pert Sofware Suite, Version 1.3e, Panalytical B.V., Almelo, The Netherlands, 2001
Vreshch V (2011) J Appl Crystallogr 44:219–220
Oxford Diffraction Ltd., Xcalibur CCD system, CrysAlis Software system Versions 1.171.37.35. Abingdon, Oxfordshire, England, 2008
Sheldrick GM (2008) Acta Crystallogr Sect A64:112–122
Farrugia LJ (1999) J Appl Crystallogr 32:837–838
Spek L (2003) J Appl Crystallogr 36:7–13
Macrae CF, Bruno IJ, Chisholm JA, Edgington PR, McCabe P, Pidcock E, Rodriguez-Monge L, Taylor R, van de Streek J, Wood PA (2008) J Appl Crystallogr 41:466–470
Becke AD (1993) J Chem Phys 98:5648–5652
Lee C, Yang W, Parr RG (1998) Phys Rev B 37:785–789
Grimme S, Ehrlich S, Goerigk L (2011) J Comput Chem 32:1456–1465
Hrenar T (2014) qcc, Quantum Chemistry Code, rev. 0.68
Primožič I, Hrenar T, Baumann K, Krišto L, Križić I, Tomić S (2014) Croat Chem Acta 87:155–162
Hrenar T, moonee, Code for manipulation and analysis of multi- and univariate data, rev. 0.6826, 2014
Gaussian 09, Revision E.01, Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Mennucci B, Petersson GA, Nakatsuji H, Caricato M, Li X, Hratchian HP, Izmaylov AF, Bloino J, Zheng G, Sonnenberg JL, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Vreven T, Montgomery Jr JA, Peralta JE, Ogliaro F, Bearpark M, Heyd JJ, Brothers E, Kudin KN, Staroverov VN, Kobayashi R, Normand J, Raghavachari K, Rendell A, Burant JC, Iyengar SS, Tomasi J, Cossi M, Rega N, Millam JM, Klene M, Knox JE, Cross JB, Bakken V, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Martin RL, Morokuma K, Zakrzewski VG, Voth GA, Salvador P, Dannenberg JJ, Dapprich S, Daniels AD, Farkas Ö, Foresman JB, Ortiz JV, Cioslowski J, Fox DJ (2009) Gaussian Inc., Wallingford, CT. http://www.gaussian.com/g_tech/g_ur/m_citation.htm
Gvozdjakova A, Ivanovičova H (1986) Chem Papers 40:797–800
Easmon J, Heinisch G, Holzer W (1989) Heterocycles 29:1399–1408
Easmon J, Puerstinger G, Roth T, Fiebig HH, Jenny M, Jaeger W, Heinisch G, Hofmann J (2001) Int J Cancer 94:89–96
Acknowledgments
We acknowledge Krešimir Molčanov, PhD, Division of Physical Chemistry, Laboratory for chemical and biological crystallography, Ruđer Bošković Institute, Zagreb, Croatia, for X-ray single-crystal diffraction data collection for compound 8. We greatly appreciate support of University of Rijeka research Grant No. 13.11.1.1.11 and access to equipment in possession of University of Rijeka within the project “Research Infrastructure for Campus-based Laboratories at University of Rijeka”, financed by European Regional Development Fund (ERDF).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
11224_2016_856_MOESM1_ESM.docx
Electronic supplementary information (ESI) is available and it contains: elemental analysis (Table S2), spectral data (Figs. S1–S27), structural formulas of L1‒L9 with abbreviated atoms that define torsional coordinates (Fig. S28) and XRPD patterns (Figs. S29, S30). Full details of the crystal structure determinations in cif format are available in the online version, at doi (to be inserted) and have also been deposited with the Cambridge Crystallographic Data Centre with deposition numbers 1494759 (L1), 1494760 (L7), 1494761 (L8) and 1494762 (L9). Copies of these last can be obtained free of charge on written application to CCDC, 12 Union Road, Cambridge, CB2 1EZ, UK (Fax: +44 1223 336033); on request by email to deposit@ccdc.cam.ac.uk or by access to http://www.ccdc.cam.ac.uk. Supplementary data associated with this article can be found, in the online version, at (to be inserted) (DOCX 2422 kb)
Rights and permissions
About this article
Cite this article
Katava, R., Pavelić, S.K., Harej, A. et al. Supramolecular solid-state structure, potential energy surfaces and evaluation of antiproliferative effect of 2-benzothiazolylhydrazone derivatives in vitro. Struct Chem 28, 709–721 (2017). https://doi.org/10.1007/s11224-016-0856-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11224-016-0856-0