Abstract
Environmental biotechnology offers several promising techniques for the rehabilitation of polluted environments. The modern industrialized world presents novel challenges to the environmental sciences, requiring a constant development and deepening of knowledge to enable the characterization of novel pollutants and a better understanding of the bioremediation strategies as well as their limiting factors. The success of bioremediation depends heavily on the survival and activities of indigenous microbial communities and their interaction with introduced microorganisms. The majority of natural microbiomes remain uncultivated; therefore, further investigations focusing on their intrinsic functions in ecosystems are needed. In this review, we aimed to provide (a) a comprehensive overview of the presence of viable but nonculturable bacteria and yet-to-be-cultivated cells in nature and their diverse awakening strategies in response to, among other factors, signalling extracellular metabolites (autoinducers, resuscitation promoting factors, and siderophores); (b) an outline of the trends in isolating unculturable bacteria; and (c) the potential applications of these hidden players in rehabilitation processes.
Similar content being viewed by others
1 Introduction
Since the dawn of microbiology in the eighteenth century, microbiologists have primarily relied on obtaining pure cultures via traditional plating techniques. However, culture-dependent methods, which require the cultivation of microorganisms, are not capable of thoroughly depicting the current microbial diversity in the biosphere (Austin 2017). Although microbial diversity on Earth is impressively rich, more than 99% of the potentially 1011–1012 microbial species remain undiscovered to date (Locey and Lennon 2016) and only a small fraction can be cultured by current techniques (Rappé and Giovannoni 2003; Pedrós-Alió and Manrubia 2016; Hofer 2018; Hahn et al. 2019). Limited microbial availability by cultivation has created an emerging need to learn more about the missing species and their functions (Epstein 2013), since they have great environmental sustainability potential for bioremediation and bio-waste industries.
Regardless of the culturing techniques, culture-independent molecular based methods, such as, denaturing and temperature gradient gel electrophoresis (DGGE/TGGE), terminal restriction fragment length polymorphism (T-RFLP), 16S rDNA clone library preparation (Marzorati et al. 2008), fatty acid methyl esters (FAME) analysis (Cavigelli et al. 1995), and next generation sequencing (NGS) technology (Smets et al. 2016), are suitable for obtaining overall information about the genetic diversity and community structures of microorganisms, including unculturable bacteria. As a culture-independent technology, metagenomics requires the harvesting of the complete genetic material from the examined sample (Handelsman 2004; Rayu et al. 2012). Combined with other omics, such as; metatranscriptomics, metabolomics, or even single-cell genomics, it can contribute to the development and production of new enzymes, supported by metabolic engineering (ME) and recombinant DNA technology. Moreover, it broadens our knowledge of yet-to-be-cultured bacteria and their potentially novel pathways for degrading environmental pollutants; therefore, it affords new insights into their promising environmental applications (Schloss and Handelsman 2003; Riesenfeld et al. 2004; Villas-Bôas and Bruheim 2007; Wilson and Piel 2013; Overmann et al. 2017).
2 The role of microorganisms in bioremediation
Due to extensive industrial and agricultural activities, accidents during transportation or improper storage, large quantities of contaminants and xenobiotics are released into the environment (Vidali 2001). The pollution of soil, water, and air is one of the most serious current environmental issues. Pollutants (including persistent organic pollutants, heavy metals, nutrients, etc.) may be very diverse in their chemical structures and toxicological effects but most of them pose a serious risk to human health and jeopardise ecological systems (Gilden et al. 2010; Kang 2014).
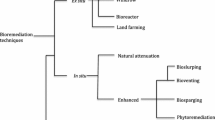
To date, a great variety of physico-chemical treatments have been available for removing these contaminants from the environment (Scullion 2006), but biological approaches are still among the most promising methods. Bioremediation based on natural processes utilizes the metabolic pathways of microorganisms (Laczi et al. 2015; Hegedüs et al. 2017; Hegedűs et al. 2017) (or plants) to neutralize pollutants and, thus, can be performed either ex situ or in situ (Vidali 2001; Perei et al. 2001; Kang 2014; Kis et al. 2015, 2017). A decrease in contamination can be achieved by stimulating the indigenous microflora of the polluted site with nutrient addition (i.e. biostimulation) or by introducing microbial degraders (i.e. bioaugmentation) into the site that requires remediation (Kis et al. 2017). In the case of complete mineralization of environmental pollutants, the whole process generates carbon dioxide, water and biomass as by-products, making bioremediation the most cost-effective and environmentally friendly remediation approach (Singh et al. 2011; Tyagi et al. 2011).
Bioremediation can be inhibited by several abiotic (temperature, pH, moisture, available electron acceptors or electron donors, etc.) and biotic (competition, predation, etc.) (Labana et al. 2005; Alvarez et al. 2006; Singh et al. 2011) factors and it has often been observed that, even though the environmental conditions for biodegradation are apparently optimal, efficiency is far below expectations. Recent studies have pointed out that strains exhibiting high pollutant biodegradation rates under controlled conditions in laboratories might be less effective and survive poorly in field-scale bioremediation (Megharaj et al. 2011; Kuppusamy et al. 2016) further supporting the existence of viable but nonculturable (VBNC) states in pollutant-degrading bacteria (Su et al. 2015b). Bacteria entering this a state in which they are alive, but have lost their culturability, could provide one of the explanations for the above observations.
3 VBNC state in bacteria
Until the first description of the phenomenon known as the ‘viable but nonculturable’ state in 1982 (Xu et al. 1982), bacterial cells deprived of their ability to grow on routinely-used laboratory media were considered to be dead. This study made the first attempt to distinguish viability from culturability. By now, many reports have revealed different physiological states in bacteria (Fig. 1) ranging from unstressed living cells to dead cells (Kell et al. 1998; Bergkessel et al. 2016; Hegedüs et al. 2017); a range which includes VBNC states. VBNC cells maintain their viability but unable to grow on routinely-used laboratory media (Oliver 2005).
Like other living organisms, microorganisms are able to interact with changing environmental conditions by triggering specific stress responses and survival mechanisms if the environmental parameters become suboptimal (Heimann 2002; Hegedüs et al. 2017). Bacteria can survive in extreme environments by forming spores (Hutchison et al. 2014) or entering a non-sporulating dormant state (Lennon and Jones 2011; Shoemaker and Lennon 2018). According to our current knowledge, a VBNC state is an adaptive strategy that serves long-term bacterial subsistence under adverse conditions (Li et al. 2014a; Oliver 2016) such as extreme temperatures; nutrient starvation; increased salinity (Oliver 1993); pH changes (Cunningham et al. 2009); osmotic stress; reactive oxygen species (ROS); and exposure to food preservatives, heavy metals (Li et al. 2014a), organic pollutants (Su et al. 2016), chlorination (Oliver et al. 2005; Chen et al. 2018), UV radiation (Ben Said et al. 2010) or even white light (Oliver 1993).
Although VBNC cells fail to grow or form colonies on laboratory media, they are very different from dead cells. VBNC cells preserve their cellular integrity, their membranes are not injured and they retain genomic or plasmid DNA. In contrast to dead cells, VBNC cells exhibit metabolic and respiratory activities, have high ATP levels, and may perform transcription and gene expression (Oliver 2005, 2010; Li et al. 2014a). Entering this survival state causes many physiological and molecular changes in VBNC cells compared to normal viable and culturable cells. Nutrient transport, and metabolic and respiratory activities are reduced in VBNC cells; cell wall and membrane composition, morphology, or gene expression level are also modified. Environmental factors, which cause VBNC states in bacteria, lead to increased crosslinking in peptidoglycan cell walls, alterations in the profile of outer membrane proteins or modifications in the composition of fatty acids in cytoplasmic membranes (Oliver 2010). Many bacterial species exhibit cell dwarfing or form coccoid-shaped cells in a VBNC state (Oliver 2000, 2005; Li et al. 2014a; Zhao et al. 2017b), suggesting that a reduced surface/volume ratio might help to minimize the energy requirement of cells (Li et al. 2014a). Morphological changes, however, cannot be considered as the sole decisive properties for defining a VBNC state.
Diverse bacterial genera were demonstrated to be capable of forming VBNC cells, implying the existence of various regulatory mechanisms underlying this process, but our knowledge of the genetic background is still limited. Two basic stress regulators, RpoS and OxyR, were found to play important role in inducing a VBNC state. Sigma factor, RpoS, is an essential regulator of general stress response. Expression of the rpoS gene is continuous in VBNC cells (Li et al. 2014a) but a lack of RpoS easily leads to cell death (Boaretti et al. 2003). RpoS plays a role in the survival of cells exposed to H2O2 (Oliver 2005) and may also affect the expression of other stress regulated genes that promote more effective stress responses and survival. OxyR is involved in regulating genes relating to oxidative stress (Oliver 2005; Li et al. 2014a).
Since the majority of microorganisms live in nutrient-limited natural habitats surrounded by biological competitors it can be hypothesized that this environmental context is responsible for the dominance of the unculturable microbial population on Earth, and a considerable proportion might be found in a VBNC state. According to a recent review, VBNC states have been described in more than 100 bacterial species belonging to almost 60 genera (Oliver 2016); however, some studies have revealed that eukaryotic cells, such as yeast Saccharomyces cerevisiae (Salma et al. 2013) and the vine spoilage yeast Brettanomyces bruxellensis could also enter this survival state in response to sulphite stress (Serpaggi et al. 2012).
4 The detection of VBNC bacteria
Although there are no specific methods available for the direct detection of VBNC bacteria, a combination of certain techniques may reveal the viability of VBNC bacteria (Table 1).
Despite the fact that they are alive, VBNC cells are no longer able to grow either on liquid or semi-solid standard bacteriological media; thus, a lack of colony formation by certainly living cells can be confirmed by using simple plating methods (Zhao et al. 2017b).
Kogure’s procedure (Kogure et al. 1979), which applied acridin orange staining to elongated living cells after exposure to a cell division inhibitor, nalidixic acid, allow the detection of normal-shaped VBNC or dormant cells under a microscope (Fakruddin et al. 2013). Once unculturability is confirmed, a wide range of fluorescent stains is available for fluorescent microscopy, which are mostly based on detecting the membrane integrity or enzymatic activity of tested cells. Counterstaining of microbes with 5-cyano-2,3-ditolyl tetrazolium chloride (CTC) and 4,6-diamino-2-phenyl indole (DAPI) is suitable for the simultaneous detection of total and viable bacteria (Ramamurthy et al. 2014). The LIVE/DEAD® BacLight™ bacteria viability kit combines two nucleic acid stains: green fluorescent SYTO® 9 for stainig living bacteria and red-fluorescent propidium iodide (PI), which passes only through the injured membrane of dead cells (Fakruddin et al. 2013). Owing to these properties, a LIVE/DEAD® BacLight™ kit is often used not only in fluorescent microscopy, but in fluorescent microplate readers, flow cytometers, and fluorimeters (Ramamurthy et al. 2014).
The combination of the direct viable count method (DVC), which applies the DNA-gyrase inhibitor novobiocin, and fluorescent in situ hybridization (FISH) made both the detection and enumeration of VBNC Helicobacter pylori cells possible (Piqueres et al. 2006).
Molecular methods that exclusively target DNA, such as normal PCR or qPCR, are not capable of differentiating viable cells from tdead ones. Combining qPCR with the microbial dye propidium monoazide (PMA), which penetrates only damaged membranes and inhibits PCR amplification by binding to the DNA of dead cells or extracellular DNA, enables the detection of nucleic acids exclusively in living cells (Ramamurthy et al. 2014; Zhao et al. 2017b). Since bacterial mRNA has a short half-life and is present only within the metabolically active cells, the latest studies have proposed a novel method for detecting cell viability by monitoring stress-related genes with reverse transcriptase PCR (RT-PCR) (Oliver 2010; Ramamurthy et al. 2014). Based on these considerations, molecular methods are not only suitable for distinguishing viable and dead cells, but when applied after a failed culturability test, can be used to detect VBNC bacteria.
5 The microbial scout hypothesis
Dormancy is a widespread phenomenon in nature that facilitates survival under hostile environmental conditions. Bacteria, including spore forming and non-sporulating species, are capable of entering a zero- or low-activity dormant state (Lennon and Jones 2011), but when conditions become optimal again, they reverse their dormancy. This reversal may be a result of external inducing effects (Dworkin and Shah 2010) or random awakening events (Paidhungat and Setlow 2000; Epstein 2009b). According to the theory proposed by Epstein (Epstein 2009a), dormant microbial populations periodically, but stochastically, form so-called ‘scout’ cells. A scout is a newly reactivated cell that has left dormancy in respone to infrequent and essentially random unknown internal events (Fig. 2). A scout is not a genetic variant and it does not differ from the typical growing cells of the population (Buerger et al. 2012a). A scout explores the available resources and initiates the transition from dormancy in the rest of the population by secreting intercellular metabolites as signals [e.g. autoinducers (AIs), resuscitation promoting factors (Rpfs), siderophores, catalases, etc.] when growth conditions are once again suitable (Epstein 2013; Pande and Kost 2017). Otherwise, the scout dies and a new scout will randomly arise in due course in the same dormant population (Buerger et al. 2012a).
Life cycle of unculturable microorganisms and their environmental potential. From time to time, uncultured and VBNC microorganisms, which make up the vast majority of microbial populations, randomly form scouts. These cells explore the available resources and can initiate the transition from a zero- or low-activity state in the rest of the population (resuscitation) by secreting certain signalling molecules. Artificial resuscitation of previously unculturable microorganisms, and utilization of their potential abilities, can open up novel prospects for environmental rehabilitation
Although some aspects of the hypothesis were controversial (Janssen 2009; Kell 2009) and the exact mechanism regulating scout formation remained undiscovered, Buerger and collegues experimentally supported the scout model of the microbial life cycle (Buerger et al. 2012a, b). The authors highlighted that the stochastic appearance of microbial scouts in a dormant bacterial community could explain, not only the random pattern of latent infections, but also the riddle of why a few cultivable bacteria exist in all VBNC populations (Bogosian and Bourneuf 2001). Recent studies produced evidence of spontaneous awakening (van Vliet 2015; Sturm and Dworkin 2015; Xu and Vetsigian 2017) and suggested that gene expression stochasticity, as a key factor in bacterial epigenetics, could impact transitions between states such as dormancy and scout formation independently of environmental cues (Bury-Moné and Sclavi 2017).
In the future, both human health sciences and environmental biotechnology could profit from the artificial awakening of inactive cells into scouts by using intercellular signalling molecules.
6 Bacterial resuscitation
Unlike dead cells, VBNC cells exhibit detectable metabolic activity, which allows them to recover culturability; hence, the whole process is reversible. The term ‘resuscitation’ was first used by Roszak and collegues in 1984 to describe the awakening of nonculturable Salmonella enteriditis cells (Roszak et al. 1984). The main challenge of resuscitation is to distinguish the regrowth of residual undetected culturable cells in a bacterial culture from the real recovery of VNBC cells (Zhao et al. 2017b).
It is hypothesized that bacteria have a so-called ‘resuscitation window’, defined by Pinto and collegues as ‘the period of time in which VBNC cells can resuscitate in response to the stimulus under study’ (Pinto et al. 2015). Although little is known about the precise length of the resuscitation window in bacteria, it varies considerably between different strains: in the case of Salmonella enteriditis the window lasts for 4 days (Roszak et al. 1984); in Micrococcus luteus, for 6 months (Mukamolova et al. 1998a) but in Citrobacter freundii for 11 years (Dhiaf et al. 2008). Supposing that a bacterial culture at a given time consists of a mixture of culturable cells and VBNC cells, under VBNC state-inducing conditions the number of culturable cells decreases, while the number of the latter increases; thus, cells are not the same age and resuscitation efficiency decreases over time (Pinto et al. 2015).
Since a VBNC state is triggered by changes in the environmental conditions mentioned previously, eliminating or neutralizing these stress factors can help to regain culturability. Favourable growth parameters, involving a favourable temperature, an increased energy source, a suitable nutrient concentration (with an ideal C/N ratio), chemical agents (e.g. ROS scavengers, antioxidants, the supernatant of actively growing cells), or co-culturing with other species, might contribute to the recovery of VBNC cells (Ramamurthy et al. 2014; Zhao et al. 2017b). In most of the cases, however, bacterial resuscitation is more complicated than simply removing the adverse factors. The resuscitation process differs between acteria (Dworkin and Shah 2010) and can be provoked by several stimuli (Table 2).
6.1 Autoinducers
A wide range of microorganisms have the ability to excrete the secondary metabolites or pheromones involved in regulating cellular differentiation, maintaining virulence, or communicating with conspecifics (Kell et al. 1995). Gene expression regulator autoinducers (AIs) are small, heat-stable, hormone-like signalling molecules mostly produced by Gram-negative bacteria that play an important role in cell-to-cell interspecies communication known as quorum sensing (Sperandio et al. 2003; Rutherford and Bassler 2012; Li et al. 2014a). AIs were discovered in Escherichia coli cultures growing in a norepinephrine containing serum-SAPI medium, in which growth rate stimulation and a reduction in lag phase were observed (Freestone et al. 1999).
Quorum sensing can be affected by several environmental factors (e.g. temperature, salinity, or pH) that have also been proved to trigger a VBNC state in bacteria, suggesting that some AIs might be involved in resuscitation (Ayrapetyan et al. 2014). Multiple bacterial strains produce autoinducer 2 (AI-2) molecules (Waters and Bassler 2006) further corroborating their potential cross-species activity (Bassler et al. 1997; Sperandio et al. 2003). Unculturable Vibrio cholerae cells were found to be resuscitated by AI-2 in aquatic reservoirs (Bari et al. 2013). Ayrapetyan and collegues proposed an AI-2-mediated resuscitation in V. vulnificus, in which the AI-2 level increased before regaining culturability and after reaching a threshold, it was able to serve as a resuscitation signal for the dormant population by stimulating rpoS expression (Ayrapetyan et al. 2014). Apart from purified autoinducers, Pinto and collegues demonstrated that the supernatant of growing cells in the late logarithmic phase also had a positive effect on bacterial resuscitation (Pinto et al. 2011).
6.2 Resuscitation promoting factors
Resuscitation promoting factor (Rpf) is a small, cytokine-like extracellular protein obtained from Microccus luteus cultures of actively growing cells on lactate minimal medium (LMM). It was first discovered in 1998 by Mukamolova and collegues (Mukamolova et al. 1998b, 2002, 2006; Su et al. 2014).
The gene family rpf is wide-spread across Gram-positive bacteria with high DNA G + C content. Aside from M. luteus, rpf gene homologues were found in other species of Actinobacteria such as Mycobacterium tuberculosis, M. leprae, M. smegmatis, Corynebacterium glutamicum and some Streptomyces species (Keep et al. 2006), which express Rpf-like proteins that share a highly conserved 70 amino acid segment called the Rpf-domain (Gupta and Srivastava 2012) and show similar characteristics and mechanisms to Rpf of M. luteus (Zhao et al. 2015). Despite the great abundance of rpf genes in Gram-positive bacteria, Panutdaporn and collegues explained the resuscitation of a food pathogen, Salmonella enterica, by a recombinant resuscitation promoting factor derived from the Gram-negative bacterium Salmonella typhimurium (Panutdaporn et al. 2006).
Like autoinducers, bacterial growth factor Rpf (or Rpf-containing M. luteus culture supernatant) also has a cross-species effect on stimulating cell growth, reducing lag phase, or helping dormant cells to revert from a VBNC state, even at picomolar concentrations (Shleeva et al. 2002) but it is sensitive to high temperatures and tryptic digestion (Mukamolova et al. 1998b). Rpf proteins display sequence homology with lysozymes and lytic transglycosylases (Cohen-Gonsaud 2004; Mukamolova et al. 2006; Nikitushkin et al. 2015), and they exhibit muralytic activity as a peptidoglycan hydrolase (Mukamolova et al. 2006; Telkov et al. 2006). In addition, a structure analysis of Rpf B obtained from M. tuberculosis revealed ubiquitin-like domains (Ruggiero et al. 2016). Boiling or exposure to ethanol can inactivate Rpf (Mukamolova et al. 2006).
Although Rpf-mediated resuscitation of VBNC cells is presumably based on the induction of cell expansion and division through the muralytic activity of Rpf, by making the cells more sensitive to environmental stimuli that support their growth (Keep et al. 2006; Mukamolova et al. 2006; Nikitushkin et al. 2016), the exact mechanism is still unclear. As an explanation for resuscitation by Rpfs, Pinto and collegues (Pinto et al. 2015) proposed three different models (Fig. 3). In the first, Rpf was considered to be a signalling molecule produced by actively-growing cells that bound to specific receptors on the surface of VBNC cells inducing resuscitation (Mukamolova et al. 1998b; Pinto et al. 2015); however, these receptors have not, so far, been detected. The second suggested that cleavage or remodelling of the peptidoglycan cell walls of dormant cells by Rpf was the initiating step for leaving VBNC state (Pinto et al. 2015). Sequence homology of Rpf with lysozymes and lytic transglycosylases further supported this theory (Telkov et al. 2006). The third model was based on the idea that, instead of being secreted into the growth medium, Rpf could be cell-bound (Mukamolova et al. 2002; Koltunov et al. 2010) and acts on the cell wall of producing cells, causing them to release small fragments of peptidoglycan that could bind to specific receptors on the surface of dormant cells and trigger reactivation from the VBNC state (Pinto et al. 2015).
Overview of the possible mechanisms for Rpf-mediated resuscitation according to Pinto and collegues (Pinto et al. 2015). a Rpf may be a signalling molecule produced by actively-growing cells that binds to specific receptors on the surface of VBNC cells, inducing resuscitation. b Cleavage or remodelling of peptidoglycan cell walls of dormant cells by Rpf can be an initiating step for transition from VBNC state. c Cell-bound Rpf may act on the cell walls of producing cells, causing them to release small fragments of peptidoglycan, which could bind to specific receptors on the surface of dormant cells and trigger reactivation from a VBNC state
6.3 Siderophores
As a central element of redox enzymes, iron is essential for basic cellular processes such as electron transport or synthesis of amino acids and DNA. Despite its important role in living organisms, the biological availability of soluble ferrous iron (Fe2+) is quite limited (Wandersman and Delepelaire 2004). Siderophores are small ferric ion (Fe3+) chelator molecules produced by microorganisms, scavenging insoluble iron then transporting it into the cells in case of iron shortage (Neilands 1995).
Siderophores also promote cell division; for example, the siderophore schizokinen can act as a growth factor by reducing the lag phase of Bacillus megaterium (Lankford et al. 1966). A lack of an available form of iron or siderophores may limit bacterial growth even in nutrient rich environments, suggesting that siderophores play an important role in the resuscitation of unculturable bacteria (Saha et al. 2016). D’Onofrio and collegues demonstrated the reactivation of dormant cells by accessing low molecular weight compounds produced by neighbouring members of the original microbial community (D’Onofrio et al. 2010). Lewis and collegues also identified siderophores from E. coli and M. luteus KLE1011 as growth factors for uncultured strains (Lewis et al. 2010). Many bacteria seem to be able to use the siderophores of other species despite their inability to produce them. This siderophore-dependence called ‘siderophore piracy’ is a widespread phenomenon in bacteria (D’Onofrio et al. 2010; Traxler et al. 2012). Considering the low cost of maintenance or the relatively high abundance of cases in which the horizontal transfer of the genes responsible for siderophore synthesis has been observed, losing these systems could provide a reductive evolutionary strategy in bacteria, known as the ‘black queen hypothesis’. According to this hypothesis, initial fast-growing strains are much more exposed to adverse environmental conditions, while slow growers are dependent on these co-existing strains and remain dormant until they can take advantage of their metabolites. This theory also supports the claim that previously unculturable bacteria cannot grow without external help (D’Onofrio et al. 2010; Morris et al. 2012).
7 Trends in isolating unculturable bacteria
Uncultivated bacteria are unable to grow in standard laboratory media due to their slow growth rates or transitions into dormancy. These species might be considered as K-strategists being adapted to limited resources and exhibit slow growth rates but having a stable existence in their habitat. Despite these bacterial species, r-strategist fast growers rapidly respond to nutrient flushing by blooming (Schmidt and Konopka 2009; Janssen 2009),; therefore, using standard cultivation techniques can result in an overgrowth of fast growers masking slow-growing or rare species (Poindexter 1981).
Based on these considerations, the first attempts to isolate previously uncultured bacteria were performed by mimicking natural conditions via decreased nutrient concentrations, decreased inoculum sizes and extended incubation times (Connon and Giovannoni 2002; Davis et al. 2005). Notably, besides the modification of isolating media formulation (substrates and nutrients) and factors that can be easily manipulated in the laboratory (e.g. pH, temperature, and oxygen availability), biotic factors such as synergistic interactions, secondary metabolite dependence or competition between species should also be taken into account during the development of new cultivation methods (Dewi Puspita et al. 2012; Pham and Kim 2012). According to the microbial scout hypothesis, Buerger and collegues suggested that given the same growth medium, a culturable but rare environmental species, with limited scout formation, might elude isolation for a long time; therefore, longer incubation is favourable for capturing novel species due to their stochastic awakening characteristics (Buerger et al. 2012b).
Since the natural milieu usually contains every nutrient required for bacterial growth, isolation techniques that stimulate the environment in the laboratory and in situ cultivation have been attracting attention during the last decade (Table 3). The main focus, however, has been on the resuscitation of VBNC bacteria (Table 4) in response to the addition of recombinant Rpf or extracellular organic matter (EOM) from M. luteus (also called supernatant Rpf or SRpf) (Su et al. 2012, 2015a; Li et al. 2014b, 2015; Zou et al. 2014; Jin et al. 2017). EOM is the sterile filtered culture supernatant of M. luteus grown on LMM and composed mainly of proteins and polysaccharides with concentrations of 25.1 and 405.7 mg L−1, respectively. Rpf has been revealed to be the dominant protein component in the EOM of M. luteus (Su et al. 2014).
8 Significance and environmental application of unculturable bacteria
Most of the microorganisms in natural environments live under suboptimal conditions. Lack of energy or resources and overcoming various stresses, such as unfavourable temperature and/or pH, the presence of competitors and predators, and similar can suppress their ability either to divide or survive (Haruta and Kanno 2015; Locey et al. 2017). Entering a dormant or low activity state serves as a survival strategy for microbes and hence a genetic and phenotypic diversity preserving mechanism. Diverse microbial seedbanks, which are large reservoirs of inactive individuals, deserve great attention and effort with a view to resuscitating these microorganisms and exploiting their highly promising potential for relevant ecological processes (Lennon and Jones 2011; Locey et al. 2017; Shoemaker and Lennon 2018).
Since the original paper regarding VBNC bacteria in 1982, many publications have attempted to describe the mechanisms underlying the formation, survival and resuscitation of VBNC bacteria. For pathogenic bacteria, the phenomenon has been deeply investigated and has become more popular, due to studies revealing the nature of latent infectious diseases in humans (Oliver 2010; Li et al. 2014a; Ramamurthy et al. 2014), focusing on the importance of food safety (Serpaggi et al. 2012; Fakruddin et al. 2013; Ayrapetyan and Oliver 2016; Shi et al. 2017; Zhao et al. 2017b) and seeking to develop new pharmacological products (Demain and Sanchez 2009). In spite of the abundance of information regarding VBNC pathogens, little is known about VBNC bacteria in polluted environments and their role in biological remediation techniques (Su et al. 2013a; Tripathi et al. 2017; Su et al. 2018c; Murugan and Vasudevan 2018). Given that most of the bacteria naturally occur in a VBNC state in the environment, these yet-to-be- cultured species are considered to be a vast unexplored pool of microbial resources (e.g. metabolic enzymes, biosurfactants, etc.) for environmental rehabilitation (Fig. 2).
8.1 Exploring the environmental potential of uncultivated bacteria
Recent studies have further supported the claim that uncultured bacteria may play an important role in the biodegradation of environmental pollutants. Metagenomic analysis showed a great abundance of aromatic-ring-hydroxylating oxygenase (RHO) genes, which were potentially responsible for polycyclic aromatic hydrocarbon (PAH) biodegradation, in uncultured microorganisms in chronically-polluted coastal sediments (Loviso et al. 2015). Exploitation of metagenomic data has proved to be beneficial for the enzymatic characterization of RHOs from yet-to-be-cultured microorganisms in polluted soils (Singleton et al. 2012; Martin et al. 2013; Chemerys et al. 2014) and marine environments (Musumeci et al. 2019) providing a better understanding of the catabolic versatility of the vast majority of uncultured microbes. Dong and collegues used shotgun metagenomics to discover uncultured bacterial and archaeal phyla capable of the anaerobic degradation of aliphatic and aromatic hydrocarbons from deep-sea sediments. Their results indicated that a rapid turnover of acetate and hydrogen can promote oxidization of organic compounds (Dong et al. 2019). Examining the bacterial composition of hydrocarbon-contaminated tropical African soil, Salam and collegues found uncultured genera that were involved in the natural attenuation of hydrocarbons (Salam et al. 2017). Furthermore, Chandra and Kumar detected uncultured bacterial members of the microbial community in a post-methanated distillery sludge (PMDS), which potentially played a significant role in neutralizing recalcitrant pollutants in distillery wastes and decreasing phyto- and genotoxicity (Chandra and Kumar 2017).
In addition to molecular approaches, the targeted isolation and resuscitation of previously uncultured bacteria by growth promoting factors have been gaining more attention in the past decade.
For a comparison of traditional plating and diffusion chamber methods, Bollmann and collegues isolated novel strains from subsurface sediments contaminated with uranium, heavy metals, nitrate, and organic pollutants using the chambers instead of the plates, and recovered isolates with potential for advantageous bioremediation (Bollmann et al. 2010). Zhao and collegues modified the microcultivation system of Ferrari and collegues (Ferrari et al. 2005) to obtain pollutant-degraders from crude oil contaminated soil. Comparing the in situ SSMS method to traditional nutrient-rich technique for isolation of phenanthrene-degrading bacteria, revealed that an in situ method that mimicks the original environment is more suitable for the growth of Sphingomonas. By contrast, traditional method resulted in the isolation of Pseudomonas. Both strains were able to degrade phenanthrene but the former seemed to be previously uncultured by nutrient-rich methods (Zhao et al. 2017a).
8.2 Potential environmental functions of VBNC bacteria
Although, there have been several examples for the resuscitation of previously VBNC bacteria from environmental samples (Ding et al. 2007, 2009; Li et al. 2007; Ding and Yokota 2010), isolation of a novel bioflocculant-producing actinomycete Arthrobacter sp. LC13T by resuscitation from a VBNC state was one of the first attempts to use Rpf for environmental purposes (Su et al. 2012; Fu et al. 2014). Since then, many studies have been carried out on the resuscitation (Table 4) and stimulation effects of Rpf (Table 5), with regard to the Rpf-containing EOM or recombinant Rpf of M. luteus. Jin and collegues resuscitated VBNC bacteria from dyeing wastewater, in which the Rpf-treated group proved to be more diverse than the Rpf-lacking control group indicating the high sensitivity of VBNC bacteria to Rpf in dyed water. All strains degraded more than 70% Congo red after 8 days. (Jin et al. 2017). A novel approach applied an Rpf-containing supernatant of M. luteus culture to stimulate VBNC or uncultured indigenous bacteria in PCB (polychlorinated biphenyl) contaminated environments. Rpf-responsive enrichment cultures degraded nearly 1500 mg L−1 biphenyl in 24 h at the optimal EOM dosage of 15% (v/v). This was the first case when Rpf was used to promote biodegradation of toxic compounds (Su et al. 2013b). Further exploring the potential environmental applications of supernatant Rpf, Liu and collegues achieved promising results for biological nutrient removal from wastewater. Although optimal dosage assessment is still needed, the presence of Rpf increased the abundance of the phyla Proteobacteria and Actinobacteria, which were responsible for the enhanced removal of phosphorus and nitrogen in activated sludge systems (Liu et al. 2016). Bounedjoum and collegues assessed a potentially functional hydrocarbon-degrader community enriched from oil-contaminated soil in response to M. luteus EOM addition (Bounedjoum et al. 2018). Among other strains, Rhodococcus biphenylivorans TG9T was isolated from a PCB-contaminated river sediment using the resuscitating and stimulating effects of EOM. EOM significantly increased both the abundance of potentially difficult-to-culture biphenyl-degraders in enrichment cultures and the biodegradation of biphenyls (Su et al. 2015a, c, d). Transcriptome analysis of R. biphenylivorans TG9T, Rhodococcus sp. TG13 and TN3 not only verified that pollutant-degrading bacteria could enter a VBNC state under cold stress and revert from it in response to the addition of supernatant Rpf but also pointed out molecular mechanisms behind these reversible processes that may vary with different bacterial species (Su et al. 2015b, 2016). Their work drew attention to the importance of preventing and controlling the formation of VBNC bacteria during bioremediation, which can be crucial in the case of bioaugmentation.
Analysing the fate of transformant soil bacteria capand colleaguesble of degrading the agricultural insecticide lindene (γ-hexachlorocyclohexane), Zhang and colleagues found that cell numbers decreased below the detection limit and after the addition of nutrients to the microcosms, they became detectable again demonstrating the low-activity state of these engineered strains when exposed to adverse environmental conditions (Zhang et al. 2012). Higher cell numbers of Rhodococcus sp. D310-1::gfp detected by fluorescence microscopy compared to colony forming units (CFUs) indicated the formation of VBNC cells in chlorimuron-ethyl-contaminated soil (Xiong et al. 2013). Fida and collegues had similar observations when a GFP-tagged PAH-degrader Novosphingobium sp. LH128 was inoculated into phenanthrene spiked soil. A rapid decline was detected in CFUs without any reduction in the number of GFP-expressing cells. Transcriptome analysis showed high expression levels of stress-related genes and phenanthrene biodegradation activity suggesting that strain LH128 entered a VBNC-like state (Fida et al. 2017). Performing proteomic analysis on the soil bacterium Cupriavidus metallidurans CH34 under water scarcity and nutrient starvation, Giagnoni and colleagues monitored metabolic changes (e.g. expression of genes related to cell shape formation and stress proteins) in C. metallidurans CH34 during its transition into a VBNC state, then its reversion to culturable cells after the addition of water or gluconate as the sole carbon source (Giagnoni et al. 2018). Their results further proved the importance of sigma factors in the formation of VBNC cells, but also suggested that such soil properties as water content or carbon sources could be associated with the intracellular metabolisms of soil bacteria, hence influencing the survival, recolonization or spatial distribution of microbial communities in restored soils.
In addition to the use of Rpf-containing EOM, the application of purified recombinant Rpf protein is starting to attract more attention. Recovery of Rhodococcus sp. DS471 cells from a VBNC state was the first report of the resuscitation ability of the recombinant Micrococcus luteus Rpf expressed in Escherichia coli (Ding et al. 2012). Aiming to map the importance of VBNC bacteria in industrial wastewater and improve sewage treatment systems, Gordonia jinhuaensis ZYR51T (as a novel species of the genus Gordonia) was isolated, along with other strains, from pharmaceutical wastewater, using Rpf (Fu et al. 2014; Li et al. 2014b; Zou et al. 2014). The novel strains Rhodococcus soli DSD51WT (Li et al. 2015) and Arthrobacter liuii DSXY973T (Yu et al. 2015) belonging to the genus Rhodococcus and Arthrobacter, respectively, were resuscitated from soil samples. Use of recombinant Rpf protein promoted the isolation of unique bacterial species from waste composting sample and enhanced the cellulase activity of enrichment cultures. Filter paper cellulase (FPCase) and carboxymethyl-cellulase (CMCase) were detected in both pure and mixed cultures. Cellulase activities were higher in both mixed and Rpf-supplemented cultures than in the pure and Rpf-lacking ones, respectively (Su et al. 2018b). Su and colleagues resuscitated functional strains from a highly nitrogen-polluted river sediment that reached ammonium and nitrate removal rates of 2.23 and 0.86 mg L−1 h−1, respectively. The results highlighted that indigenous VBNC bacteria in polluted rivers and water bodies can be considered useful resources for biological nitrogen removal (Su et al. 2019c). A 1% (v/v) concentration of purified Rpf proved to be an enhancing additive for bacteria in activated sludge facilitating their activity during the biological treatment of a phenolic wastewater under high salinity conditions. Rpf-treatment not only shortened the whole domestication process for activated sludge, but promoted the biodegradation of 1800 mg L−1 phenol at 60 g L−1 NaCl concentration in 18 days (Su et al. 2018a). Better phenol removal performance was also achieved in membrane bioreactors (MBRs) with Rpf addition. Nearly 1500 mg L−1 of phenol was degraded within 100 h in phenol-laden saline wastewater probably due to the resuscitation and stimulation of gammaproteobacterial and alphaproteobacterial populations (Su et al. 2019b). In a search for further halotolerant phenol-degrader bacteria, several strains were isolated from a sewage treatment tank (Li et al. 2018) and activated sludge samples (Su et al. 2019b) by Rpf. Neutralization of phenolic wastewaters by applying Bacillus sp. SAS19, either in a mixed consortium with Corynebacterium sp. SAS21 (Li et al. 2018) or immobilized in porous gels (Ke et al. 2018), represented the first application of a previously VBNC bacterium. Castellaniella sp. SPC4, another former VBNC bacterium resuscitated by Rpf, was found to be capable of effectively catabolizing 3,3′,4,4′-tetrachlorobiphenyl (PCB 77). The strain maintained the capability of PCB biodegradation even without Rpf supplementation, further proving the degradative potential of VBNC bacteria in contaminated environments (Su et al. 2019a). The study of their capabilities is one of the major future tasks of environmental biotechnology.
Considering the results mentioned above and the acknowledged fact that Rpf is able to resuscitate potential pollutant-degrading VBNC bacteria, its application can be integrated into biostimulation methods of bioremediation (which raises the question of the role of non-protein components in EOM). By contrast, it is assumed that the outcome of bioaugmentation, which involves the inoculation of efficient pollutant-degrader bacteria into polluted, heavily depends not only on the survival and efficiency of these strains, but also on their synergistic degradation aptitudes and transitions into VBNC states (Su et al. 2015b).
9 Conclusion
Nowadays, the most crucial challenges of bioremediation are to recover the hidden players in microbial diversity and overcome the obstacles to moving pollutant-degrader microorganisms from laboratory flasks to field sites. It is essential to enhance their survival and biodegradation capabilities in order to intensify their bioconversion rates, even for field remediation processes. The future success of environmental biotechnology depends heavily on developing and improving databases of the microorganisms and enzymes that are responsible for biodegradation of environmental pollutants. A deeper understanding of the metabolic pathways involved in the microbial degradation of recalcitrant contaminants requires a better knowledge about the unculturable majority of bacteria, their relationships with coexisting members of the microbial community, and how they can be integrated as key players in environmental rehabilitation processes.
References
Alvarez PJJ, Illman WA, (Walter A, Wiley InterScience (Online service) (2006) Bioremediation and natural attenuation: process fundamentals and mathematical models. Wiley, Chichester
Austin B (2017) The value of cultures to modern microbiology. Antonie van Leeuwenhoek Int J Gen Mol Microbiol. https://doi.org/10.1007/s10482-017-0840-8
Ayrapetyan M, Oliver JD (2016) The viable but non-culturable state and its relevance in food safety. Curr Opin Food Sci 8:127–133. https://doi.org/10.1016/j.cofs.2016.04.010
Ayrapetyan M, Williams TC, Oliver JD (2014) Interspecific quorum sensing mediates the resuscitation of viable but nonculturable vibrios. Appl Environ Microbiol 80:2478–2483. https://doi.org/10.1128/AEM.00080-14
Bari SMN, Roky MK, Mohiuddin M et al (2013) Quorum-sensing autoinducers resuscitate dormant Vibrio cholerae in environmental water samples. Proc Natl Acad Sci USA 110:9926–9931. https://doi.org/10.1073/pnas.1307697110
Bassler BL, Greenberg EP, Stevens AM (1997) Cross-species induction of luminescence in the quorum-sensing bacterium Vibrio harveyi. J Bacteriol 179:4043–4045. https://doi.org/10.1128/JB.179.12.4043-4045.1997
Ben Said M, Masahiro O, Hassen A (2010) Detection of viable but non cultivable Escherichia coli after UV irradiation using a lytic Qβ phage. Ann Microbiol 60:121–127. https://doi.org/10.1007/s13213-010-0017-4
Bergkessel M, Basta DW, Newman DK (2016) The physiology of growth arrest: uniting molecular and environmental microbiology. Nat Rev Microbiol 14:549–562. https://doi.org/10.1038/nrmicro.2016.107
Boaretti M, Lleò MDM, Bonato B et al (2003) Involvement of rpoS in the survival of Escherichia coli in the viable but non-culturable state. Environ Microbiol 5:986–996. https://doi.org/10.1046/j.1462-2920.2003.00497.x
Bogosian G, Bourneuf EV (2001) A matter of bacterial life and death. EMBO Rep 2:770–774. https://doi.org/10.1093/embo-reports/kve182
Bollmann A, Palumbo AV, Lewis K, Epstein SS (2010) Isolation and physiology of bacteria from contaminated subsurface sediments. Appl Environ Microbiol 76:7413–7419. https://doi.org/10.1128/AEM.00376-10
Bounedjoum N, Bodor A, Laczi K et al (2018) Assessment of potentially functional hydrocarbon-degrader bacterial communities in response to Micrococcus luteus EOM using culture-dependent and culture-independent methods. New Biotechnol 44:S134–S135. https://doi.org/10.1016/j.nbt.2018.05.1091
Buerger S, Spoering A, Gavrish E et al (2012a) Microbial scout hypothesis, stochastic exit from dormancy, and the nature of slow growers. Appl Environ Microbiol 78:3221–3228. https://doi.org/10.1128/AEM.07307-11
Buerger S, Spoering A, Gavrish E et al (2012b) Microbial scout hypothesis and microbial discovery. Appl Environ Microbiol 78:3229–3233. https://doi.org/10.1128/AEM.07308-11
Bury-Moné S, Sclavi B (2017) Stochasticity of gene expression as a motor of epigenetics in bacteria: from individual to collective behaviors. Res Microbiol. https://doi.org/10.1016/j.resmic.2017.03.009
Cavigelli MA, Robertson GP, Klug MJ (1995) Fatty acid methyl ester (FAME) profiles as measures of soil microbial community structure. Plant Soil 170:99–113. https://doi.org/10.1007/BF02183058
Chandra R, Kumar V (2017) Detection of androgenic-mutagenic compounds and potential autochthonous bacterial communities during in situ bioremediation of post-methanated distillery sludge. Front Microbiol 8:1–18. https://doi.org/10.3389/fmicb.2017.00887
Chemerys A, Pelletier E, Cruaud C et al (2014) Characterization of novel polycyclic aromatic hydrocarbon dioxygenases from the bacterial metagenomic DNA of a contaminated soil. Appl Environ Microbiol 80:6591–6600. https://doi.org/10.1128/AEM.01883-14
Chen S, Li X, Wang Y et al (2018) Induction of Escherichia coli into a VBNC state through chlorination/chloramination and differences in characteristics of the bacterium between states. Water Res 142:279–288. https://doi.org/10.1016/J.WATRES.2018.05.055
Cohen-Gonsaud M (2004) Resuscitation-promoting factors possess a lysozyme-like domain. Trends Biochem Sci 29:7–10. https://doi.org/10.1016/j.tibs.2003.10.009
Connon SA, Giovannoni SJ (2002) High-throughput methods for culturing microorganisms in very-low-nutrient media yield diverse new marine isolates. Appl Environ Microbiol 68:3878–3885. https://doi.org/10.1128/AEM.68.8.3878-3885.2002
Cunningham E, O’Byrne C, Oliver JD (2009) Effect of weak acids on Listeria monocytogenes survival: evidence for a viable but nonculturable state in response to low pH. Food Control 20:1141–1144. https://doi.org/10.1016/j.foodcont.2009.03.005
D’Onofrio A, Crawford JM, Stewart EJ et al (2010) Siderophores from neighboring organisms promote the growth of uncultured bacteria. Chem Biol 17:254–264. https://doi.org/10.1016/j.chembiol.2010.02.010
Davis KER, Joseph SJ, Janssen PH (2005) Effects of growth medium, inoculum size, and incubation time on culturability and isolation of soil bacteria. Appl Environ Microbiol 71:826–834. https://doi.org/10.1128/AEM.71.2.826-834.2005
Demain AL, Sanchez S (2009) Microbial drug discovery: 80 years of progress. J Antibiot (Tokyo) 62:5–16. https://doi.org/10.1038/ja.2008.16
Dewi Puspita I, Kamagata Y, Tanaka M et al (2012) Are uncultivated bacteria really uncultivable? Microbes Environ 27:356–366. https://doi.org/10.1264/jsme2.ME12092
Dhiaf A, Bakhrouf A, Witzel K-P (2008) Resuscitation of eleven-year VBNC Citrobacter. J Water Health 6:565–568
Ding L, Yokota A (2010) Curvibacter isolated from well. Water 271:267–271
Ding L, Hirose T, Yokota A (2007) Amycolatopsis echigonensis sp. nov. and Amycolatopsis niigatensis sp. nov., novel actinomycetes isolated from a filtration substrate. Int J Syst Evol Microbiol 57:1747–1751. https://doi.org/10.1099/ijs.0.64791-0
Ding L, Hirose T, Yokota A (2009) Four novel Arthrobacter species isolated from filtration substrate. Int J Syst Evol Microbiol 59:856–862. https://doi.org/10.1099/ijs.0.65301-0
Ding L, Zhang P, Hong H et al (2012) Cloning and expression of Micrococcus luteus IAM 14879 Rpf and its role in the recovery of the VBNC state in Rhodococcus sp. DS471. Wei Sheng Wu Xue Bao 52:77–82
Dong X, Greening C, Rattray JE et al (2019) Metabolic potential of uncultured bacteria and archaea associated with petroleum seepage in deep-sea sediments. Nat Commun 10:1816. https://doi.org/10.1038/s41467-019-09747-0
Dworkin J, Shah IM (2010) Exit from dormancy in microbial organisms. Nat Rev Microbiol 8:890–896. https://doi.org/10.1038/nrmicro2453
Epstein SS (2009a) General model of microbial uncultivability. Springer, Berlin, pp 131–159
Epstein SS (2009b) Microbial awakenings. Nature 457:1083. https://doi.org/10.1038/4571083a
Epstein SS (2013) The phenomenon of microbial uncultivability. Curr Opin Microbiol 16:636–642. https://doi.org/10.1016/j.mib.2013.08.003
Fakruddin M, Bin Mannan KS, Andrews S (2013) Viable but nonculturable bacteria: food safety and public health perspective. ISRN Microbiol 2013:1–6. https://doi.org/10.1155/2013/703813
Ferrari BC, Binnerup SJ, Gillings M (2005) Microcolony cultivation on a soil substrate membrane system selects for previously uncultured soil bacteria. Appl Environ Microbiol 71:8714–8720. https://doi.org/10.1128/AEM.71.12.8714-8720.2005
Fida TT, Moreno-Forero SK, Breugelmans P et al (2017) Physiological and transcriptome response of the polycyclic aromatic hydrocarbon degrading Novosphingobium sp. LH128 after inoculation in soil. Environ Sci Technol. https://doi.org/10.1021/acs.est.6b03822
Freestone PP, Haigh RD, Williams PH et al (1999) Stimulation of bacterial growth by heat-stable, norepinephrine-induced autoinducers. FEMS Microbiol Lett 172:53–60. https://doi.org/10.1111/j.1574-6968.1999.tb13449.x
Fu H, Wei Y, Zou Y et al (2014) Research progress on the Actinomyces arthrobacter. Adv Microbiol 4:747–753
Gavrish E, Bollmann A, Epstein S, Lewis K (2008) A trap for in situ cultivation of filamentous actinobacteria. J Microbiol Methods 72:257–262. https://doi.org/10.1016/j.mimet.2007.12.009
Giagnoni L, Arenella M, Galardi E et al (2018) Bacterial culturability and the viable but non-culturable (VBNC) state studied by a proteomic approach using an artificial soil. Soil Biol Biochem 118:51–58. https://doi.org/10.1016/J.SOILBIO.2017.12.004
Gilden RC, Huffling K, Sattler B (2010) Pesticides and health risks. JOGNN J Obstet Gynecol Neonatal Nurs 39:103–110. https://doi.org/10.1111/j.1552-6909.2009.01092.x
Gupta RK, Srivastava R (2012) Resuscitation promoting factors: a family of microbial proteins in survival and resuscitation of dormant mycobacteria. Indian J Microbiol 52:114–121. https://doi.org/10.1007/s12088-011-0202-6
Hahn MW, Koll U, Schmidt J (2019) Isolation and cultivation of bacteria. Springer, Cham, pp 313–351
Handelsman J (2004) Metagenomics: application of genomics to uncultured microorganisms. Microbiol Mol Biol Rev 68:669–685. https://doi.org/10.1128/MMBR.68.4.669-685.2004
Haruta S, Kanno N (2015) Survivability of microbes in natural environments and their ecological impacts. Microbes Environ 30:123–125. https://doi.org/10.1264/jsme2.ME3002rh
Hegedüs B, Kós PB, Bende G et al (2017) Starvation- and xenobiotic-related transcriptomic responses of the sulfanilic acid-degrading bacterium, Novosphingobium resinovorum SA1. Appl Microbiol Biotechnol. https://doi.org/10.1007/s00253-017-8553-5
Hegedűs B, Kós PB, Bálint B et al (2017) Complete genome sequence of Novosphingobium resinovorum SA1, a versatile xenobiotic-degrading bacterium capable of utilizing sulfanilic acid. J Biotechnol 241:76–80. https://doi.org/10.1016/j.jbiotec.2016.11.013
Heimann JD (2002) The extracytoplasmic function (ECF) sigma factors. Adv Microb Physiol 46:47–110
Hofer U (2018) The majority is uncultured. Nat Rev Microbiol 16:716–717. https://doi.org/10.1038/s41579-018-0097-x
Hutchison EA, Miller DA, Angert ER (2014) Sporulation in bacteria: beyond the standard model. Microbiol Spectr. https://doi.org/10.1128/microbiolspec.TBS-0013-2012.Correspondence
Janssen PH (2009) Dormant microbes: scouting ahead or plodding along? Nature 458:831. https://doi.org/10.1038/458831a
Jin Y, Gan G, Yu X et al (2017) Isolation of viable but non-culturable bacteria from printing and dyeing wastewater bioreactor based on resuscitation promoting factor. Curr Microbiol 74:787–797. https://doi.org/10.1007/s00284-017-1240-z
Kaeberlein T, Lewis K, Epstein SS (2002) Isolating “uncultivable” microorganisms in pure culture in a simulated natural environment. Science 296:1127–1129. https://doi.org/10.1126/science.1070633
Kang JW (2014) Removing environmental organic pollutants with bioremediation and phytoremediation. Biotechnol Lett 36:1129–1139. https://doi.org/10.1007/s10529-014-1466-9
Ke Q, Zhang Y, Wu X et al (2018) Sustainable biodegradation of phenol by immobilized Bacillus sp. SAS19 with porous carbonaceous gels as carriers. J Environ Manag 222:185–189. https://doi.org/10.1016/J.JENVMAN.2018.05.061
Keep NH, Ward JM, Cohen-Gonsaud M, Henderson B (2006) Wake up! Peptidoglycan lysis and bacterial non-growth states. Trends Microbiol 14:271–276. https://doi.org/10.1016/j.tim.2006.04.003
Kell D (2009) Dormant microbes: time to revive some old ideas. Nature 458:831. https://doi.org/10.1038/458831b
Kell DB, Kaprelyants AS, Grafen A (1995) Pheromones, social behaviour and the functions of secondary metabolism in bacteria. Trends Ecol Evol 10:126–129. https://doi.org/10.1016/S0169-5347(00)89013-8
Kell DB, Kaprelyants AS, Weichart DH et al (1998) Viability and activity in readily culturable bacteria: a review and discussion of the practical issues. Antonie van Leeuwenhoek Int J Gen Mol Microbiol 73:169–187. https://doi.org/10.1023/A:1000664013047
Kis Á, Laczi K, Zsíros S et al (2015) Biodegradation of animal fats and vegetable oils by Rhodococcus erythropolis PR4. Int Biodeterior Biodegrad 105:114–119. https://doi.org/10.1016/j.ibiod.2015.08.015
Kis ÁE, Laczi K, Zsíros S et al (2017) Characterization of the Rhodococcus sp. MK1 strain and its pilot application for bioremediation of diesel oil-contaminated soil. Acta Microbiol Immunol Hung 64:463–482. https://doi.org/10.1556/030.64.2017.037
Kogure K, Simidu U, Taga N (1979) A tentative direct microscopic method for counting living marine bacteria. Can J Microbiol 25:415–420. https://doi.org/10.1139/m79-063
Koltunov V, Greenblatt CL, Goncharenko AV et al (2010) Structural changes and cellular localization of resuscitation-promoting factor in environmental isolates of Micrococcus luteus. Microb Ecol 59:296–310. https://doi.org/10.1007/s00248-009-9573-1
Kuppusamy S, Palanisami T, Megharaj M et al (2016) In-situ remediation approaches for the management of contaminated sites: a comprehensive overview. Springer, Berlin, pp 1–115
Labana S, Singh OV, Basu A et al (2005) A microcosm study on bioremediation of p-nitrophenol-contaminated soil using Arthrobacter protophormiae RKJ100. Appl Microbiol Biotechnol. https://doi.org/10.1007/s00253-005-1926-1
Laczi K, Kis Á, Horváth B et al (2015) Metabolic responses of Rhodococcus erythropolis PR4 grown on diesel oil and various hydrocarbons. Appl Microbiol Biotechnol 99:9745–9759. https://doi.org/10.1007/s00253-015-6936-z
Lankford CE, Walker JR, Reeves JB et al (1966) Inoculum-dependent division lag of Bacillus cultures and its relation to an endogenous factor(s) (“schizokinen”). J Bacteriol 91:1070–1079
Lennon JT, Jones SE (2011) Microbial seed banks: the ecological and evolutionary implications of dormancy. Nat Rev Microbiol 9:119–130. https://doi.org/10.1038/nrmicro2504
Lewis K, Epstein S, D’Onofrio A, Ling LL (2010) Uncultured microorganisms as a source of secondary metabolites. J Antibiot (Tokyo) 63:468–476. https://doi.org/10.1038/ja.2010.87
Li B, Furihata K, Ding LX, Yokota A (2007) Rhodococcus kyotonensis sp. nov., a novel actinomycete isolated from soil. Int J Syst Evol Microbiol 57:1956–1959. https://doi.org/10.1099/ijs.0.64770-0
Li L, Mendis N, Trigui H et al (2014a) The importance of the viable but non-culturable state in human bacterial pathogens. Front Microbiol 5:1. https://doi.org/10.3389/fmicb.2014.00258
Li SH, Jin Y, Cheng J et al (2014b) Gordonia jinhuaensis sp. nov., a novel actinobacterium, isolated from a VBNC (viable but non-culturable) state in pharmaceutical wastewater. Antonie van Leeuwenhoek Int J Gen Mol Microbiol 106:347–356. https://doi.org/10.1007/s10482-014-0207-3
Li SH, Yu XY, Park DJ et al (2015) Rhodococcus soli sp. nov., an actinobacterium isolated from soil using a resuscitative technique. Antonie van Leeuwenhoek Int J Gen Mol Microbiol 107:357–366. https://doi.org/10.1007/s10482-014-0334-x
Li Z, Zhang Y, Wang Y et al (2018) A new approach of Rpf addition to explore bacterial consortium for enhanced phenol degradation under high salinity conditions. Curr Microbiol 75:1046–1054. https://doi.org/10.1007/s00284-018-1489-x
Liu Y, Su X, Lu L et al (2016) A novel approach to enhance biological nutrient removal using a culture supernatant from Micrococcus luteus containing resuscitation-promoting factor (Rpf) in SBR process. Environ Sci Pollut Res 23:4498–4508. https://doi.org/10.1007/s11356-015-5603-3
Locey KJ, Lennon JT (2016) Scaling laws predict global microbial diversity. Proc Natl Acad Sci USA 113:5970–5975. https://doi.org/10.1073/pnas.1521291113
Locey KJ, Fisk MC, Lennon JT (2017) Microscale insight into microbial seed banks. Front Microbiol 7:2040. https://doi.org/10.3389/fmicb.2016.02040
Loviso CL, Lozada M, Guibert LM et al (2015) Metagenomics reveals the high polycyclic aromatic hydrocarbon-degradation potential of abundant uncultured bacteria from chronically polluted subantarctic and temperate coastal marine environments. J Appl Microbiol 119:411–424. https://doi.org/10.1111/jam.12843
Martin F, Malagnoux L, Violet F et al (2013) Diversity and catalytic potential of PAH-specific ring-hydroxylating dioxygenases from a hydrocarbon-contaminated soil. Appl Microbiol Biotechnol 97:5125–5135. https://doi.org/10.1007/s00253-012-4335-2
Marzorati M, Wittebolle L, Boon N et al (2008) How to get more out of molecular fingerprints: practical tools for microbial ecology. Environ Microbiol 10:1571–1581. https://doi.org/10.1111/j.1462-2920.2008.01572.x
Megharaj M, Ramakrishnan B, Venkateswarlu K et al (2011) Bioremediation approaches for organic pollutants: a critical perspective. Environ Int 37:1362–1375. https://doi.org/10.1016/j.envint.2011.06.003
Morris JJ, Lenski RE, Zinser ER (2012) The Black Queen Hypothesis: evolution of dependencies through adaptive gene loss. MBio 3:e00036-12. https://doi.org/10.1128/mBio.00036-12
Mukamolova GV, Yanopolskaya ND, Kell DB, Kaprelyants AS (1998a) On resuscitation from the dormant state of Micrococcus luteus. Antonie Van Leeuwenhoek 73:237–243. https://doi.org/10.1023/A:1000881918216
Mukamolova GV, Kaprelyants AS, Young DI et al (1998b) A bacterial cytokine. Proc Natl Acad Sci 95:8916–8921. https://doi.org/10.1073/pnas.95.15.8916
Mukamolova GV, Turapov OA, Kazarian K et al (2002) The rpf gene of Micrococcus luteus encodes an essential secreted growth factor. Mol Microbiol 46:611–621. https://doi.org/10.1046/j.1365-2958.2002.03183.x
Mukamolova GV, Murzin AG, Salina EG et al (2006) Muralytic activity of Micrococcus luteus Rpf and its relationship to physiological activity in promoting bacterial growth and resuscitation. Mol Microbiol 59:84–98. https://doi.org/10.1111/j.1365-2958.2005.04930.x
Murugan K, Vasudevan N (2018) Intracellular toxicity exerted by PCBs and role of VBNC bacterial strains in biodegradation. Ecotoxicol Environ Saf 157:40–60. https://doi.org/10.1016/J.ECOENV.2018.03.014
Musumeci MA, Loviso CL, Lozada M et al (2019) Substrate specificities of aromatic ring-hydroxylating oxygenases of an uncultured gammaproteobacterium from chronically-polluted subantarctic sediments. Int Biodeterior Biodegrad 137:127–136. https://doi.org/10.1016/J.IBIOD.2018.12.005
Neilands JB (1995) Siderophores: structure and function of microbial iron transport compounds. J Biol Chem 270:26723–26726. https://doi.org/10.1074/JBC.270.45.26723
Nichols D, Cahoon N, Trakhtenberg EM, et al (2010) Use of ichip for high-throughput in situ cultivation of “uncultivable” microbial species. Appl Environ Microbiol 76:2445–2450. https://doi.org/10.1128/AEM.01754-09
Nikitushkin VD, Demina GR, Shleeva MO et al (2015) A product of RpfB and RipA joint enzymatic action promotes the resuscitation of dormant mycobacteria. FEBS J 282:2500–2511. https://doi.org/10.1111/febs.13292
Nikitushkin VD, Demina GR, Kaprelyants AS (2016) Rpf proteins are the factors of reactivation of the dormant forms of actinobacteria. Biochemistry 81:1719–1734. https://doi.org/10.1134/S0006297916130095
Oliver JD (1993) Formation of viable but nonculturable cells. In: Kjelleberg S (ed) Starvation in bacteria. Springer, Boston, MA
Oliver J (2000) The viable but nonculturable state and cellular resuscitation. In: Bell CR, Brylinsky M, Johnson-Green P (eds) Microbial systems: new frontiers. Atlantic Canada Society for Microbial Ecology, Halifax, pp 723–730
Oliver JD (2005) The viable but nonculturable state in bacteria. J Microbiol 43:93–100
Oliver JD (2010) Recent findings on the viable but nonculturable state in pathogenic bacteria. FEMS Microbiol Rev 34:415–425. https://doi.org/10.1111/j.1574-6976.2009.00200.x
Oliver JD (2016) The viable but nonculturable state in bacteria. Status update. This dormant form of bacteria was first appreciated in 1982; now skeptics recognize this state as a bacterial response to stress and a strategy for survival. J Microbiol 43:93–100
Oliver JD, Dagher M, Linden K (2005) Induction of Escherichia coli and Salmonella typhimurium into the viable but nonculturable state following chlorination of wastewater. J Water Health 3:249–257
Overmann J, Abt B, Sikorski J (2017) Present and future of culturing bacteria. Annu Rev Microbiol 71:711–730. https://doi.org/10.1146/annurev-micro-090816-093449
Paidhungat M, Setlow P (2000) Role of ger proteins in nutrient and nonnutrient triggering of spore germination in Bacillus subtilis. J Bacteriol 182:2513–2519. https://doi.org/10.1128/JB.182.9.2513-2519.2000
Pande S, Kost C (2017) Special issue: from one to many bacterial unculturability and the formation of intercellular metabolic networks. Trends Microbiol. https://doi.org/10.1016/j.tim.2017.02.015
Panutdaporn N, Kawamoto K, Asakura H, Makino S-I (2006) Resuscitation of the viable but non-culturable state of Salmonella enterica serovar Oranienburg by recombinant resuscitation-promoting factor derived from Salmonella typhimurium strain LT2. Int J Food Microbiol 106:241–247. https://doi.org/10.1016/j.ijfoodmicro.2005.06.022
Pedrós-Alió C, Manrubia S (2016) The vast unknown microbial biosphere. Proc Natl Acad Sci USA 113:6585–6587. https://doi.org/10.1073/pnas.1606105113
Perei K, Rákhely G, Kiss I et al (2001) Biodegradation of sulfanilic acid by Pseudomonas paucimobilis. Appl Microbiol Biotechnol 55:101–107. https://doi.org/10.1007/s002530000474
Pham VHT, Kim J (2012) Cultivation of unculturable soil bacteria. Trends Biotechnol 30:475–484. https://doi.org/10.1016/j.tibtech.2012.05.007
Pinto D, Almeida V, Almeida Santos M, Chambel L (2011) Resuscitation of Escherichia coli VBNC cells depends on a variety of environmental or chemical stimuli. J Appl Microbiol 110:1601–1611. https://doi.org/10.1111/j.1365-2672.2011.05016.x
Pinto D, Santos MA, Chambel L (2015) Thirty years of viable but nonculturable state research: unsolved molecular mechanisms. Crit Rev Microbiol 41:61–76. https://doi.org/10.3109/1040841X.2013.794127
Piqueres P, Moreno Y, Alonso JL, Ferrús MA (2006) A combination of direct viable count and fluorescent in situ hybridization for estimating Helicobacter pylori cell viability. Res Microbiol 157:345–349. https://doi.org/10.1016/j.resmic.2005.09.003
Poindexter JS (1981) Oligotrophy: feast and famine existence. In: Alexander M (ed) Advances in microbial ecology. Plenum Press, New York, pp 63–89
Ramamurthy T, Ghosh A, Pazhani GP, Shinoda S (2014) Current perspectives on viable but non-culturable (VBNC) pathogenic bacteria. Front Public Heal 2:1–9. https://doi.org/10.3389/fpubh.2014.00103
Rappé MS, Giovannoni SJ (2003) The uncultured microbial majority. Annu Rev Microbiol 57:369–394. https://doi.org/10.1146/annurev.micro.57.030502.090759
Rappé MS, Connon SA, Vergin KL, Giovannoni SJ (2002) Cultivation of the ubiquitous SAR11 marine bacterioplankton clade. Nature 418(6898):630–633. https://doi.org/10.1038/nature00917
Rayu S, Karpouzas DG, Singh BK (2012) Emerging technologies in bioremediation: constraints and opportunities. Biodegradation 23:917–926. https://doi.org/10.1007/s10532-012-9576-3
Riesenfeld CS, Schloss PD, Handelsman J (2004) Metagenomics: genomic analysis of microbial communities. Annu Rev Genet 38:525–552. https://doi.org/10.1146/annurev.genet.38.072902.091216
Roszak DB, Grimes DJ, Colwell RR (1984) Viable but nonrecoverable stage of Salmonella enteritidis in aquatic systems. Can J Microbiol 30:334–338. https://doi.org/10.1139/m84-049
Ruggiero A, Squeglia F, Romano M et al (2016) The structure of Resuscitation promoting factor B from M. tuberculosis reveals unexpected ubiquitin-like domains. Biochim Biophys Acta Gen Subj 1860:445–451. https://doi.org/10.1016/j.bbagen.2015.11.001
Rutherford ST, Bassler BL (2012) Bacterial quorum sensing: its role in virulence and possibilities for its control. Cold Spring Harb Perspect Med 2:a012427. https://doi.org/10.1101/cshperspect.a012427
Saha M, Sarkar S, Sarkar B et al (2016) Microbial siderophores and their potential applications: a review. Environ Sci Pollut Res 23:3984–3999. https://doi.org/10.1007/s11356-015-4294-0
Salam LB, Ilori MO, Amund OO et al (2017) Characterization of bacterial community structure in a hydrocarbon-contaminated tropical African soil. Environ Technol. https://doi.org/10.1080/09593330.2017.1317838
Salma M, Rousseaux S, Sequeira-Le Grand A et al (2013) Characterization of the viable but nonculturable (VBNC) state in Saccharomyces cerevisiae. PLoS ONE 8:e77600. https://doi.org/10.1371/journal.pone.0077600
Schloss PD, Handelsman J (2003) Biotechnological prospects from metagenomics. Curr Opin Biotechnol 14:303–310. https://doi.org/10.1016/S0958-1669(03)00067-3
Schmidt TM, Konopka AE (2009) Physiological and ecological adaptations of slow-growing, heterotrophic microbes and consequences for cultivation. Springer, Berlin, pp 257–276
Scullion J (2006) Remediating polluted soils. Naturwissenschaften 93:51–65. https://doi.org/10.1007/s00114-005-0079-5
Serpaggi V, Remize F, Recorbet G et al (2012) Characterization of the “viable but nonculturable” (VBNC) state in the wine spoilage yeast Brettanomyces. Food Microbiol 30:438–447. https://doi.org/10.1016/j.fm.2011.12.020
Shi X, Xie Y, Zhou X (2017) Food-borne Pathogenic Bacteria. In: Jen JJS, Chen J (eds) Food safety in China: science, technology, management and regulation. Wiley, New Jersey, pp 65–82
Shleeva MO, Bagramyan K, Telkov MV et al (2002) Formation and resuscitation of “non-culturable” cells of Rhodococcus rhodochrous and Mycobacterium tuberculosis in prolonged stationary phase. Microbiology 148:1581–1591. https://doi.org/10.1099/00221287-148-5-1581
Shoemaker WR, Lennon JT (2018) Evolution with a seed bank: the population genetic consequences of microbial dormancy. Evol Appl 11:60–75. https://doi.org/10.1111/eva.12557
Singh A, Parmar N, Kuhad RC (2011) Bioaugmentation, biostimulation and biocontrol. Springer, Berlin
Singleton DR, Hu J, Aitken MD (2012) Heterologous expression of polycyclic aromatic hydrocarbon ring-hydroxylating dioxygenase genes from a novel pyrene-degrading betaproteobacterium. Appl Environ Microbiol 78:3552–3559. https://doi.org/10.1128/AEM.00173-12
Smets W, Leff JW, Bradford MA et al (2016) A method for simultaneous measurement of soil bacterial abundances and community composition via 16S rRNA gene sequencing. Soil Biol Biochem 96:145–151. https://doi.org/10.1016/j.soilbio.2016.02.003
Sperandio V, Torres AG, Jarvis B et al (2003) Bacteria-host communication: the language of hormones. Proc Natl Acad Sci USA 100:8951–8956. https://doi.org/10.1073/pnas.1537100100
Sturm A, Dworkin J (2015) Phenotypic diversity as a mechanism to exit cellular dormancy. Curr Biol 25:2272–2277. https://doi.org/10.1016/j.cub.2015.07.018
Su X, Shen X, Ding L, Yokota A (2012) Study on the flocculability of the Arthrobacter sp., an actinomycete resuscitated from the VBNC state. World J Microbiol Biotechnol 28:91–97. https://doi.org/10.1007/s11274-011-0795-2
Su X, Chen X, Hu J et al (2013a) Exploring the potential environmental functions of viable but non-culturable bacteria. World J Microbiol Biotechnol 29:2213–2218. https://doi.org/10.1007/s11274-013-1390-5
Su X, Shen H, Yao X et al (2013b) A novel approach to stimulate the biphenyl-degrading potential of bacterial community from PCBs-contaminated soil of e-waste recycling sites. Bioresour Technol 146:27–34. https://doi.org/10.1016/j.biortech.2013.07.028
Su X, Liu Y, Hu J et al (2014) Optimization of protein production by Micrococcus luteus for exploring pollutant-degrading uncultured bacteria. Springerplus 3:117. https://doi.org/10.1186/2193-1801-3-117
Su X, Liu Y, Hashmi MZ et al (2015a) Rhodococcus biphenylivorans sp. nov., a polychlorinated biphenyl-degrading bacterium. Antonie van Leeuwenhoek Int J Gen Mol Microbiol 107:55–63. https://doi.org/10.1007/s10482-014-0303-4
Su X, Sun F, Wang Y et al (2015b) Identification, characterization and molecular analysis of the viable but nonculturable Rhodococcus biphenylivorans. Sci Rep 5:18590. https://doi.org/10.1038/srep18590
Su X, Zhang Q, Hu J et al (2015c) Enhanced degradation of biphenyl from PCB-contaminated sediments: the impact of extracellular organic matter from Micrococcus luteus. Appl Microbiol Biotechnol 99:1989–2000. https://doi.org/10.1007/s00253-014-6108-6
Su XM, Liu YD, Hashmi MZ et al (2015d) Culture-dependent and culture-independent characterization of potentially functional biphenyl-degrading bacterial community in response to extracellular organic matter from Micrococcus luteus. Microb Biotechnol 8:569–578. https://doi.org/10.1111/1751-7915.12266
Su X, Guo L, Ding L et al (2016) Induction of viable but nonculturable state in Rhodococcus and transcriptome analysis using RNA-seq. PLoS ONE 11:e0147593. https://doi.org/10.1371/journal.pone.0147593
Su X, Wang Y, Xue B et al (2018a) Resuscitation of functional bacterial community for enhancing biodegradation of phenol under high salinity conditions based on Rpf. Bioresour Technol 261:394–402. https://doi.org/10.1016/J.BIORTECH.2018.04.048
Su X, Zhang S, Mei R et al (2018b) Resuscitation of viable but non-culturable bacteria to enhance the cellulose-degrading capability of bacterial community in composting. Microb Biotechnol 11:527–536. https://doi.org/10.1111/1751-7915.13256
Su XM, Bamba AM, Zhang S et al (2018c) Revealing potential functions of VBNC bacteria in polycyclic aromatic hydrocarbons biodegradation. Lett Appl Microbiol 66:277–283. https://doi.org/10.1111/lam.12853
Su X, Li S, Cai J et al (2019a) Aerobic degradation of 3,3′,4,4′-tetrachlorobiphenyl by a resuscitated strain Castellaniella sp. SPC4: kinetics model and pathway for biodegradation. Sci Total Environ 688:917–925. https://doi.org/10.1016/J.SCITOTENV.2019.06.364
Su X, Wang Y, Xue B et al (2019b) Impact of resuscitation promoting factor (Rpf) in membrane bioreactor treating high-saline phenolic wastewater: performance robustness and Rpf-responsive bacterial populations. Chem Eng J 357:715–723. https://doi.org/10.1016/J.CEJ.2018.09.197
Su X, Xue B, Wang Y et al (2019c) Bacterial community shifts evaluation in the sediments of Puyang River and its nitrogen removal capabilities exploration by resuscitation promoting factor. Ecotoxicol Environ Saf 179:188–197. https://doi.org/10.1016/J.ECOENV.2019.04.067
Telkov MV, Demina GR, Voloshin SA et al (2006) Proteins of the Rpf (resuscitation promoting factor) family are peptidoglycan hydrolases. Biochemistry 71:414–422. https://doi.org/10.1134/S0006297906040092
Traxler MF, Seyedsayamdost MR, Clardy J, Kolter R (2012) Interspecies modulation of bacterial development through iron competition and siderophore piracy. Mol Microbiol 86:628–644. https://doi.org/10.1111/mmi.12008
Tripathi V, Edrisi SA, Chen B et al (2017) Biotechnological advances for restoring degraded land for sustainable development land restoration for regaining essential ecosystem services. Trends Biotechnol. https://doi.org/10.1016/j.tibtech.2017.05.001
Tyagi M, da Fonseca MMR, de Carvalho CCCR (2011) Bioaugmentation and biostimulation strategies to improve the effectiveness of bioremediation processes. Biodegradation 22:231–241. https://doi.org/10.1007/s10532-010-9394-4
van Vliet S (2015) Bacterial dormancy: how to decide when to wake up. Curr Biol 25:R753–R755. https://doi.org/10.1016/j.cub.2015.07.039
Vidali M (2001) Bioremediation. An overview*. Pure Appl Chem 73:1163–1172
Villas-Bôas SG, Bruheim P (2007) The potential of metabolomics tools in bioremediation studies. Omi A J Integr Biol 11:305–313. https://doi.org/10.1089/omi.2007.0005
Wandersman C, Delepelaire P (2004) Bacterial iron sources: from siderophores to hemophores. Annu Rev Microbiol 58:611–647. https://doi.org/10.1146/annurev.micro.58.030603.123811
Waters CM, Bassler BL (2006) The Vibrio harveyi quorum-sensing system uses shared regulatory components to discriminate between multiple autoinducers. Genes Dev 20:2754–2767. https://doi.org/10.1101/gad.1466506
Wilson MC, Piel J (2013) Metagenomic approaches for exploiting uncultivated bacteria as a resource for novel biosynthetic enzymology. Chem Biol 20:636–647. https://doi.org/10.1016/j.chembiol.2013.04.011
Xiong M, Hu Z, Zhang Y et al (2013) Survival of GFP-tagged Rhodococcus sp. D310-1 in chlorimuron-ethyl-contaminated soil and its effects on the indigenous microbial community. J Hazard Mater 252–253:347–354. https://doi.org/10.1016/j.jhazmat.2013.02.054
Xu Y, Vetsigian K (2017) Phenotypic variability and community interactions of germinating Streptomyces spores. Sci Rep 7:1–13
Xu H-S, Roberts N, Singleton FL et al (1982) Survival and viability of nonculturableEscherichia coli andVibrio cholerae in the estuarine and marine environment. Microb Ecol 8:313–323. https://doi.org/10.1007/BF02010671
Yu XY, Zhang L, Ren B et al (2015) Arthrobacter liuii sp. nov., resuscitated from Xinjiang desert soil. Int J Syst Evol Microbiol 65:896–901. https://doi.org/10.1099/ijs.0.000037
Zengler K, Toledo G, Rappe M, et al (2002) Cultivating the uncultured. Proc Natl Acad Sci U S A 99:15681–15686. https://doi.org/10.1073/pnas.252630999
Zhang X, Nesme J, Simonet P, Frostegård Å (2012) Fate of invading bacteria in soil and survival of transformants after simulated uptake of transgenes, as evaluated by a model system based on lindane degradation. Res Microbiol 163:200–210. https://doi.org/10.1016/j.resmic.2012.01.007
Zhang D, Berry JP, Zhu D, et al (2015) Magnetic nanoparticle-mediated isolation of functional bacteria in a complex microbial community. ISME J 9:603–614. https://doi.org/10.1038/ismej.2014.161
Zhao S, Song X, Zhao Y et al (2015) Protective and therapeutic effects of the resuscitation-promoting factor domain and its mutants against Mycobacterium tuberculosis in mice. Pathog Dis 73:e31908. https://doi.org/10.1093/femspd/ftu025
Zhao H, Zhang Y, Xiao X et al (2017a) Different phenanthrene-degrading bacteria cultured by in situ soil substrate membrane system and traditional cultivation. Int Biodeterior Biodegrad 117:269–277. https://doi.org/10.1016/j.ibiod.2016.12.016
Zhao X, Zhong J, Wei C et al (2017b) Current perspectives on viable but non-culturable state in foodborne pathogens. Front Microbiol. https://doi.org/10.3389/fmicb.2017.00580
Zou Y, Fu H, Chen Y et al (2014) Viable but non-culturable bacteria in bioreactor-based pharmaceutical wastewater. Agric Sci Technol 15:1299–1303
Acknowledgements
Open access funding provided by University of Szeged (SZTE, University of Szeged Open Access Fund 4587). The project was supported by the European Union and co-financed by the European Social Fund (Grant Agreement No. EFOP-3.6.2-16-2017-00010) in the frame of the Széchényi program of the Hungarian State.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bodor, A., Bounedjoum, N., Vincze, G.E. et al. Challenges of unculturable bacteria: environmental perspectives. Rev Environ Sci Biotechnol 19, 1–22 (2020). https://doi.org/10.1007/s11157-020-09522-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11157-020-09522-4