Abstract
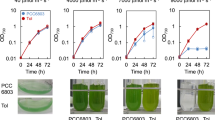
Experimental evolution is a powerful tool for clarifying phenotypic and genotypic changes responsible for adaptive evolution. In this study, we isolated acid-adapted Synechocystis sp. PCC 6803 (Synechocystis 6803) strains to identify genes involved in acid tolerance. Synechocystis 6803 is rarely found in habitants with pH < 5.75. The parent (P) strain was cultured in BG-11 at pH 6.0. We gradually lowered the pH of the medium from pH 6.0 to pH 5.5 over 3 months. Our adapted cells could grow in acid stress conditions at pH 5.5, whereas the parent cells could not. We performed whole-genome sequencing and compared the acid-adapted and P strains, thereby identifying 11 SNPs in the acid-adapted strains, including in Fo F1-ATPase. To determine whether the SNP genes responded to acid stress, we examined gene expression in the adapted strains using quantitative reverse-transcription polymerase chain reaction. sll0914, sll1496, sll0528, and sll1144 expressions increased under acid stress in the P strain, whereas sll0162, sll0163, slr0623, and slr0529 expressions decreased. There were no differences in the SNP genes expression levels between the P strain and two adapted strains, except for sll0528. These results suggest that SNPs in certain genes are involved in acid stress tolerance in Synechocystis 6803.




Similar content being viewed by others
Abbreviations
- QRT-PCR:
-
Real-time quantitative reverse-transcription polymerase chain reaction
- Synechocystis 6803:
-
Synechocystis sp. PCC 6803
References
Amachi S, Ishikawa K, Toyoda S, Kagawa Y, Yokota A et al (1998) Characterization of a mutant of Lactococcus lactis with reduced membrane-bound ATPase activity under acidic conditions. Biosci Biotechnol Biochem 62:1574–1580
Anfelt J, Hallström B, Nielsen J, Uhlén M, Hudson EP (2013) Using transcriptomics to improve butanol tolerance of Synechocystis sp. strain PCC 6803. Appl Environ Microbiol 79:7419–7427
Chen K, Wallis JW, McLellan MD, Larson DE, Kalicki JM, Pohl CS, McGrath SD, Wendl MC, Zhang Q, Locke DP, Shi X, Fulton RS, Ley TJ, Wilson RK, Ding L, Mardis ER (2009) BreakDancer: an algorithm for high-resolution mapping of genomic structural variation. Nat Methods 6:677–681
Chiu SW, Chen SY, Wong HC (2008) Localization and expression of MreB in Vibrio parahaemolyticus under different stresses. Appl Environ Microbiol 74:7016–7022
Cotter PD, Hill C (2003) Surviving the acid test: responses of gram-positive bacteria to low pH. Microbiol Mol Biol Rev 67:429–453
Dettman JR, Rodrigue N, Melnyk AH, Wong A, Bailey SF, Kassen R (2012) Evolutionary insight from whole-genome sequencing of experimentally evolved microbes. Mol Ecol 21:2058–2077
Francius G, Polyakov P, Merlin J, Abe Y, Ghigo JM, Merlin C, Beloin C, Duval JF (2011) Bacterial surface appendages strongly impact nanomechanical and electrokinetic properties of Escherichia coli cells subjected to osmotic stress. PLoS One 6:e20066
Fulda S, Mikkat S, Huang F, Huckauf J, Marin K, Norling B, Hagemann M (2006) Proteome analysis of salt stress response in the cyanobacterium Synechocystis sp. strain PCC 6803. Proteomics 6:2733–2745
Giotis ES, Blair IS, McDowell DA (2007) Morphological changes in Listeria monocytogenes subjected to sublethal alkaline stress. Int J Food Microbiol 120:250–258
Hasan N, Ahmad F, Wu HF (2013) Monitoring the heat stress response of Escherichia coli via NiO nanoparticle assisted MALDI-TOF mass spectrometry. Talanta 103:38–46
Hihara Y, Kamei A, Kanehisa M, Kaplan A, Ikeuchi M (2001) DNA microarray analysis of cyanobacterial gene expression during acclimation to high light. Plant Cell 13:793–806
Hihara Y, Sonoike K, Kanehisa M, Ikeuchi M (2003) DNA microarray analysis of redox-responsive genes in the genome of the cyanobacterium Synechocystis sp. strain PCC 6803. J Bacteriol 185:1719–1725
Hosoya-Matsuda N, Motohashi K, Yoshimura H, Nozaki A, Inoue K, Ohmori M, Hisabori T (2005) Anti-oxidative stress system in cyanobacteria. Significance of type II peroxiredoxin and the role of 1-Cys peroxiredoxin in Synechocystis sp. strain PCC 6803. J Biol Chem 280:840–846
Jan G, Leverrier P, Pichereau V, Boyaval P (2001) Changes in protein synthesis and morphology during acid adaptation of Propionibacterium freudenreichii. Appl Environ Microbiol 67:2029–2036
Kallas T, Castenholz RW (1982) Internal pH and ATP-ADP pools in the cyanobacterium Synechococcus sp. during exposure to growth-inhibiting low pH. J Bacteriol 149:229–236
Kanesaki Y, Suzuki I, Allakhverdiev SI, Mikami K, Murata N (2002) Salt stress and hyperosmotic stress regulate the expression of different sets of genes in Synechocystis sp. PCC 6803. Biochem Biophys Res Commun 290:339–348
Kanesaki Y, Shiwa Y, Tajima N, Suzuki M, Watanabe S, Sato N, Ikeuchi M, Yoshikawa H (2012) Identification of substrain-specific mutations by massively parallel whole-genome resequencing of Synechocystis sp. PCC 6803. DNA Res 19:67–79
Kaniya Y, Kizawa A, Miyagi A, Kawai-Yamada M, Uchimiya H, Kaneko Y, Nishiyama Y, Hihara Y (2013) Deletion of the transcriptional regulator cyAbrB2 deregulates primary carbon metabolism in Synechocystis sp. PCC 6803. 2013. Plant Physiol 162:1153–1163
Kobayashi M, Ishizuka T, Katayama M, Kanehisa M, Bhattacharyya-Pakrasi M, Pakrasi HB, Ikeuchi M (2004) Response to oxidative stress involves a novel peroxiredoxin gene in the unicellular cyanobacterium Synechocystis sp. PCC 6803. Plant Cell Physiol 45:290–299
Kochian LV (1995) Cellular mechanisms of aluminum toxicity and resistance in plants. Annu Rev Plant Physiol Plant Mol Biol 46:237–260
Lee DH, Palsson BØ (2010) Adaptive evolution of Escherichia coli K-12 MG1655 during growth on a nonnative carbon source, L-1,2-propanediol. Appl Environ Microbiol 76:4158–4168
Li H, Durbin R (2010) Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26:589–595
Li H, Ruan J, Durbin R (2008) Mapping short DNA sequencing reads and calling variants using mapping quality scores. Genome Res 18:1851–1858
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R (2009) 1000 Genome project data processing subgroup. Bioinformatics 25:2078–2079
Li Y, Rao N, Yang F, Zhang Y, Yang Y, Liu HM, Guo F, Huang J (2014) Biocomputional construction of a gene network under acid stress in Synechocystis sp. PCC 6803. Res Microbiol 165:420–428
Marbouty M, Mazouni K, Saguez C, Cassier-Chauvat C, Chauvat F (2009) Characterization of the Synechocystis strain PCC 6803 penicillin-binding proteins and cytokinetic proteins FtsQ and FtsW and their network of interactions with ZipN. J Bacteriol 191:5123–5133
Miseon B, Sangnam O, Kwang-Sei L, Younghoon K, Sejong O (2014) The involvement of ATPase activity in the acid tolerance of Lactobacillus rhamnosus strain GG. Int J Dairy Tech 67:229–236
Naranjo MA, Forment J, Roldán M, Serrano R, Vicente O (2006) Overexpression of Arabidopsis thaliana LTL1, a salt-induced gene encoding a GDSL-motif lipase, increases salt tolerance in yeast and transgenic plants. Plant Cell Environ 29:1890–1900
Nitta K, Nagayama K, Danev R, Kaneko Y (2009) Visualization of BrdU-labelled DNA in cyanobacterial cells by Hilbert differential contrast transmission electron microscopy. J Microsc 234:118–123
Ohta H, Shibata Y, Haseyama Y, Yoshino Y, Suzuki T, Kagasawa T, Kamei A, Ikeuchi M, Enami I (2005) Identification of genes expressed in response to acid stress in Synechocystis sp. PCC 6803 using DNA microarrays. Photosynth Res 84:225–230
Oud B, van Maris AJ, Daran JM, Pronk JT (2012) Genome-wide analytical approaches for reverse metabolic engineering of industrially relevant phenotypes in yeast. FEMS Yeast Res 12:183–196
Pianetti A, Battistelli M, Citterio B, Parlani C, Falcieri E, Bruscolini F (2009) Morphological changes of Aeromonas hydrophila in response to osmotic stress. Micron 40:426–433
Qiu D, Lin X (2002) Effects of simulated acid rain on chloroplast activity in Dimorcarpus longana Lour. cv. wulongling leaves. Ying Yong Sheng Tai Xue Bao 13:1559–1562
Sakiyama T, Ueno H, Homma H, Numata O, Kuwabara T (2006) Purification and characterization of a hemolysin-like protein, Sll1951, a nontoxic member of the RTX protein family from the Cyanobacterium Synechocystis sp. strain PCC 6803. J Bacteriol 188:3535–3542
Sakiyama T, Araie H, Suzuki I, Shiraiwa Y (2011) Functions of a hemolysin-like protein in the cyanobacterium Synechocystis sp PCC 6803. Arch Microbiol 193:565–571
Sambe M, Kitayama S, Moriyama A, Uchiyama J, Ohta H (2013) The role of sll1558 and sll1496 genes under acid stress conditions in Cyanobacterium Synechocystis sp. PCC6803. Photosynth Res Food Fuel Future 429:579–582
Sánchez-Riego AM, López-Maury L, Florencio FJ (2014) Genomic responses to arsenic in the cyanobacterium Synechocystis sp. PCC 6803. PLoS One 5:e96826
Santos JM, Freire P, Vicente M, Arraiano CM (1999) The stationary-phase morphogene bolA from Escherichia coli is induced by stress during early stages of growth. Mol Microbiol 32:789–798
Shapiguzov A, Lyukevich AA, Allakhverdiev SI, Sergeyenko TV, Suzuki I, Murata N, Los D (2005) Osmotic shrinkage of cells of Synechocystis sp. PCC 6803 by water efflux via aquaporins regulates osmostress-inducible gene expression. Microbiology 151:447–455
Shiwa Y, Matsumoto T, Yoshikawa H (2013) Identification of laboratory-specific variations of Bacillus subtilis strains used in Japan. Biosci Biotechnol Biochem 77:2073–2076
Stanier RY, Kunisawa R, Mandel M, Cohen-Bazire G (1971) Purification and properties of unicellular blue-green algae (order Chroococcales). Bacteriol Rev 35:171–205
Summerfield TC, Sherman LA (2008) Global transcriptional response of the alkali-tolerant cyanobacterium Synechocystis sp. strain PCC 6803 to a pH10 environment. Appl Environ Microbiol 74:5276–5284
Tahara H, Uchiyama J, Yoshihara T, Matsumoto K, Ohta H (2012) Role of Slr1045 in environmental stress tolerance and lipid transport in the cyanobacterium Synechocystis sp. PCC6803. Biochim Biophys Acta 1817:1360–1366
Trautner C, Vermaas WF (2013) The sll1951 gene encodes the surface layer protein of Synechocystis sp. strain PCC 6803. J Bacteriol 195:5370–5380
Uchiyama J, Asakura R, Kimura M, Moriyama A, Tahara H, Kobayashi Y, Kubo Y, Yoshihara T, Ohta H (2012) Slr0967 and Sll0939 induced by the SphR response regulator in Synechocystis sp. PCC 6803 are essential for growth under acid stress conditions. Biochim Biophys Acta 1817:1270–1276
Uchiyama J, Asakura R, Moriyama A, Kubo Y, Shibata Y, Yoshino Y, Tahara H, Matsuhashi A, Sato S, Nakamura Y, Tabata S, Ohta H (2014) Sll0939 is induced by Slr0967 in the cyanobacterium Synechocystis sp. PCC6803 and is essential for growth under various stress conditions. Plant Physiol Biochem 81:36–43
Wang C, Xing D, Zeng L, Ding C, Chen Q (2005) Effect of artificial acid rain and SO2 on characteristics of delayed light emission. Luminescence 20:51–56
Wong A, Rodrigue N, Kassen R (2012) Genomics of adaptation during experimental evolution of the opportunistic pathogen Pseudomonas aeruginosa. PLoS Genet 8:e1002928
Yamauchi Y, Kaniya Y, Kaneko Y, Hihara Y (2011) Physiological roles of the cyAbrB transcriptional regulator pair Sll0822 and Sll0359 in Synechocystis sp. strain PCC 6803. J Bacteriol 193:3702–3709
Zeng QL, Huang XH, Zhou Q (2005) Effect of acid rain on seed germination of rice, wheat and rape. Huan Jing Ke Xue 26:181–184
Zhang X, Chen G, Qin C, Wang Y, Wei D (2012) Slr0643, an S2P homologue, is essential for acid acclimation in the cyanobacterium Synechocystis sp. PCC 6803. Microbiology 158:2765–2780
Zhou Q, Huang X, Liu X (2002) Stress effects of simulant acid rain on three woody plants. Huan Jing Ke Xue 23:42–46
Acknowledgments
This study was supported by the Program for Development of Strategic Research Center in Private Universities, which was supported by MEXT (S1001020). This study was supported by MEXT-Supported Program for the Strategic Research Foundation at Private Universities, 2013–2017 (S1311017). The authors would like to thank Enago (www.enago.jp) for the English language review.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Uchiyama, J., Kanesaki, Y., Iwata, N. et al. Genomic analysis of parallel-evolved cyanobacterium Synechocystis sp. PCC 6803 under acid stress. Photosynth Res 125, 243–254 (2015). https://doi.org/10.1007/s11120-015-0111-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11120-015-0111-3