Abstract
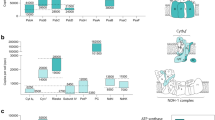
We developed here the quantitative and objective method to analyze chlorophyll fluorescence from the cyanobacterium Synechocystis sp. PCC 6803 in the aim of systematic examination of gene function. The overall similarity of the chlorophyll fluorescence induction kinetics was evaluated for 499 mutants. Mutants of 333 genes showed the difference in the fluorescence kinetics from that of wild type, indicating the wide interaction of photosynthesis with other metabolisms. Hierarchical clustering of the similarity of the mutants enables us to group together the mutants having defect in the regulation of photosystem stoichiometry as well as those having defects in respiration or other functions. Furthermore, wild-type cells treated with inhibitors of respiration and mutants of genes involved in respiration shared similar induction kinetics. Apparently, quantitative comparison of the induction kinetics could be useful to analyze the function of genes as well as to predict the target sites of various chemicals.






Similar content being viewed by others
Abbreviations
- PCC:
-
Pasteur culture collection
- CCD:
-
Charge-coupled device
- TES:
-
N-tris(hydroxymethyl)methyl-2-aminoethanesulfonic acid
References
Alonso JM, Stepanova AN, Leisse TJ, Kim CJ, Chen H, Shinn P, Stevenson DK, Zimmerman J, Barajas P, Cheuk R, Gadrinab C, Heller C, Jeske A, Koesema E, Meyers CC, Parker H, Prednis L, Ansari Y, Choy N, Deen H, Geralt M, Hazari N, Hom E, Karnes M, Mulholland C, Ndubaku R, Schmidt I, Guzman P, Aguilar-Henonin L, Schmid M, Weigel D, Carter DE, Marchand T, Risseeuw E, Brogden D, Zeko A, Crosby WL, Berry CC, Ecker JR (2003) Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301:653–657
Baker NR (2008) Chlorophyll fluorescence: a probe of photosynthesis in vivo. Annu Rev Plant Biol 59:89–113
Fujimori T, Higuchi M, Sato H, Aiba H, Muramatsu M, Hihara Y, Sonoike K (2005) The mutant of sll1961, which encodes a putative transcriptional regulator, has a defect in regulation of photosystem stoichiometry in the cyanobacterium Synechocystis sp. PCC 6803. Plant Physiol 139:408–416
Genty B, Briantais JM, Baker NR (1989) The relationship between the quantum yield of photosynthetic electron transport and quenching of chlorophyll fluorescence. Biochim Biophys Acta 990:87–92
Govindjee (1995) Sixty-three years since kautsky: chlorophyll a fluorescence. Aust J Plant Physiol 22:131–160
Hashimoto M, Endo T, Peltier G, Tasaka M, Shikanai T (2003) A nucleus-encoded factor, CRR2, is essential for the expression of chloroplast ndhB in Arabidopsis. Plant J 36:541–549
Hihara Y, Sonoike K, Ikeuchi M (1998) A novel gene, pmgA, specifically regulates photosystem stoichiometry in the cyanobacterium Synechocystis species PCC 6803 in response to high light. Plant Physiol 117:1205–1216
Hihara Y, Kamei A, Kanehisa M, Kaplan A, Ikeuchi M (2001) DNA microarray analysis of cyanobacterial gene expression during acclimation to high light. Plant Cell 13:793–806
Ihaka R, Gentleman R (1996) R: a language for data analysis and graphics. J Comput Graph Stat 5:299–314
Kamath RS, Fraser AG, Dong Y, Poulin G, Durbin R, Gotta M, Kanapin A, Le Bot N, Moreno S, Sohrmann M, Welchman DP, Zipperlen P, Ahringer J (2003) Systematic functional analysis of the Caenorhabditis elegans genome using RNAi. Nature 421:231–237
Kitajima M, Butler WL (1975) Quenching of chlorophyll fluorescence and primary photochemistry in chloroplasts by dibromothymoquinone. Biochim Biophys Acta 376:105–115
Kuromori T, Wada T, Kamiya A, Yuguchi M, Yokouchi T, Imura Y, Takabe H, Sakurai T, Akiyama K, Hirayama T, Okada K, Shinozaki K (2006) A trial of phenome analysis using 4000 Ds-insertional mutants in gene-coding regions of Arabidopsis. Plant J 47:640–651
Li XP, Bjorkman O, Shih C, Grossman AR, Rosenquist M, Jansson S, Niyogi KK (2000) A pigment-binding protein essential for regulation of photosynthetic light harvesting. Nature 403:391–395
Los DA, Suzuki I, Zinchenko VV, Murata N (2007) Stress responses in Synechocystis: regulated genes and regulatory systems. In: Herrero A, Flores E (eds) Cyanobacteria: molecular biology, genomics and evolution. Horizon Scientific Press, Norfolk, pp 117–157
Marin K, Kanesaki Y, Los DA, Murata N, Suzuki I, Hagemann M (2004) Gene expression profiling reflects physiological processes in salt acclimation of Synechocystis sp. strain PCC 6803. Plant Physiol 136:3290–3300
Munshi MK, Kobayashi Y, Shikanai T (2005) Identification of a novel protein, CRR7, required for the stabilization of the chloroplast NAD(P)H dehydrogenase complex in Arabidopsis. Plant J 44:1036–1044
Munshi MK, Kobayashi Y, Shikanai T (2006) Chlororespiratory reduction 6 is a novel factor required for accumulation of the chloroplast NAD(P)H dehydrogenase complex in Arabidopsis. Plant Physiol 141:737–744
Niyogi KK, Bjorkman O, Grossman AR (1997) Chlamydomonas xanthophyll cycle mutants identified by video imaging of chlorophyll fluorescence quenching. Plant Cell 9:1369–1380
Niyogi KK, Grossman AR, Bjorkman O (1998) Arabidopsis mutants define a central role for the xanthophyll cycle in the regulation of photosynthetic energy conversion. Plant Cell 10:1121–1134
Oxborough K (2004) Imaging of chlorophyll a fluorescence: theoretical and practical aspects of an emerging technique for the monitoring of photosynthetic performance. J Exp Bot 55:1195–1205
Ozaki H, Sonoike K (2008) Quantitative analysis of chlorophyll fluorescence kinetics of the cyanobacterium Synechocystis sp. PCC 6803. In: Allen JF, Grantt E, Golbeck JH, Osmond B (eds) Photosynthesis. Energy from the Sun: 14th International Congress on Photosynthesis. Springer, New York, pp 1571–1574
Ozaki H, Ikeuchi M, Ogawa T, Fukuzawa H, Sonoike K (2007) Large-scale analysis of chlorophyll fluorescence kinetics in Synechocystis sp. PCC 6803: identification of the factors involved in the modulation of photosystem stoichiometry. Plant Cell Physiol 48:451–458
Rippka R, Deruelles J, Waterbury JB, Herdman M, Stanier RY (1979) Generic assignments, strain histories and properties of pure cultures of cyanobacteria. J Gen Microbiol 111:1–61
Ross-Macdonald P, Coelho PS, Roemer T, Agarwal S, Kumar A, Jansen R, Cheung KH, Sheehan A, Symoniatis D, Umansky L, Heidtman M, Nelson FK, Iwasaki H, Hager K, Gerstein M, Miller P, Roeder GS, Snyder M (1999) Large-scale analysis of the yeast genome by transposon tagging and gene disruption. Nature 402:413–418
Scherens B, Goffeau A (2004) The uses of genome-wide yeast mutant collections. Genome Biol 5:229
Schreiber U, Bilger W, Neubauer C (1994) Chlorophyll fluorescence as a nonintrusive indicator for rapid assessment of in vivo photosynthesis. In: Schulze CD, Caldwell MM (eds) Ecophysiology of photosynthesis. Springer, Berlin, pp 45–70
Strasser RJ, Srivastava A, Tsimilli-Michael M (2000) The fluorescence transient as a tool to characterize and screen photosynthetic samples. In: Yunus M, Pathre U, Mohanty P (eds) Probing photosynthesis. Taylor & Francis, London, pp 445–483
Winzeler EA, Shoemaker DD, Astromoff A, Liang H, Anderson K, Andre B, Bangham R, Benito R, Boeke JD, Bussey H, Chu AM, Connelly C, Davis K, Dietrich F, Dow SW, El Bakkoury M, Foury F, Friend SH, Gentalen E, Giaever G, Hegemann JH, Jones T, Laub M, Liao H, Liebundguth N, Lockhart DJ, Lucau-Danila A, Lussier M, M’Rabet N, Menard P, Mittmann M, Pai C, Rebischung C, Revuelta JL, Riles L, Roberts CJ, Ross-MacDonald P, Scherens B, Snyder M, Sookhai-Mahadeo S, Storms RK, Véronneau S, Voet M, Volckaert G, Ward TR, Wysocki R, Yen GS, Yu K, Zimmermann K, Philippsen P, Johnston M, Davis RW (1999) Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285:901–906
Acknowledgements
The authors thank to the staff of Kazusa DNA Research Institute for helpful discussion. This work was supported by Grant-in-Aid for Genome Biology from the Ministry of Education, Culture, Sports, Science and Technology to K.S.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ozaki, H., Sonoike, K. Quantitative analysis of the relationship between induction kinetics of chlorophyll fluorescence and function of genes in the cyanobacterium Synechocystis sp. PCC 6803. Photosynth Res 101, 47–58 (2009). https://doi.org/10.1007/s11120-009-9462-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11120-009-9462-y