Abstract
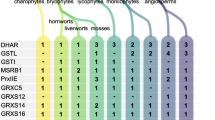
The availability of the Arabidopsis genome revealed the complexity of the gene families implicated in dithiol disulfide exchanges. Most non-green organisms present less dithiol oxidoreductase genes. The availability of the almost complete genome sequence of rice now allows a systematic search for thioredoxins, glutaredoxins and their reducers. This shows that all redoxin families previously defined for Arabidopsis have members in the rice genome and that all the deduced rice redoxins fall within these families. This establishes that the redoxin classification applies both to dicots and monocots. Nevertheless, within each redoxin type the number of members is not the same in these two higher plants and it is not always possible to define orthologues between rice and Arabidopsis. The sequencing of two unicellular algae (Chlamydomonas and Ostreococcus) genomes are almost finished. This allowed us to follow the origin of the different gene families in the green lineage. It appears that most thioredoxin and glutaredoxin types, their chloroplastic, mitochondrial and cytosolic reducers are always present in these unicellular organisms. Nevertheless, striking differences appear in comparison to higher plant redoxins. Some thioredoxin types are not present in these algal genomes including thioredoxins o, clot and glutaredoxins CCxC. Numerous redoxins, including the cytosolic thioredoxins, do not fit with the corresponding higher plant classification. In addition both algae present a NADPH-dependent thioredoxin reductase with a selenocysteine which is highly similar to the animal thioredoxin reductases, a type of thioredoxin reductase not present in higher plants.







Similar content being viewed by others
Change history
30 October 2007
An Erratum to this paper has been published: https://doi.org/10.1007/s11120-007-9273-y
References
Bick JA, Setterdahl AT, Knaff DB, Chen Y, Pitcher LH, Zilinskas BA, Leustek T (2001) Regulation of the plant-type 5’-adenylyl sulfate reductase by oxidative stress. Biochemistry 40:9040–9048
Brehelin C, Meyer EH, de Souris J, Bonnard G, Meyer Y (2003) Resemblance and dissemblance of arabidopsis type ii peroxiredoxins: similar sequences for divergent gene expression, protein localization, and activity. Plant Physiol 132:2045–2057
Broin M, Rey P (2003) Potato plants lacking the cdsp32 plastidic thioredoxin exhibit overoxidation of the bas1 2-cysteine peroxiredoxin and increased lipid peroxidation in thylakoids under photooxidative stress. Plant Physiol 132:1335–1343
Collin V, Issakidis-Bourguet E, Marchand C, Hirasawa M, Lancelin J, Knaff DB, Miginiac-Maslow M (2003) The arabidopsis plastidial thioredoxins: new functions and new insights into specificity. J Biol Chem 278:23747–23752
Collin V, Lamkemeyer P, Miginiac-Maslow M, Hirasawa M, Knaff DB, Dietz K, Issakidis-Bourguet E (2004) Characterization of plastidial thioredoxins from arabidopsis belonging to the new y-type. Plant Physiol 136:4088–4095
Creissen G, Reynolds H, Xue Y, Mullineaux P (1995) Simultaneous targeting of pea glutathione reductase and of a bacterial fusion protein to chloroplasts and mitochondria in transgenic tobacco. Plant J 8:167–175
Dai S, Schwendtmayer C, Schurmann P, Ramaswamy S, Eklund H (2000) Redox signaling in chloroplasts: cleavage of disulfides by an iron–sulfur cluster. Science 287:655–658
Gelhaye E, Rouhier N, Jacquot J (2004) The thioredoxin h system of higher plants. Plant Physiol Biochem 42:265–271
Giordano E, Peluso I, Rendina R, Digilio A, Furia M (2003) The clot gene of Drosophila melanogaster encodes a conserved member of the thioredoxin-like protein superfamily. Mol Genet Genomics 268:692–697
Gonnet GH, Hallett MT, Korostensky C, Bernardin L (2000) Darwin v. 2.0: an interpreted computer language for the biosciences. Bioinformatics 16:101–103
Hartman H, Syvanen M, Buchanan BB (1990) Contrasting evolutionary histories of chloroplast thioredoxins f and m. Mol Biol Evol 7:247–254
Isakov N, Witte S, Altman A (2000) Picot-hd: a highly conserved protein domain that is often associated with thioredoxin and glutaredoxin modules. Trends Biochem Sci 25:537–539
Laloi C, Rayapuram N, Chartier Y, Grienenberger JM, Bonnard G, Meyer Y (2001) Identification and characterization of a mitochondrial thioredoxin system in plants. Proc Natl Acad Sci USA 98:14144–14149
Laughner BJ, Sehnke PC, Ferl RJ (1998) A novel nuclear member of the thioredoxin superfamily. Plant Physiol 118:987–996
Lemaire SD, Collin V, Keryer E, Quesada A, Miginiac-Maslow M (2003) Characterization of thioredoxin y, a new type of thioredoxin identified in the genome of Chlamydomonas reinhardtii. FEBS Lett 543:87–92
Lemaire S (2004) The glutaredoxin family in oxygenic photosynthetic organisms. Photosynth Res 79:305–318
Lemaire S, Miginiac-Maslow M (2004) The thioredoxin superfamily in Chlamydomonas reinhardtii. Photosynth Res 82:203–220
Lemaire SD, Quesada A, Merchan F, Corral JM, Igeno MI, Keryer E, Issakidis-Bourguet E, Hirasawa M, Knaff DB, Miginiac-Maslow M (2005) NADP-malate dehydrogenase from unicellular green alga Chlamydomonas reinhardtii. A first step toward redox regulation?. Plant Physiol 137:514–521
Lennartz K, Plucken H, Seidler A, Westhoff P, Bechtold N, Meierhoff K (2001) Hcf164 encodes a thioredoxin-like protein involved in the biogenesis of the cytochrome b(6)f complex in arabidopsis. Plant Cell 13:2539–2551
Michelet L, Zaffagnini M, Marchand C, Collin V, Decottignies P, Tsan P, Lancelin JM, Trost P, Miginiac-Maslow M, Noctor G, Lemaire SD (2005) Glutathionylation of chloroplast thioredoxin f is a redox signaling mechanism in plants. Proc Natl Acad Sci USA 102(45):16478–16483
Miranda-Vizuete A, Spyrou G (2002) Genomic organization and identification of a novel alternative splicing variant of mouse mitochondrial thioredoxin reductase (trxr2) gene. Mol Cells 13:488–492
Meyer Y, Vignols F, Reichheld JP (2002) Classification of plant thioredoxins by sequence similarity and intron position. Methods Enzymol 347:394–402
Novoselov SV, Rao M, Onoshko NV, Zhi H, Kryukov GV, Xiang Y, Weeks DP, Hatfield DL, Gladyshev VN (2002) Selenoproteins and selenocysteine insertion system in the model plant cell system, Chlamydomonas reinhardtii. EMBO J 21:3681–3693
Rey P, Cuine S, Eymery F, Garin J, Court M, Jacquot J, Rouhier N, Broin M (2005) Analysis of the proteins targeted by cdsp32, a plastidic thioredoxin participating in oxidative stress responses. Plant J 41:31–42
Rivas S, Rougon-Cardoso A, Smoker M, Schauser L, Yoshioka H, Jones JDG (2004) Citrx thioredoxin interacts with the tomato cf-9 resistance protein and negatively regulates defence. EMBO J 23:2156–2165
Reichheld J, Meyer E, Khafif M, Bonnard G, Meyer Y (2005) Atntrb is the major mitochondrial thioredoxin reductase in arabidopsis thaliana. FEBS Lett 579:337–342
Rouhier N, Gelhaye E, Jacquot JP (2002) Glutaredoxin-dependent peroxiredoxin from poplar: protein-protein interaction and catalytic mechanism. J Biol Chem 277:13609–13614
Rouhier N, Vlamis-Gardikas A, Lillig CH, Berndt C, Schwenn J, Holmgren A, Jacquot J (2003) Characterization of the redox properties of poplar glutaredoxin. Antioxid Redox Signal 5:15–22
Rouhier N, Gelhaye E, Jacquot J (2004) Plant glutaredoxins: still mysterious reducing systems. Cell Mol Life Sci 61:1266–1277
Rouhier N, Villarejo A, Srivastava M, Gelhaye E, Keech O, Droux M, Finkemeier I, Samuelsson G, Dietz KJ, Jacquot J, Wingsle G (2005) Identification of plant glutaredoxin targets. Antioxid Redox Signal 7:919–929
Rouhier N, Couturier J, Jacquot J (2006) Genome-wide analysis of plant glutaredoxin systems. J Exp Bot
Sahrawy M, Hecht V, Lopez-Jaramillo J, Chueca A, Chartier Y, Meyer Y (1996) Intron position as an evolutionary marker of thioredoxins and thioredoxin domains. J Mol Evol 42:422–431
Sarkar N, Lemaire S, Wu-Scharf D, Issakidis-Bourguet E, Cerutti H (2005) Functional specialization of Chlamydomonas reinhardtii cytosolic thioredoxin h1 in the response to alkylation-induced DNA damage. Eukaryot Cell 4:262–273
Schürmann P (2002) Ferredoxin-dependent thioredoxin reductase: a unique iron–sulfur protein. Methods Enzymol 347:403–411
Serrato AJ, Perez-Ruiz JM, Spinola MC, Cejudo FJ (2004) A novel nadph thioredoxin reductase, localized in the chloroplast, which deficiency causes hypersensitivity to abiotic stress in arabidopsis thaliana. J Biol Chem 279:43821–43827
Stork T, Michel K, Pistorius EK, Dietz K (2005) Bioinformatic analysis of the genomes of the cyanobacteria synechocystis sp. pcc 6803 and synechococcus elongatus pcc 7942 for the presence of peroxiredoxins and their transcript regulation under stress. J Exp Bot 56:3193–3206
Su D, Novoselov SV, Sun Q, Moustafa ME, Zhou Y, Oko R, Hatfield DL, Gladyshev VN (2005) Mammalian selenoprotein thioredoxin-glutathione reductase roles in disulfide bond formation and sperm maturation. J Biol Chem 280:26491–26498
Vignols F, Mouaheb N, Thomas D, Meyer Y (2003) Redox control of hsp70-co-chaperone interaction revealed by expression of a thioredoxin-like arabidopsis protein. J Biol Chem 278:4516–4523
Xing S, Rosso MG, Zachgo S (2005) Roxy1, a member of the plant glutaredoxin family, is required for petal development in arabidopsis thaliana. Development 132:1555–1565
Acknowledgments
We would like to thank Stephane Lemaire (Orsay) for interesting discussions about redoxin genes in Chlamydomonas and Richard Cooke (Perpignan) both for discussions on comparative genomics and for correcting this manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Meyer, Y., Riondet, C., Constans, L. et al. Evolution of redoxin genes in the green lineage. Photosynth Res 89, 179–192 (2006). https://doi.org/10.1007/s11120-006-9095-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11120-006-9095-3