Abstract
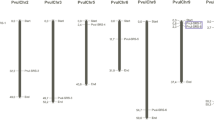
The SABATH gene family is a plant-specific class of small molecule methyltransferases. The first goal of this study was to conduct a genome-wide analysis and in silico characterization of SABATH gene family members in Phaseolus vulgaris. The second goal was to see how the members of the SABATH gene family react to salt and drought stress, as well as their expression patterns when melatonin is added to the mix. Within the scope of the study, 18 Pvul-SABATH proteins were discovered in the Phaseolus vulgaris genome using various in silico methods. These proteins ranged in size from 30.18 to 42.73 kDa and were made up of 268 to 387 amino acids. The Pvul-SABATH proteins had isoelectric points ranging from 4.95 (Pvul-SABATH-8) to 6.08 (Pvul-SABATH-2 and -3). The Pvul-SABATH genes were found to have at least three but no more than four exons. According to the results of the phylogenetic analysis, Pvul-SABATH proteins were found to be clustered in three major groups with Arabidopsis thaliana, Glycine max species, and various methyltransferase members. There was segmental duplication between the Pvul-SABATH-1 and Pvul-SABATH-5 genes, as well as the Pvul-SABATH-10 and Pvul-SABATH-14 genes. By comparing the expression level of Pvul-SABATH genes, their different expression levels depending on the bean cultivars were discovered. Moreover, these findings suggested that Pvul-SABATH genes may play a role in the growth and development process of the plants in response to various biotic and abiotic stresses. The findings of this study, which is the first to look at the SABATH gene family in Phaseolus vulgaris, are thought to be beneficial in plant biotechnology and molecular biology.













Similar content being viewed by others
References
Arnao MB, Hernandez-Ruiz J (2009) Protective effect of melatonin against chlorophyll degradation during the senescence of barley leaves. J Pineal Res 46(1):58–63. https://doi.org/10.1111/j.1600-079X.2008.00625.x
Arnao, MB, Hernández‐Ruiz J (2013) Growth conditions determine different melatonin levels in L upinus albus L. J Pineal Res 55(2):149–155. https://doi.org/10.1111/jpi.12055
Attieh J, Djiana R, Koonjul P, Etienne C, Sparace SA, Saini HS (2002) Cloning and functional expression of two plant thiol methyltransferases: a new class of enzymes involved in the biosynthesis of sulfur volatiles. Plant Mol Biol 50(3):511–521. https://doi.org/10.1023/a:1019865829534
Bailey TL, Williams N, Misleh C, Li WW (2006) MEME: discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res 34:W369–W373. https://doi.org/10.1093/nar/gkl198
Broughton WJ, Hernandez G, Blair M, Beebe S, Gepts P, Vanderleyden J (2003) Beans (Phaseolus spp.) – model food legumes. Plant Soil 252(1):55–128.
Büyük I, Aras S (2017) Genome-wide in silico identification, characterization and transcriptional analysis of the family of growth-regulating factors in common bean (Phaseolus vulgaris l.) subjected to polyethylene glycol-induced drought stress. Arch Biol Sci 69(1):5–14.
Chaiprasongsuk M, Zhang C, Qian P, Chen XL, Li GL, Trigiano RN, Guo H, Chen F (2018) Biochemical characterization in Norway spruce (Picea abies) of SABATH methyltransferases that methylate phytohormones. Phytochemistry 149:146–154. https://doi.org/10.1016/j.phytochem.2018.02.010
Chen CJ, Chen H, Zhang Y, Thomas HR, Frank MH, He YH, Xia R (2020) TBtools: an ıntegrative toolkit developed for ınteractive analyses of big biological data. Mol Plant 13(8):1194–1202. https://doi.org/10.1016/j.molp.2020.06.009
Conesa A, Gotz S, Garcia-Gomez JM, Terol J, Talon M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21(18):3674–3676. https://doi.org/10.1093/bioinformatics/bti610
Crooks GE, Hon G, Chandonia JM, Brenner SE (2004) WebLogo: a sequence logo generator. Genome Res 14(6):1188–1190. https://doi.org/10.1101/gr.849004
D Auria JC, Chen F, Pichersky E (2003) The SABATH family of MTs in Arabidopsis thaliana and other plant species. Recent Adv Phytochem 37:253–284
Demsar J et al (2013) Orange: Data Mining Toolbox in Python. J Mach Learn Res 14:2349–2353
FAO (2020) FAOSTAT Statistical Database. In. http://www.fao.org/faostat/en/#data
Guo Y, Qiao DH, Yang C, Chen J, Li Y, Liang SH, Lin KQ, Chen ZW (2020) Genome-wide identification and expression analysis of SABATH methyltransferases in tea plant (Camellia sinensis): insights into their roles in plant defense responses. Plant Signal Behav 15(10). https://doi.org/10.1080/15592324.2020.1804684
Han XM, Yang Q, Liu YJ, Yang ZL, Wang XR, Zeng QY, Yang HL (2018) Evolution and function of the Populus SABATH family reveal that a single amino acid change results in a substrate switch. Plant Cell Physiol 59(2):392–403. https://doi.org/10.1093/pcp/pcx198
Hiz MC, Canher B, Niron H, Turet M (2014) Transcriptome analysis of salt tolerant common bean (Phaseolus vulgaris L.) under saline conditions. Plos One 9(3). https://doi.org/10.1371/journal.pone.0092598
Horton P, Park KJ, Obayashi T, Fujita N, Harada H, Adams-Collier CJ, Nakai K (2007) WoLF PSORT: protein localization predictor. Nucleic Acids Res 35:W585–W587. https://doi.org/10.1093/nar/gkm259
Hu B, Jin JP, Guo AY, Zhang H, Luo JC, Gao G (2015) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31(8):1296–1297. https://doi.org/10.1093/bioinformatics/btu817
Huangfu L et al (2020) Exogenous melatonin promotes rice seed germination under salinity through regulating antioxidants and metabolic homeostasis. https://doi.org/10.21203/rs.2.21896/v1
Juretic N, Hoen DR, Huynh ML, Harrison PM, Bureau TE (2005) The evolutionary fate of MULE-mediated duplications of host gene fragments in rice. Genome Res 15(9):1292–1297. https://doi.org/10.1101/gr.4064205
Kapteyn J, Qualley AV, Xie ZZ, Fridman E, Dudareva N, Gang DR (2007) Evolution of cinnamate/p-coumarate carboxyl methyltransferases and their role in the biosynthesis of methylcinnamate. Plant Cell 19(10):3212–3229. https://doi.org/10.1105/tpc.107.054155
Kelley LA, Mezulis S, Yates CM, Wass MN, Sternberg MJE (2015) The Phyre2 web portal for protein modeling, prediction and analysis. Nat Protoc 10(6):845–858. https://doi.org/10.1038/nprot.2015.053
Kollner TG, Lenk C, Zhao N, Seidl-Adams I, Gershenzon J, Chen F, Degenhardt J (2010) Herbivore-ınduced SABATH methyltransferases of maize that methylate anthranilic acid using S-adenosyl-L-methionine. Plant Physiol 153(4):1795–1807. https://doi.org/10.1104/pp.110.158360
Lamesch P et al (2012) The Arabidopsis Information Resource (TAIR): improved gene annotation and new tools. Nucleic Acids Res 40(D1):D1202–D1210. https://doi.org/10.1093/nar/gkr1090
Lescot M, Dehais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouze P, Rombauts S (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30(1):325–327. https://doi.org/10.1093/nar/30.1.325
Letunic I, Bork P (2011) Interactive Tree Of Life v2: online annotation and display of phylogenetic trees made easy. Nucleic Acids Res 39:W475–W478. https://doi.org/10.1093/nar/gkr201
Li J, Liu J, Zhu T, Zhao C, Li L, Chen M (2019) The role of melatonin in salt stress responses. Int J Mol Sci 20(7):1735. https://doi.org/10.3390/ijms20071735
Li C, Wang P, Wei Z, Liang D, Liu C, Yin L, Jia D, Fu M, Ma F (2012) The mitigation effects of exogenous melatonin on salinity-induced stress in Malus hupehensis. J Pineal Res 53: 298-306. https://doi.org/10.1111/j.1600-079X.2012.00999.x
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(T)(-Delta Delta C) method. Methods 25(4):402–408. https://doi.org/10.1006/meth.2001.1262
Lynch M, Conery JS (2003) The evolutionary demography of duplicate genes. Genome Evolution:35–44.
Michael S, Stéphanie R, Jürgen K-V (2013) Auxin: simply complicated. J Exp Bot 64(9):2565–2577. https://doi.org/10.1093/jxb/ert139
Mortazavi A, Williams BA, Mccue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5(7):621–628. https://doi.org/10.1038/nmeth.1226
Murata J, Roepke J, Gordon H, De Luca V (2008) The leaf epidermome of Catharanthus roseus reveals its biochemical specialization. Plant Cell 20(3):524–542. https://doi.org/10.1105/tpc.107.056630
Ogawa M, Herai Y, Koizumi N, Kusano T, Sano H (2001) 7-Methylxanthine methyltransferase of coffee plants – gene isolation and enzymatic properties. J Biol Chem 276(11):8213–8218
Pardo-Hernández M, López-Delacalle M, Rivero RM (2020) ROS and NO Regulation by Melatonin Under Abiotic Stress in Plants. Antioxidants 9(11):1078. https://doi.org/10.3390/antiox9111078
Qin GJ et al (2005) An indole-3-acetic acid carboxyl methyltransferase regulates Arabidopsis leaf development. Plant Cell 17(10):2693–2704. https://doi.org/10.1105/tpc.105.034959
Quevillon E, Silventoinen V, Pillai S, Harte N, Mulder N, Apweiler R, Lopez R (2005) InterProScan: protein domains identifier. Nucleic Acids Res 33:W116–W120. https://doi.org/10.1093/nar/gki442
Ramakrishna A, Ravishankar GA (2011) Influence of abiotic stress signals on secondary metabolites in plants. Plant Signal Behav 6:1720–1731. https://doi.org/10.4161/psb.6.11.17613
Rivas-San Vicente M, Plasencia J. (2011) Salicylic acid beyond defence: its role in plant growth and development. J Exp Bot 62(10):3321–3338
Schmutz J et al (2014) A reference genome for common bean and genome-wide analysis of dual domestications. Nat Genet 46(7):707–713. https://doi.org/10.1038/ng.3008
Seo HS, Song JT, Cheong JJ, Lee YH, Lee YW, Hwang I, Lee JS, Choi YD (2001) Jasmonic acid carboxyl methyltransferase: a key enzyme for jasmonate-regulated plant responses. P Natl Acad Sci USA 98(8):4788–4793. https://doi.org/10.1073/pnas.081557298
Shafi A, Singh AK, Zahoor I (2021) Melatonin: Role in Abiotic Stress Resistance and Tolerance. In: Aftab T, Hakeem KR (ed) Plant Growth Regulators, Springer, Cham, pp 239–273. https://doi.org/10.1007/978-3-030-61153-8_12
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504
Shen J, Chen D, Zhang X, Song L, Dong J, Xu Q, Wang W (2021) Mitigation of salt stress response in upland cotton (Gossypium hirsutum) by exogenous melatonin. J Plant Res 134(4):857–871
Shi H, Jiang C, Ye T, Tan DX, Reiter RJ, Zhang H, Chan Z (2015) Comparative physiological, metabolomic, and transcriptomic analyses reveal mechanisms of improved abiotic stress resistance in bermudagrass [Cynodon dactylon (L). Pers.] by exogenous melatonin. J Exp Bot 66(3):681–694. https://doi.org/10.1093/jxb/eru373
Singewar K, Moschner CR, Hartung E, Fladung M (2021) Genome-wide bioinformatics analysis revealed putative substrate specificities of SABATH and MES family members in silver birch (Betula pendula). Silvae Genet 70(1):57–74. https://doi.org/10.2478/sg-2021-0005
Suyama M, Torrents D, Bork P (2006) PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res 34:W609–W612. https://doi.org/10.1093/nar/gkl315
Takahashi F, Kuromori T, Urano K, Yamaguchi-Shinozaki K, Shinozaki K (2020) Drought stress responses and resistance in plants: From cellular responses to long-distance intercellular communication. Front Plant Sci 11:556972. https://doi.org/10.3389/fpls.2020.556972
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: Molecular Evolutionary Genetics Analysis Using Maximum Likelihood, Evolutionary Distance, and Maximum Parsimony Methods. Mol Biol Evol 28(10):2731–2739. https://doi.org/10.1093/molbev/msr121
Tan DX, Hardeland R, Manchester LC, Korkmaz A, Ma SR, Rosales-Corral S, Reiter RJ (2012) Functional roles of melatonin in plants, and perspectives in nutritional and agricultural science. J Exp Bot 63(2):577–597. https://doi.org/10.1093/jxb/err256
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X Windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25(24):4876–4882. https://doi.org/10.1093/nar/25.24.4876
Valliyodan B et al (2019) Construction and comparison of three reference-quality genome assemblies for soybean. Plant J 100(5):1066–1082. https://doi.org/10.1111/tpj.14500
Varbanova M et al (2007) Methylation of gibberellins by Arabidopsis GAMT1 and GAMT2. Plant Cell 19(1):32–45. https://doi.org/10.1105/tpc.106.044602
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93(1):77–78. https://doi.org/10.1093/jhered/93.1.77
Wang B, Wang SQ, Wang ZZ (2017) Genome-wide comprehensive analysis the molecular phylogenetic evaluation and tissue-specific expression of SABATH gene family in Salvia miltiorrhiza. Genes-Basel 8(12). https://doi.org/10.3390/genes8120365
Wang P, Yin LH, Liang D, Li C, Ma FW, Yue ZY (2012) Delayed senescence of apple leaves by exogenous melatonin treatment: toward regulating the ascorbate-glutathione cycle. J Pineal Res 53(1):11–20. https://doi.org/10.1111/j.1600-079X.2011.00966.x
Weeda S, Zhang N, Zhao X, Ndip G, Guo Y, Buck GA, Ren S (2014) Arabidopsis transcriptome analysis reveals key roles of melatonin in plant defense systems. PloS one 9(3): e93462. https://doi.org/10.1371/journal.pone.0093462
Wei XM, Tao KL, Zhang JW, Lu SG, Chen SY, Liao JG (2021) Identification of SABATH family members in Solanum lycopersicum and their expression patterns under abiotic/biotic stresses. Plant Mol Biol Rep 39(2):403–418
Yang L, Bu S, Zhao S, Wang N, Xiao J, He F, Gao X (2022) Transcriptome and physiological analysis of increase in drought stress tolerance by melatonin in tomato. Plos One 17(5), e0267594
Yang Y, Yuan JS, Ross J, Noel JP, Pichersky E, Chen F (2006) An Arabidopsis thaliana methyltransferase capable of methylating farnesoic acid. Arch Biochem Biophys 448(1–2):123–132. https://doi.org/10.1016/j.abb.2005.08.006
Yang ZH (2007) PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24(8):1586–1591. https://doi.org/10.1093/molbev/msm088
Yang ZH, Nielsen R (2000) Estimating synonymous and nonsynonymous substitution rates under realistic evolutionary models. Mol Biol Evol 17(1):32–43. https://doi.org/10.1093/oxfordjournals.molbev.a026236
Zhang HJ, Zhang NA, Yang RC, Wang L, Sun QQ, Li DB, Guo YD (2014) Melatonin promotes seed germination under high salinity by regulating antioxidant systems, ABA and GA 4 interaction in cucumber (C ucumis sativus L.). J Pineal Res 57(3): 269–279. https://doi.org/10.1111/jpi.12167
Zhao N (2008) Comparative functional genomics of the SABATH family of methyltransferases in plants
Zhao N, Boyle B, Duval I, Ferrer JL, Lin H, Seguin A, Mackay J, Chen F (2009) SABATH methyltransferases from white spruce (Picea glauca): gene cloning, functional characterization and structural analysis. Tree Physiol 29(7):947–957. https://doi.org/10.1093/treephys/tpp023
Zhao N, Ferrer JL, Moon HS, Kapteyn J, Zhuang XF, Hasebe M, Stewart CN, Gang DR, Chen F (2012) A SABATH methyltransferase from the moss Physcomitrella patens catalyzes S-methylation of thiols and has a role in detoxification. Phytochemistry 81:31–41. https://doi.org/10.1016/j.phytochem.2012.06.011
Zhao C, Guo H, Wang J, Wang Y, Zhang R (2021) Melatonin enhances drought tolerance by regulating leaf stomatal behavior, carbon and nitrogen metabolism, and related gene expression in maize plants. Front Plant Sci 12
Acknowledgements
We are grateful to ETU High Technology Application and Research Center and Atatürk University Faculty of Agriculture for providing suitable working conditions for this study. We thank Dr. Fatma Necmiye Kaci for English language editing.
Funding
This work was supported by Erzurum Technical University Scientific Research Projects Grant Number 2021/1.
Author information
Authors and Affiliations
Contributions
ASA, EI, MA, and IB conducted the experiment and wrote the paper. EG, SM, AGK, and EY participated in making the bioinformatic analysis and stress applications; all the authors participated in the discussion of the article.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key Message
The SABATH gene family is a plant-specific class of small small-molecule methyltransferases. They may be important for the growth and development of plants under different stress conditions because they catalyze the methylation process.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Aygören, A.S., Güneş, E., Muslu, S. et al. Genome-Wide Analysis and Characterization of SABATH Gene Family in Phaseolus vulgaris Genotypes Subject to Melatonin under Drought and Salinity Stresses. Plant Mol Biol Rep 41, 242–259 (2023). https://doi.org/10.1007/s11105-022-01363-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-022-01363-5