Abstract
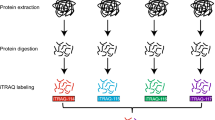
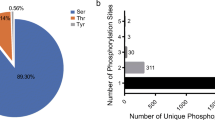
Legumes are unique in their ability to establish symbiotic interactions with rhizobacteria, providing a source of assimilable nitrogen; this symbiosis is regulated by complex signaling process between the plant and the bacteria. The participation of specific protein kinases during the initial steps of the nodulation process has been established. However, their substrates or the signaling networks implicated are not fully understood. Herein, a phosphoproteomic analysis of Phaseolus vulgaris roots treated for 24 h with specific Nod factors was performed using an immobilized metal ion affinity chromatography enrichment and two-dimensional gel electrophoresis approach with mass spectrometry identification. A total of 33 protein spots showing more than 1.5-fold shift were identified (17 protein spots in which the relative abundance increased and 16 that decreased). The majority of the identified root phosphoproteins displaying an increased relative abundance are presumed to have functions related to the biosynthesis and folding of proteins, energy metabolism, or cytoskeleton rearrangements, which reflect the metabolic status of the roots as being part of the developmental processes leading to nodule initiation and the importance of cytoskeleton rearrangement in the P. vulgaris–rhizobia symbiosis. The proteins in which relative abundance decreased are associated with defense and oxido-reduction processes, which could indicate a suppression of plant defense responses during the establishment of the rhizobia–legume interaction and an increase of reactive oxygen species production.




Similar content being viewed by others
References
Afzal A, Srour A, Natarajan A, Saini N, Iqbal M, Geisler M, El Shemy A, Lightfoot D (2012) Resistance to soybean Cyst Nematode: Rhg1. J Plant Gen Sci 1:39–45
Aker J, de Vries S (2008) Plasma membrane receptor complexes. Plant Physiol 147:1560–1564
Capoen W, Goormachtig S, De Rycke R, Schroeyers K, Holsters M (2005) SrSymRK, a plant receptor essential for symbiosome formation. P Natl Acad Sci USA 102:10369–10374
Cárdenas L, Domínguez J, Quinto C, López-Lara IM, Lugtenberg BJ, Spaink HP, Rademaker GJ, Haverkamp J, Thomas-Oates J (1995) Isolation, chemical structures and biological activity of the lipo-chitin oligosaccharide nodulation signals from Rhizobium etli. Plant Molecular Biology 29:453–464
Cárdenas L, Vidali L, Domínguez J, Pérez H, Sánchez F, Hepler P, Quinto C (1998) Rearrangement of actin microfilaments in plant root hairs responding to Rhizobium etli nodulation signals. Plant Physiol 116:871–877
Cárdenas L, Feijó JA, Kunkel JG, Sánchez F, Holdaway-Clarke T, Hepler P, Quinto C (1999) Rhizobium nod factors induce increases in intracellular free calcium and extracellular calcium influxes in bean root hairs. The Plant Journal 19:347–352
Cárdenas L, Martinez A, Sanchez F, Quinto C (2008) Fast, transient and specific intracellular ROS changes in living root hair cells responding to Nod factors (NFs). Plant J 56:802–813
Carol J, Dolan L (2006) The role of reactive oxygen species in cell growth: lessons from root hairs. J Exp Bot 57:1829–1834
Collins MO, Yu L, Coba MP, Husi H, Campuzano I, Blackstock WP, Choudhary JS, Grant SG (2005) Proteomic analysis of in vivo phosphorylated synaptic proteins. J Biol Chem 280:5972–5982
Dalton D, Russell S, Hanus F, Pascoe G, Evans H (1986) Enzymatic reactions of ascorbate and glutathione that prevent peroxide damage in soybean root nodules. P Natl Acad Sci USA 83:3811–3815
De Carvalho-Niebel F, Timmers A, Chabaud M, Defaux-Petras A, Barker DG (2002) The Nod factor-elicited annexin MtAnn1 is preferentially localised at the nuclear periphery in symbiotically activated root tissues of Medicagotruncatula. Plant J 32:343–352
Dubrovska A, Soucheinyskyi S (2005) Efficient enrichment of intact phosphorylated proteins by modified immobilized metal-affinity chromatography. Proteomics 5:4678–4683
Ehrhardt D, Wais R, Long S (1996) Calcium spiking in plant root hairs responding to rhizobium nodulation signals. Cell 85:673–681
Endre G, Kereszt A, Kevei Z, Mihacea S, Kalo P, G and Kiss (2002) A receptor kinase gene regulating symbiotic nodule development. Nature 417:962–966
Esseling J, Lhuissier F, Emons A (2003) Nod factors-induced root hair curling: continuous polar growth towards the point of Nod factor application. Plant Physiol 132:1982–1988
Farag M, Huhman D, Dixon R, Sumner L (2008) Metabolomics reveals novel pathways and differential mechanistic and elicitor-specific responses in phenylpropanoid and isoflavonoid biosynthesis in Medicagotruncatula cell cultures. Plant Physiol 146:387–402
Felle H, Kondorosi E, Kondorosi A, Schultze M (1998) The role of ion fluxes in nod factor signalling in Medicago sativa. Plant J 13:455–463
Goetting-Minesky MP, Mullin BC (1994) Differential gene expression in an actinorhizal symbiosis: evidence for a nodule-specific cysteine proteinase. Proc Natl Acad Sci U S A 91:9891–9895
Guerrera IC, Predic-Atkinson J, Kleiner O, Soskic V, Godovac-Zimmermann J (2005) Enrichment of phosphoproteins for proteomic analysis using immobilized Fe (III)-affinity adsorption chromatography. J Proteome Res 4:1545–1553
Kaetzel M, Mo Y, Mealy T, Campos B, Bergsma-Schutter W, Brisson A, Dedman J (2001) Phosphorylation mutants elucidate the mechanism of annexin IV-mediated membrane aggregation. Biochemistry 40:41925–44199
Koltai H, Dhandaydham M, Opperman C, Thomas J, Bird D (2001) Overlapping plant signal transduction pathways induced by a parasitic nematode and rhizobial endosymbiont. Mol Plant-Microbe Interac 14:1168–1177
Konopka-Postupolska D, Greg-Clark G, Hofmann A (2011) Structure, function and membrane interactions of plant annexins: an update. Plant Sci 181:230–241
Libault M, Farmer A, Joshi T, Takahashi K, Langley RJ, Franklin L, He J et al (2010) An integrated transcriptome atlas of the crop model Glycine max and its use in comparative analyses in plants. Plant J 63:86–99
Limpens E, Mirabella R, Fedorova E, Franken C, Franssen H, Bisseling T, Geurts R (2005) Formation of organelle-like N2-fixing symbiosomes in legume root nodules is controlled by DMI2. P Natl Acad Sci USA 102:10375–10380
Lukowitz W, Gillmor C, Scheible W (2000) Positional cloning in Arabidopsis. Why it feels good to have a genome initiative working for you. Plant Physiol 123:795–805
Madeo F, Schlauer J, Zischka H, Mecke D, Fröhlich K (1998) Tyrosine phosphorylation regulates cell cycle-dependent nuclear localization of Cdc48p. Mol Biol Cell 9:131–141
Madsen E, Madsen L, Radutoiu S, Olbryt M, Rakwalska M, Szczyglowski K, Sato S et al (2003) A receptor kinase gene of the LysM type is involved in legume perception of rhizobial signals. Nature 425:637–640
Manning G, Whyte D, Martinez R, Hunter T, Sudarsanam S (2002) The protein kinase complement of the human genome. Science 298:1912–1934
Miller JB, Pratap A, Miyahara A, Zhou L, Bornemann S, Morris RJ, Oldroyd GE (2013) Calcium/Calmodulin-dependent protein kinase is negatively and positively regulated by calcium, providing a mechanism for decoding calcium responses during symbiosis signaling. Plant Cell 25:5053–5066
Montiel J, Nava N, Cárdenas L, Sánchez-López R, Arthikala MK, Santana O, Sánchez F, Quinto C (2012) A Phaseolus vulgaris NADPH oxidase gene is required for root infection by Rhizobia. Plant Cell Physiol 53:1751–1767
Nakagami H, Sugiyama N, Mochida K, Daudi A, Yoshida Y, Toyoda Y, Tomita M et al (2010) Large-scale comparative phosphoproteomics identifies conserved phosphorylation sites in plants. Plant Physiol 153:1161–1174
Nguyen T, Brechenmacher L, Aldrich J, Clauss T, Gritsenko M, Hixson K, Libault M et al (2012) Quantitative phosphoproteomic analysis of soybean root hairs inoculated with Bradyrhizobium japonicum. Mol Cel Proteomics 11:1140–1155
Oh C, Lee H, Kim H, An C (2004) Isolation and characterization of a root nodule-specific cysteine proteinase cDNA from soybean. J Plant Biol 47:216–220
Parida A, Das A (2005) Salt tolerance and salinity effects on plants: a review. Ecotox Environ Safe 60:324–349
Peleg-Grossman S, Volpin H, Levine A (2007) Root hair curling and Rhizobium infection in Medicago truncatula are mediated by phosphatidylinositide-regulated endocytosis and reactive oxygen species. J Exp Bot 58:1637–1649
Radutoiu S, Madsen LH, Madsen EB, Felle HH, Umehara Y, Gronlund M, Sato S et al (2003) Plant recognition of symbiotic bacteria requires two LysM receptor-like kinases. Nature 425:585–592
Ramu K, Peng H, Cook D (2002) Nod factor induction of reactive oxygen species production is correlated with expression of the early nodulin gene rip1 in Medicago truncatula. Mol Plant-Microbe Interact 15:522–528
Rose C, Venkateshwaran M, Volkening J (2012) Rapid phosphoproteomic and transcriptomic changes in the rhizobia-legume symbiosis. Mol Cel Proteomics 11:724–744
Sánchez-López R, Jáuregui D, Nava N, Alvarado-Affantranger X, Montiel J, Santana O, Sanchez F, Quinto C (2011) Down-regulation of SymRK correlates with a deficiency in vascular bundle development in Phaseolus vulgaris nodules. Plant Cell Environ 34:2109–2121
Serna-Sanz A, Parniske M, Peck SC (2011) Phosphoproteome analysis of Lotus japonicusroots reveals shared and distinct components of symbiosis and defense. Mol Plant-Microbe Interact 24:932–937
Stracke S, Kistner C, Yoshida S, Mulder L, Sato S, Kaneko T, Tabata S et al (2002) A plant receptor-like kinase required for both bacterial and fungal symbiosis. Nature 417:959–962
Timmers A, Auriac M, Truchet G (1999) Refined analysis of early symbiotic steps of the Rhizobium-Medicago interaction in relationship with microtubular cytoskeleton rearrangements. Development 126:3617–3628
Acknowledgments
This work was supported by Dirección General de Asuntos del Personal Académico – Universidad Nacional Autónoma de México (IN201312). We thank Drs. Federico Sánchez and Rosana Sánchez-Lopez for critical reading of the manuscript and Olivia Santana for the purification of R. etli Nod factors.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jáuregui-Zúñiga, D., Ortega-Ortega, Y., Pedraza-Escalona, M. et al. Phosphoproteomic Analysis in Phaseolus vulgaris Roots Treated with Rhizobium etli Nodulation Factors. Plant Mol Biol Rep 34, 961–969 (2016). https://doi.org/10.1007/s11105-016-0978-y
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-016-0978-y